Experiences and lessons learnt from bootstrapping randomeffects predictions

Experiences and lessons learnt from bootstrapping randomeffects predictions Robert Grant Senior Research Fellow, Kingston University & St George’s

Topics • • The problem: quality of stroke care Predictions from random effects Bootstrapping do-file Spotting errors from xtmelogit postestimation

The problem: quality of stroke care • • National clinical audit of stroke 203 hospitals provide data 10, 617 patients from the 2008 data analysed 26 binary quality indicators used in scoring

The problem: quality of stroke care National clinical audit of stroke 203 hospitals provide data 10, 617 patients from the 2008 data analysed 26 binary quality indicators used in scoring Is inter-hospital variation adequately summarised? • Can we depict uncertainty around scores / ranks of hospitals? • • •

“Multilevel principal components analysis” • Run mixed-effects models to adjust each indicator • Get predictions of each hospital’s performance as BLUPs (level-2 residuals) • Summarise by principal components analysis • Rank the scores on the component(s) • Bootstrap to capture correlation between indicators

Predictions from random effects • Logistic model • BLUPs – predictions of individual ui • Random effect distribution (mean 0, SD estimated) acts as empirical Bayes prior • Data in each cluster provide likelihood • BLUP can be mode (xtmelogit) or mean (gllamm) of the posterior distribution
![Predictions from random effects • Profile likelihood for the cluster effects Stata 11 [XT] Predictions from random effects • Profile likelihood for the cluster effects Stata 11 [XT]](http://slidetodoc.com/presentation_image_h/e91c89bf9bb82f19e4c4d82bfac244fa/image-7.jpg)
Predictions from random effects • Profile likelihood for the cluster effects Stata 11 [XT] manual, p. 277

Predictions from random effects • xtmelogit outcome covariate 1 covariate 2 || hospital: • predict mode_blup, reffects • gllamm outcome covariate 1 covariate 2, i(hospital) family(binomial) link(logit) adapt • gllapred mean_blup, u • xtmelogit. . . • predict offset, xb • statsby mle=_b[_cons], by(hospital) saving(ml): logit outcome, offset(offset)

Bootstrapping do-file • Program with bsample / bstat • Save individual resample and ‘parameters’ • Can be broken into, run on multiple machines

Do-file • Assemble your observed values as a single matrix pca mode 1 mode 2 mode 3 mode 4 mode 5 mode 6 mode 7 mode 8 mode 9 mode 10 mode 11 mode 12 mode 13 mode 14 mode 15 mode 16 mode 17 mode 18 mode 19 mode 20 mode 21 mode 22 mode 23 mode 24 mode 25 mode 26 if pickone, covariance components(1) matrix obsload=e(L) forvalues i=1/26 { scalar obsload`i'=obsload[`i', 1] } scalar obsload 27=e(rho) matrix obspca=(obsload 1, obsload 2, obsload 3, obsload 4, obsload 5, obsload 6, obsload 7, obsload 8, obsload 9, obsload 10, obsload 11, obsload 12, obsload 13, obsload 14, obsload 15, obsload 16, obsload 17, obsload 18, obsload 19, obsload 20, obsload 21, obsload 22, obsload 23, obsload 24, obsload 25, obsload 26, obsload 27) predict obsscore if pickone, score mkmat obsscore if pickone matrix define obs=obspca, obsscore'

Do-file • The `iteration’ macro lets you see how many resamples the computer has done. • Define your bootstrap program global iteration=1 capture: program drop myboot_mode program define myboot_mode, rclass display as result "Now running resample number $iteration" global iteration=$iteration + 1 preserve bsample, strata(hospital) capture: drop pickone egen pickone=tag(hospital) * save bsample_$iteration. dta, replace * Code then follows for the models, the PCA etc.

Do-file • now you use simulate to run your program many times and to save the outputs global startdate="`c(current_date)'" global starttime="`c(current_time)'" simulate load 1=bootload 1 load 2=bootload 2 load 3=bootload 3 * …omitting many, many lines of tedious code… score 203=bootscore 203, noisily reps(80) seed(1635) saving(modescorebootstrap. dta, replace): myboot_mode bstat, stat(obs) n(10617) estat bootstrap, all global iteration=$iteration - 1 display as result "This program ran $iteration bootstrap resamples" display as result "Starting on $startdate at $starttime" display as result "Ending on `c(current_date)' at `c(current_time)'"

