Exercises Pairwise alignment Homology search BLAST Multiple alignment

Exercises • • • Pairwise alignment Homology search (BLAST) Multiple alignment (CLUSTAL W) Iterative Profile Search: Profile Search – – Pfam Prosite PSI-BLAST SAM

Overview Exercises Query Sequence Unknown Blast Sequence to search for close homologs PFAM PROSITE Search p. FAM, Prosite for conserved motifs You detected homology with an annotated protein family Clustal. W Make a multiple sequence alignment HMMer, PSSM PSI-blast Generate profile or HMM Profile Search database for remote homologs
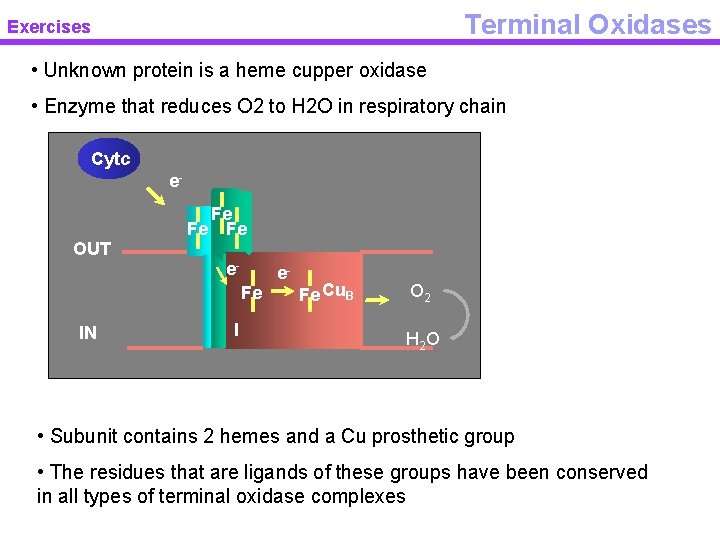
Terminal Oxidases Exercises • Unknown protein is a heme cupper oxidase • Enzyme that reduces O 2 to H 2 O in respiratory chain Cytc e. Fe Fe Fe OUT e. Fe IN I e- Fe Cu. B O 2 H 2 O • Subunit contains 2 hemes and a Cu prosthetic group • The residues that are ligands of these groups have been conserved in all types of terminal oxidase complexes
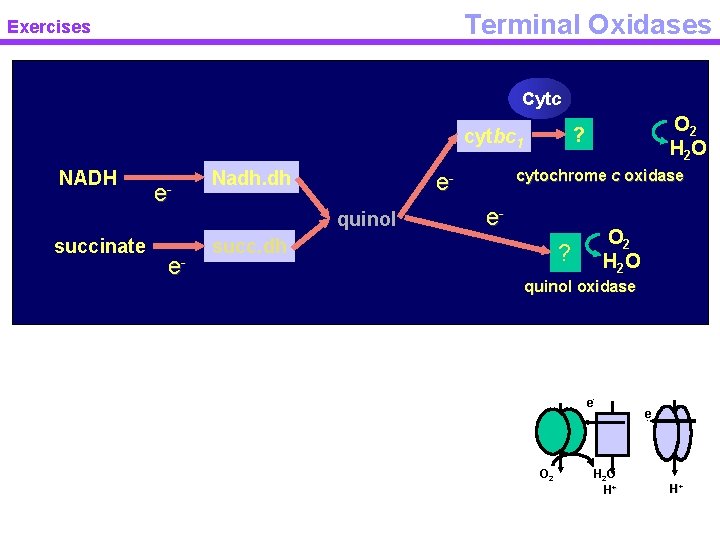
Terminal Oxidases Exercises Cytc NADH e- Nadh. dh succinate e- cytochrome c oxidase equinol O 2 H 2 O ? cytbc 1 e- succ. dh O 2 H 2 O ? quinol oxidase e- O 2 H 2 O H+ e- H+
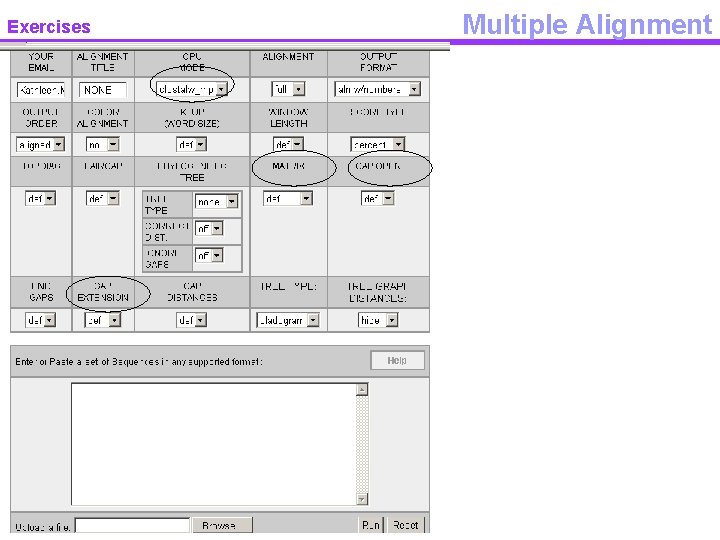
Exercises Multiple Alignment

Multiple Alignment Exercises Multiple alignment: standard gap cost Prosite pattern Ligands Cu center Ligands hemes