Spotting errors • xtmelogit is faster than gllamm because mode doesn’t require full integration • A few extremely large BLUPs arise from nonconvergence of the profile likelihood • Can pick them out and draw up bootstrap confidence intervals from the remainder (may not be so easy in every situation? )
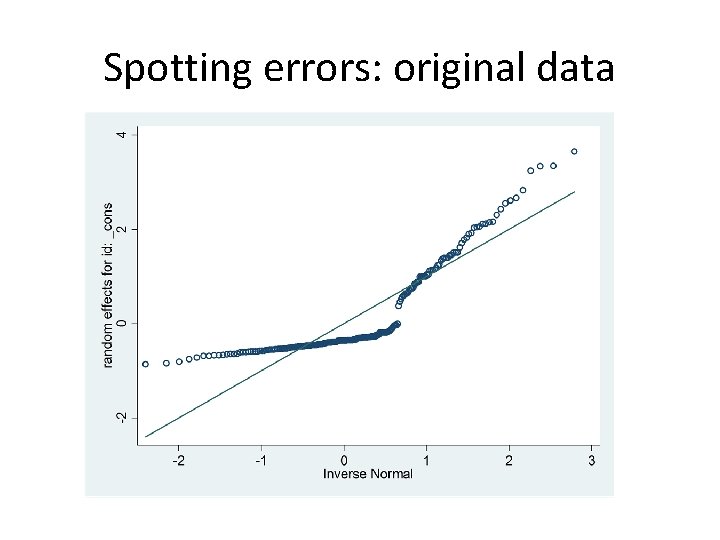
Spotting errors: original data
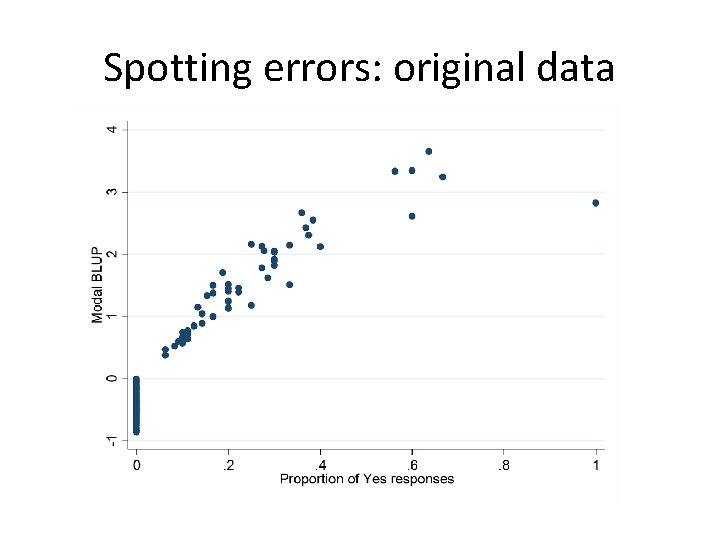
Spotting errors: original data
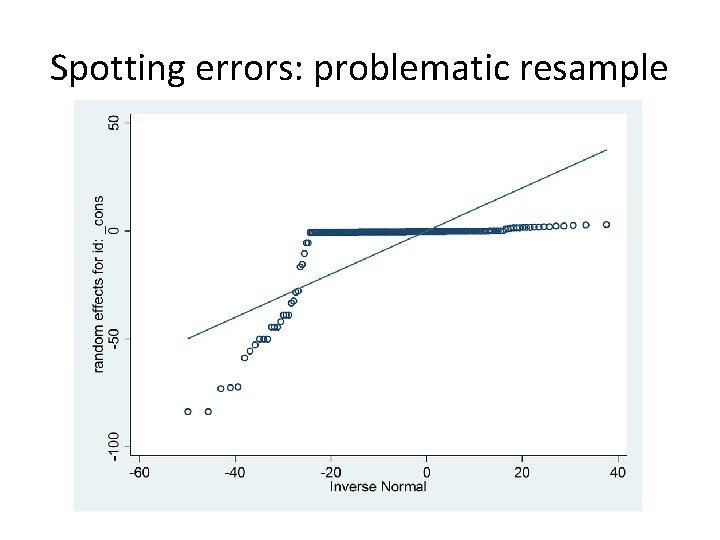
Spotting errors: problematic resample
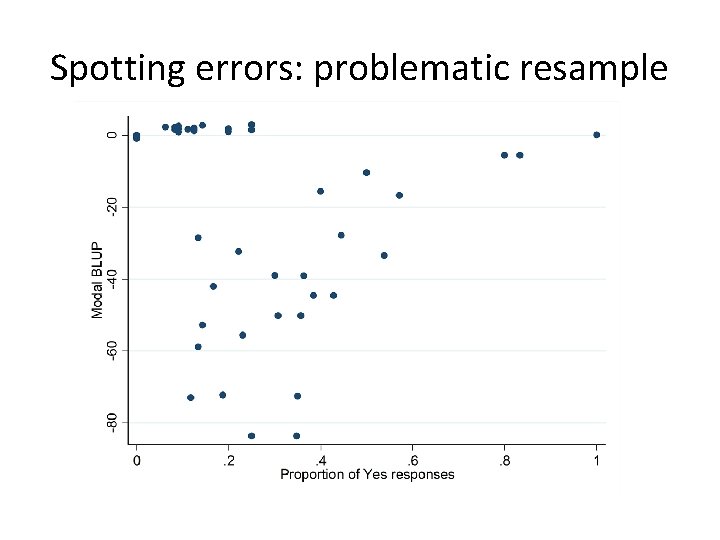
Spotting errors: problematic resample
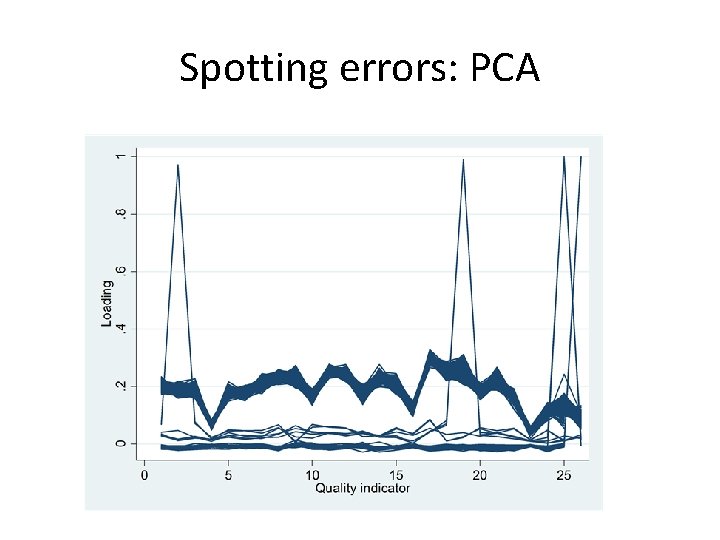
Spotting errors: PCA