Exercises Multiple alignment: large gap cost Prosite pattern Multiple Alignment Ligands Cu center Ligands hemes

Exercises Tree based on subselection Phylogenetic Tree

Exercises PROSITE
Exercises Prosite

Exercises Prosite
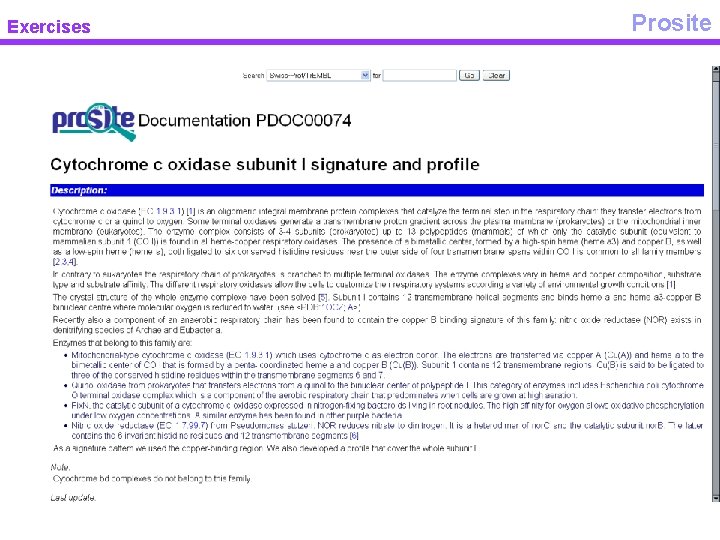
Exercises Prosite

Exercises Prosite
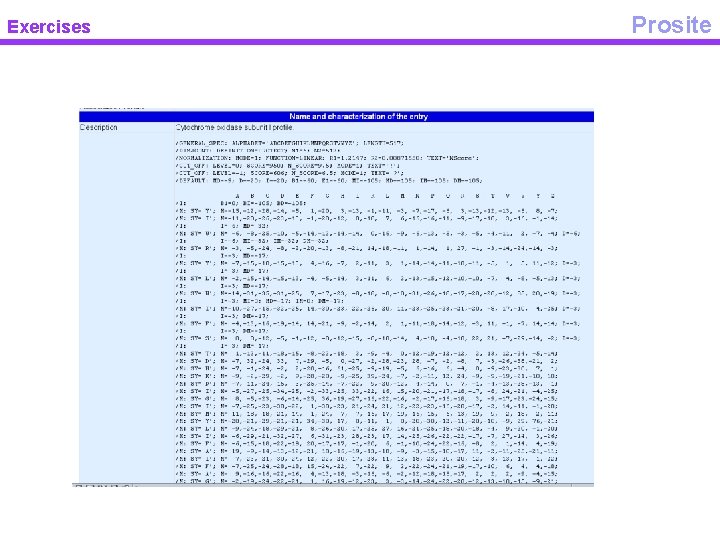
Exercises Prosite
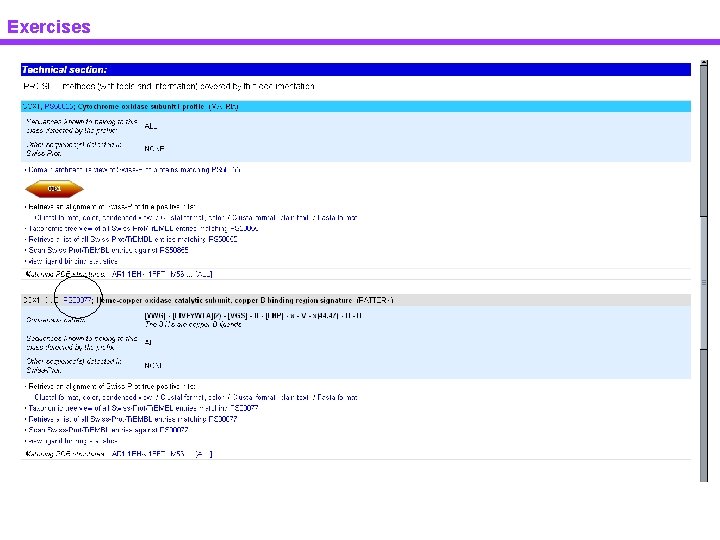
Exercises