Spotting errors • Consider multivariate outliers which may not be univariate ones • Methods suggested by Gnanadesikan & Kettenring (Biometrics 1972; 28(1): 81 -124. ) • Mahalanobis distances • Minor principal component scores
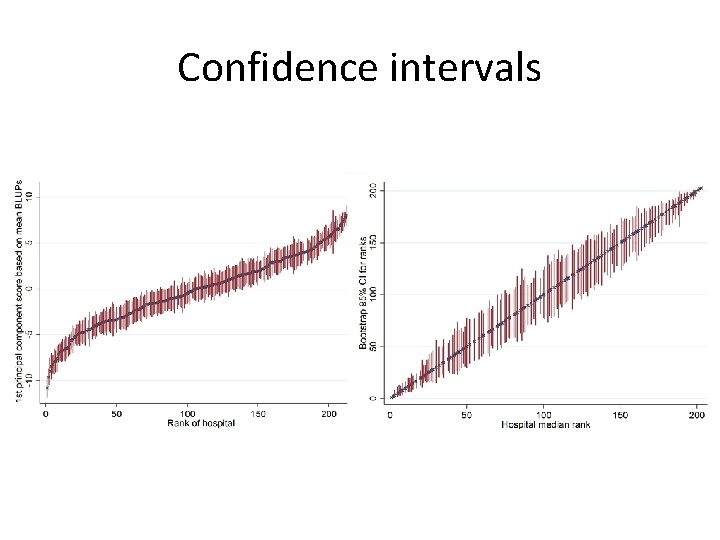
Confidence intervals

Confidence intervals • Normal approximate confidence intervals were often inappropriate • Percentile, bias-corrected (BC) and Bca confidence intervals often disagreed • Sometime degenerate to a point • The problem is mostly in the ranks • Because of an implicitly discontinuous transformation – no Edgeworth expansion?

Alternatives • Multivariate AND multilevel modelling (SEM? ) in a single step • Goldstein, Bonnet & Rocher. J Educ Behav Stat (2007); 32: 252 • runmlwin (MCMC) • Fully Bayesian specification

Acknowledgements • Thanks to Kristin Mac. Donald & colleagues at Stata. Corp for looking into the xtmelogit postestimation errors • Thanks to former colleagues at the Royal College of Physicians for supplying the stroke audit data • Thanks to Chris Frost & Rebecca Jones at LSHTM for guidance on the early versions of these analyses which went into my MSc.
- Slides: 23