Exercises

Exercises Prosite

Exercises Prosite domain
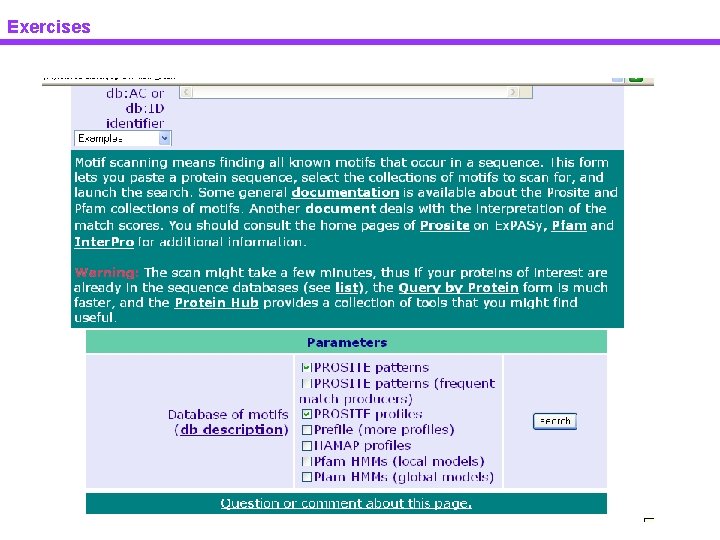
Exercises

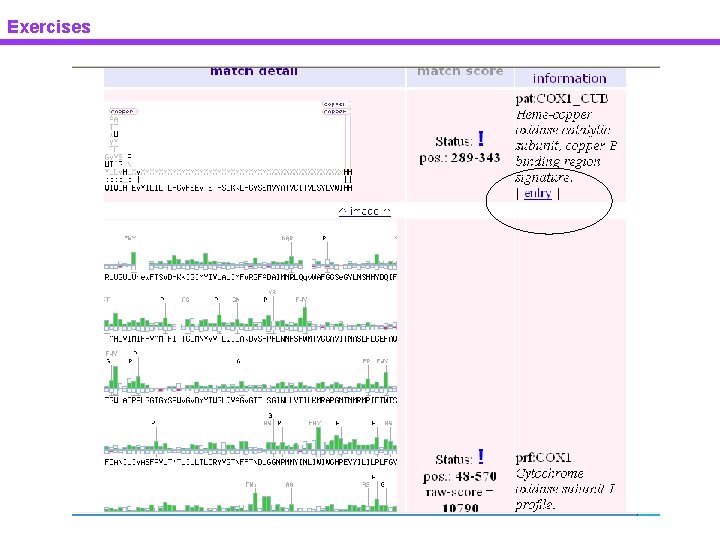
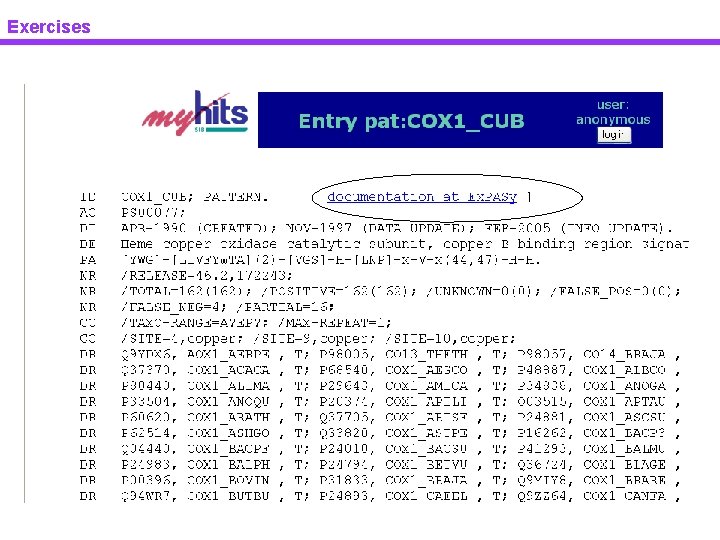
Exercises Pattern & profile
Exercises
Exercises

Exercises

Exercises p. FAM
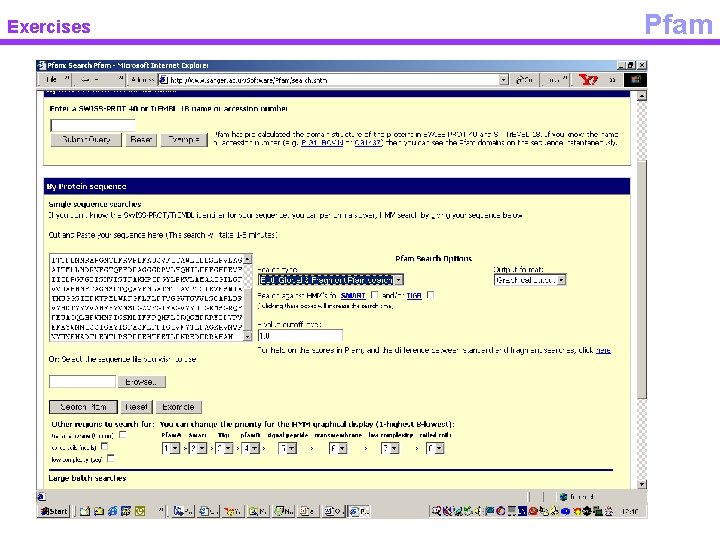
Exercises Pfam
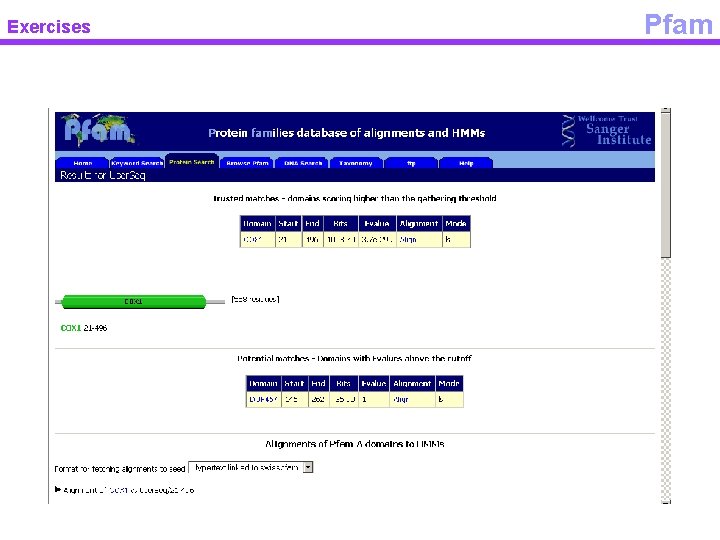
Exercises Pfam

Exercises Pfam
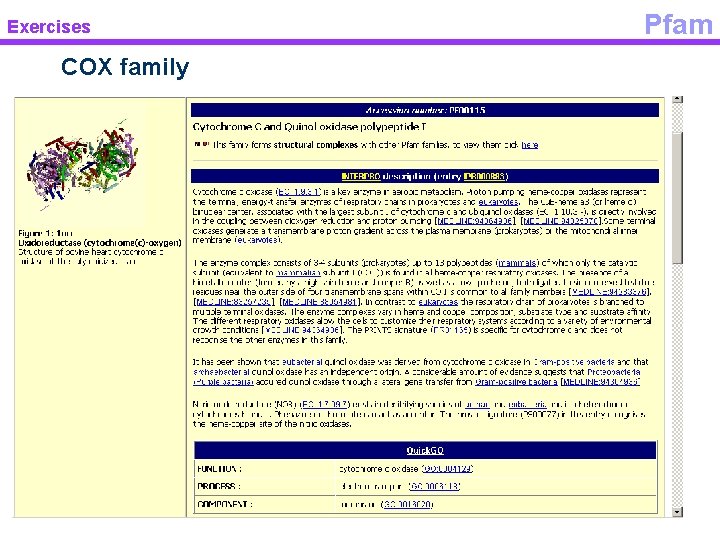
Exercises COX family Pfam
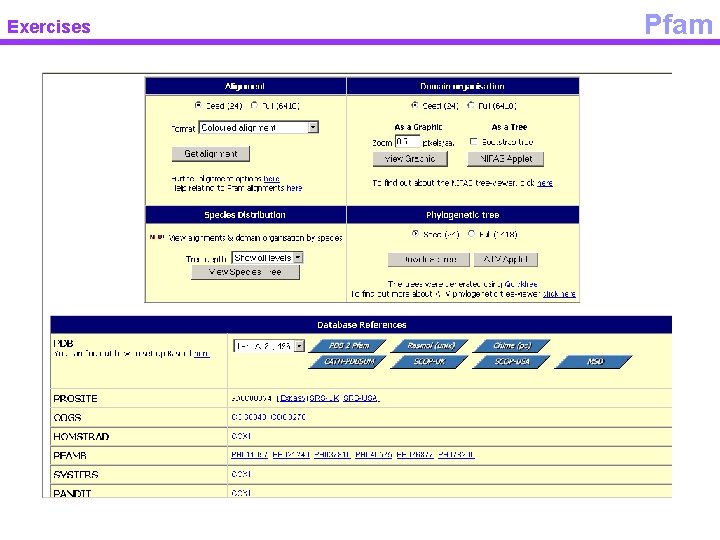
Exercises Pfam

Exercises Pfam

Exercises Pfam
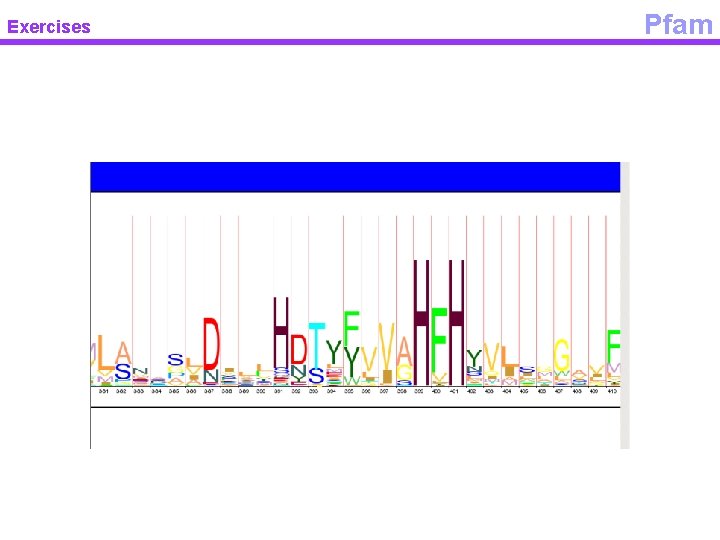
Exercises Pfam
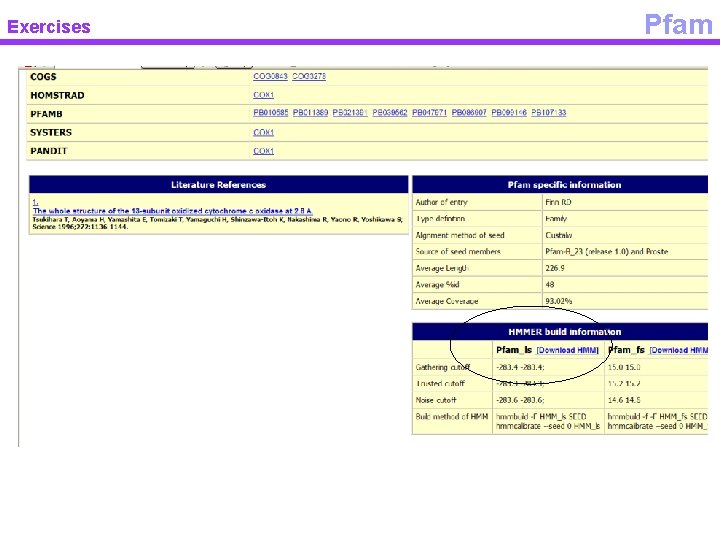
Exercises Pfam
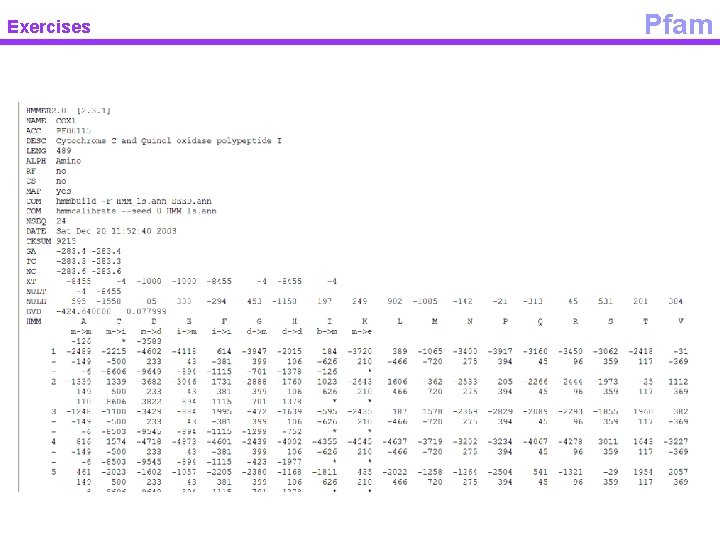
Exercises Pfam

Exercises BLOCKS
Exercises • Blocks
Exercises
Exercises
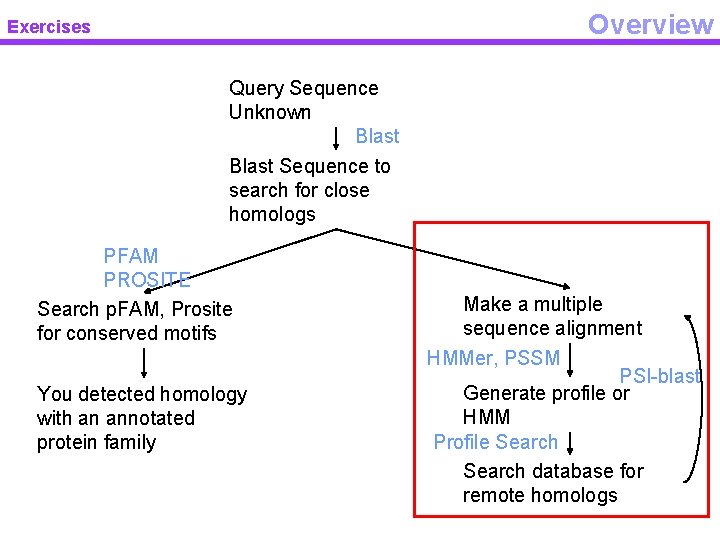
Overview Exercises Query Sequence Unknown Blast Sequence to search for close homologs PFAM PROSITE Search p. FAM, Prosite for conserved motifs You detected homology with an annotated protein family Make a multiple sequence alignment HMMer, PSSM PSI-blast Generate profile or HMM Profile Search database for remote homologs

Exercises PSI-BLAST

Exercises PSI-BLAST • PSI BLAST – Start from a single sequence – Blast it against NCBI – Select high scoring hits – Perform multiple alignment – Construct profile – Iterate and find remote homologs • Usually cut the sequence in pieces • Avoid to give as input multi domain proteins
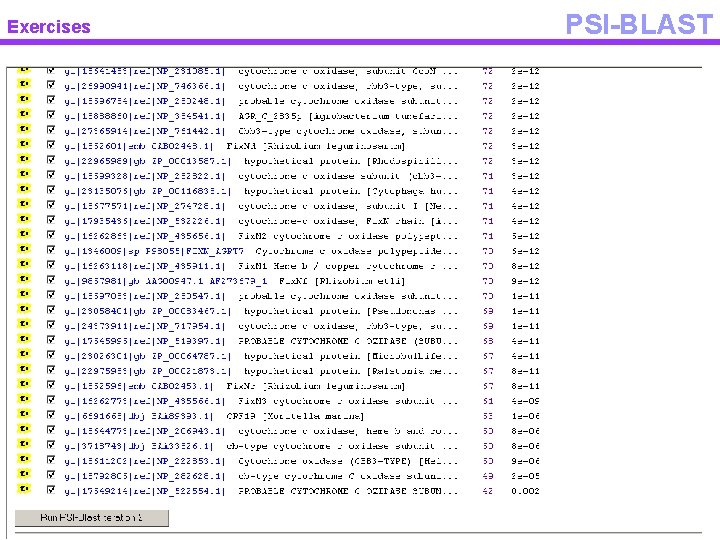
Exercises PSI-BLAST
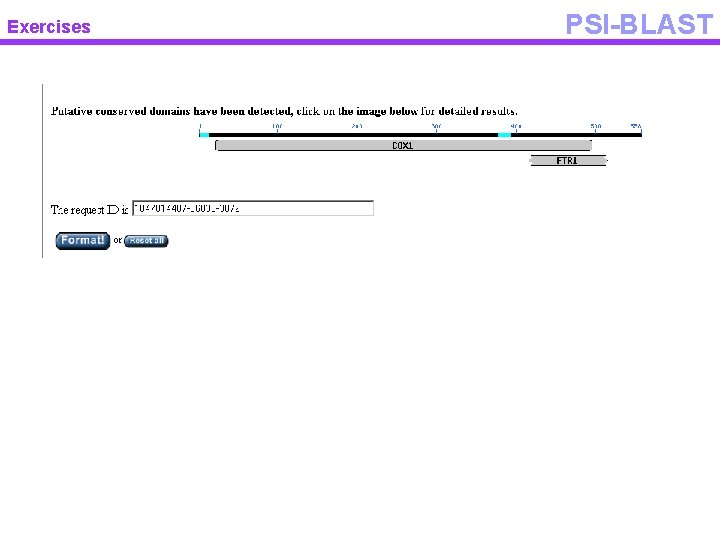
Exercises PSI-BLAST

Exercises PSI-BLAST

Exercises PSI-BLAST
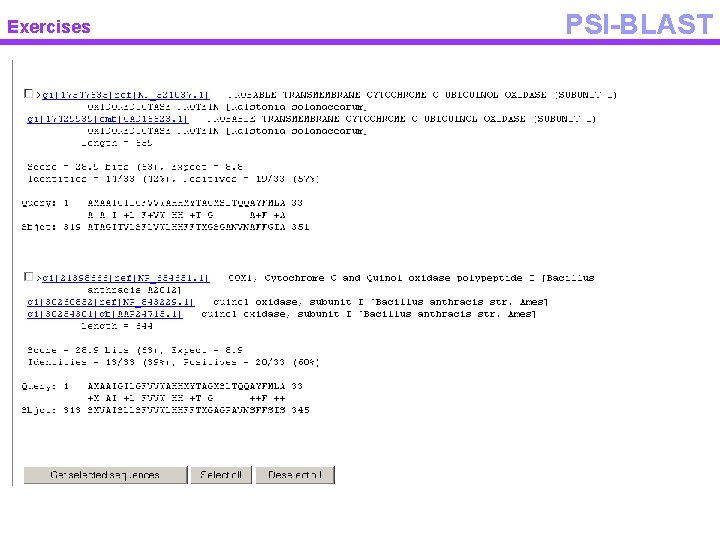
Exercises PSI-BLAST
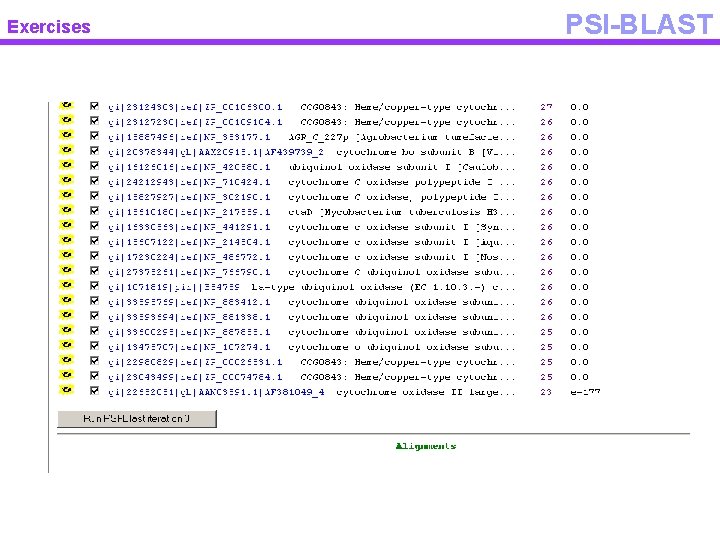
Exercises PSI-BLAST

Exercises SAM

Exercises SAM http: //www. cse. ucsc. edu/research/compbio/HMM-apps/HMM-applications. html

Exercises SAM

Exercises SAM
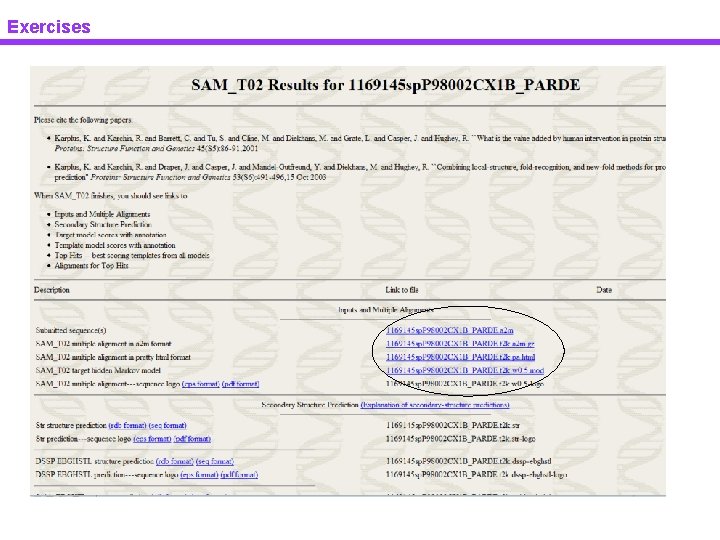
Exercises
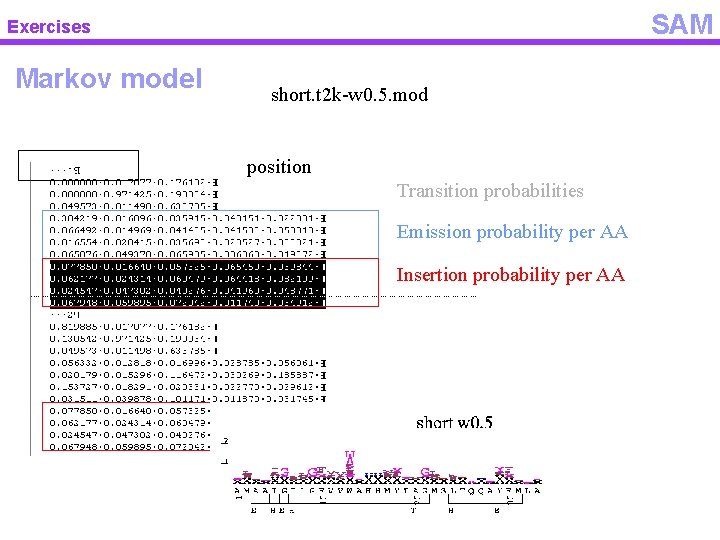
SAM Exercises Markov model short. t 2 k-w 0. 5. mod position Transition probabilities Emission probability per AA Insertion probability per AA

SAM Exercises input targets
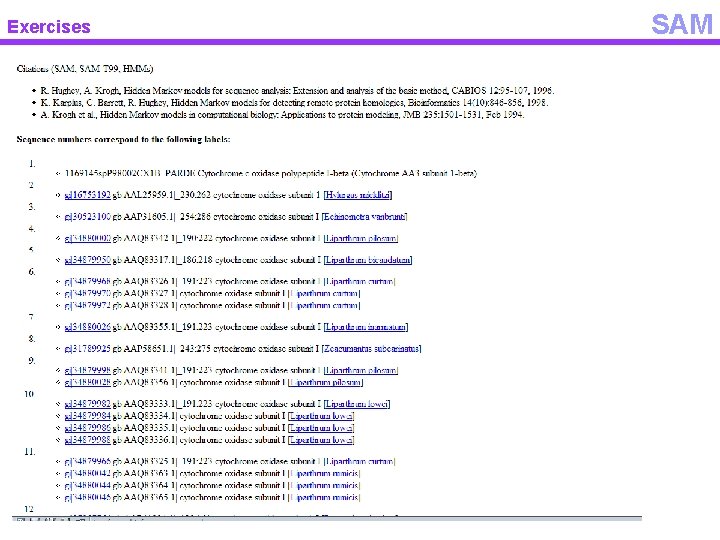
Exercises SAM

Exercises SAM
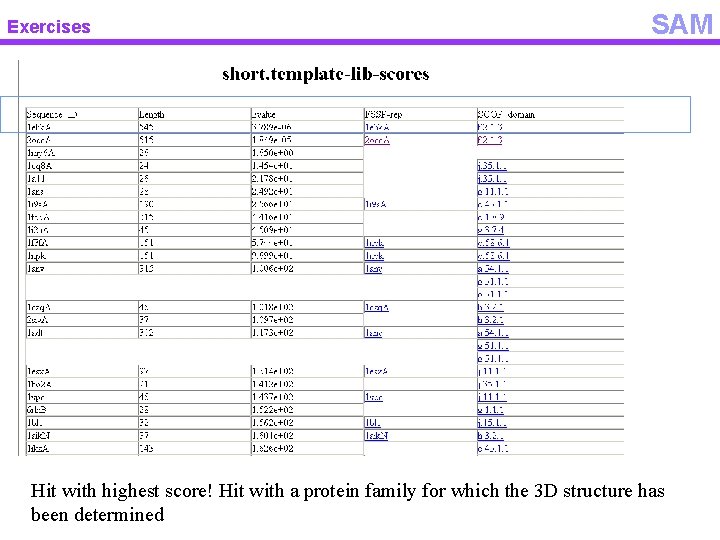
Exercises SAM Hit with highest score! Hit with a protein family for which the 3 D structure has been determined
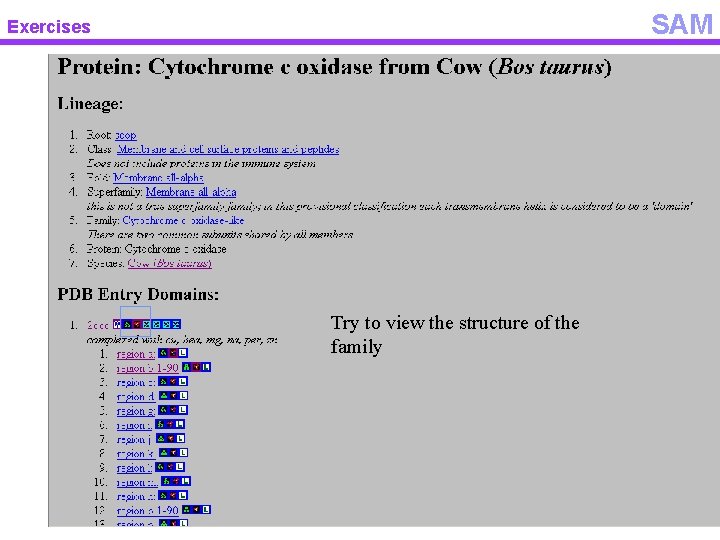
SAM Exercises Try to view the structure of the family
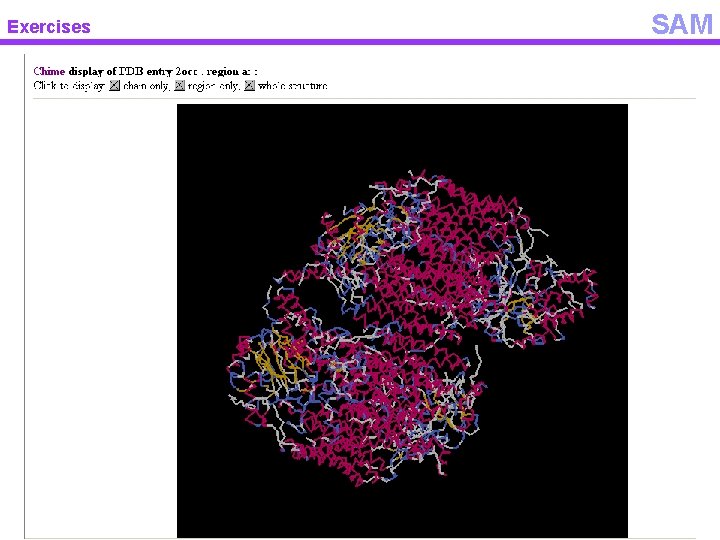
Exercises SAM
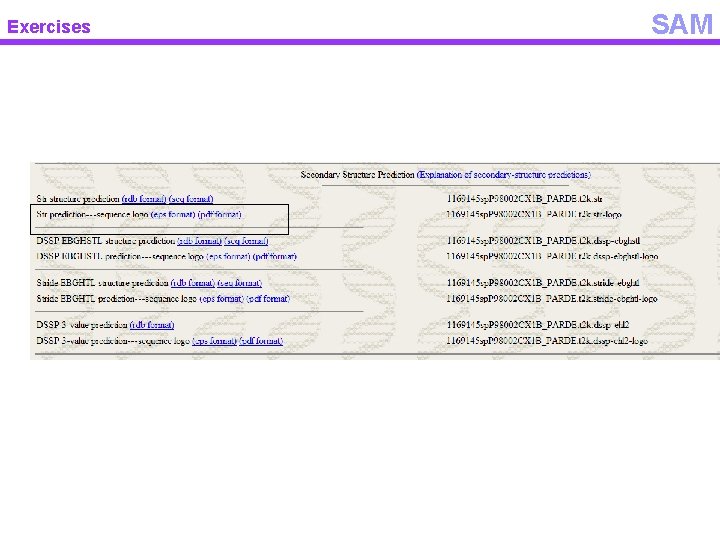
Exercises SAM
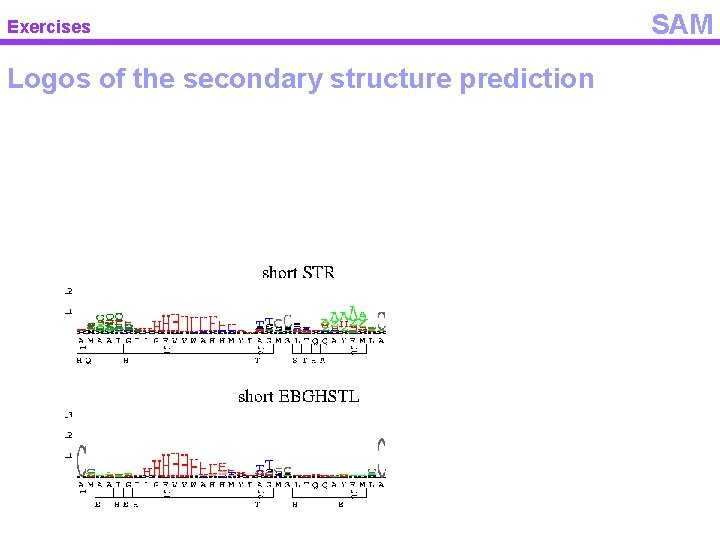
Exercises Logos of the secondary structure prediction SAM

Exercises SAM
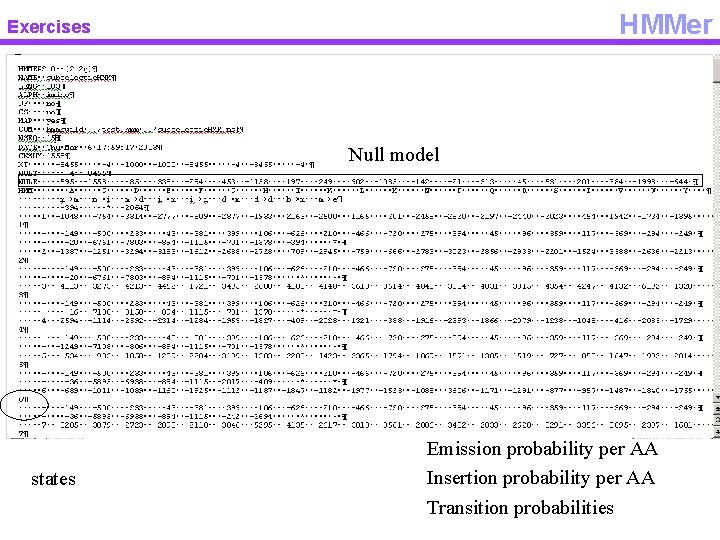
HMMer Exercises Null model states Emission probability per AA Insertion probability per AA Transition probabilities
Exercises
Exercises

Exercises

Exercises

Exercises
- Slides: 68