Evolutionary Models CS 498 SS Saurabh Sinha Models

Evolutionary Models CS 498 SS Saurabh Sinha

Models of nucleotide substitution • The DNA that we study in bioinformatics is the end(? ? )-product of evolution • Evolution is a very complicated process • Very simplified models of this process can be studied within a probabilistic framework • Allows testing of various hypotheses about the evolutionary process, from multi-species data Source: Ewens and Grant, Chapter 14.

Diversity in a population • There IS genetic variation between individuals in a population • But relatively little variation at nucl. level • E. g. , two humans differ at the nucl. level at one in 500 to 1000 nucls. • Roughly speaking, a single nucleotide dominates the population at a particular position in the genome

Substitution • Over long time periods, the nucleotide at a given position remains the same • But periodically, this nucleotide changes (over the entire population) • This is called “substitution”, i. e. , replacement of the predominant nucl. for that position with another predominant nucl.

Markov Chain to model substitution • Markov chain to describe the substitution process at a position • States are “a”, “c”, “g”, “t” • The chain “runs” in certain units of time, i. e. , the state may change from one time point to the next time point • The unit of time (difference between successive time points) may be arbitrary, e. g. , 20000 generations.

Markov Chain to model substitution • A symbol such as “pag” is the probability of a change from “a” to “g” in one unit of time • When studying two extant species, the evolutionary model has to provide the joint probability of the two species’ data • Sometimes, this is done by computing probability of the ancestor, starting from one extant species, and then the probability of the other extant species, starting from the ancestor • If we want to do this, the evolutionary process (model) must be “time reversible”: P(x)P(x->y) = P(y)P(y->x)

Jukes Cantor Model • Markov chain with four states: a, c, g, t • Transition matrix P given by: a g c t a 1 -3 g 1 -3 c 1 -3 t 1 -3

Jukes Cantor Model • is a parameter depending on what a “time unit” means. If time unit represents more #generations, will be larger • must be less than 1/3 though

Jukes Cantor Model • Whatever the current nucl is, each of the other three nucls are equally likely to substitute for it

Understanding the J-C Model • Consider a transition matrix P, and a probability vector v (a row vector) • What does w = v P represent ? • If v is the probability distribution of the 4 nucls (at a position) now, w is the prob. distr. at the next time step.

Understanding the J-C model • Suppose we can find a vector such that P = • If the probability distribution is , it will continue to remain at future times • This is called the stationary distribution of the Markov Chain

Understanding the J-C model • Check that = (0. 25, 0. 25) satisfies P = • Therefore, if a position evolves as per this model, for long enough, it will be equally likely to have any of the 4 nucls! • This is the very long term prediction, but can we write down what the position will be as a function of time (steps) ?

Spectral Decomposition • Recall that we found a such that P= • Such a vector is called an “eigenvector” of P, and the corresponding “eigenvalue” is 1. • In general, if v P = v (for scalar ), is called an eigenvalue, and v is a left eigenvector of P

Spectral decomposition • Similarly, if P u. T = u. T, then u is called a right eigenvector • In general, there may be multiple eigenvalues j and their corresponding left and right eigenvectors vj and uj • We can write P as

Spectral decomposition • Then, for any positive integer, it is true that • Why is Pn interesting to us ? • Because it tells us what the probability distribution will be after n time steps • If we started with v, then Pnv will be the prob. distr. after n steps

Back to the J-C model • We reasoned that = (. 25, . 25) is a left eigenvector for the eigenvalue 1. • Actually, the J-C transition matrix has this eigenvalue and the eigenvalue (1 -4 ), and if we do the math we get the spectral decomposition of P as:

Back to the J-C model • So, if we started with (1, 0, 0, 0), i. e. , an “a”, the probability that we’ll see an “a” at that position after n time steps is: 0. 25+0. 75(1 -4 )n • And the probability that the “a” would have mutated to say “c” is: 0. 25 - 0. 25(1 -4 )n

Substitution probability • As a function of time n, we therefore get • Pr(x -> y) = 0. 25 + 0. 75 (1 -4 )n if x = y • and = 0. 25 - 0. 25 (1 -4 )n otherwise • If n -> , we get back our (0. 25, 0. 25) calculation

More advanced models • The J-C model made highly “symmetric” assumptions, in its formulation of the transition matrix P • In reality, for example, “transitions” are more common than “transversions” – What are these? Purine = A or G. Pyrimidine = C or T. Transition is substitution in the same category; transversion is substitution across categories – Purines are similarly sized, and pyrimidines are similarly sized. More likely to be replaced by similar sized nucl. • The “Kimura” model captures this transition/transversion bias
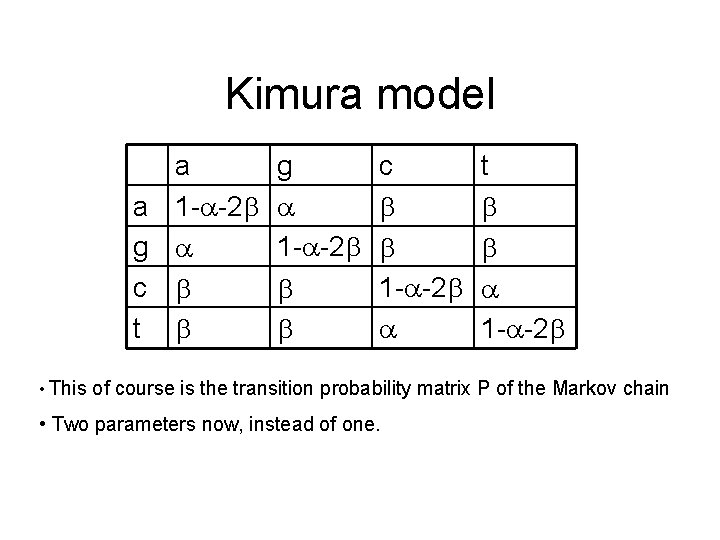
Kimura model a a 1 - -2 g c t • This g 1 - -2 c 1 - -2 t 1 - -2 of course is the transition probability matrix P of the Markov chain • Two parameters now, instead of one.

Kimura model • Again, one of the eigenvalues is 1, and the left eigenvector corresponding to it is = (. 25, . 25) • So again, the stationary distribution is uniform • P(x -> x) =. 25+. 25(1 -4 )n+. 5(1 -2( + ))n • P(x -> y) =. 25+. 25(1 -4 )n+. 5(1 -2( + ))n if x is a purine and y is the other purine

Even more advanced models • Get to greater levels of realism • Kimura model still has a uniform stationary distribution, which is not true of real data • One extension: purine to pyrimidine subst. prob. is different from pyrimidine to purine subst. prob. – This leads to a non-uniform stationary probability
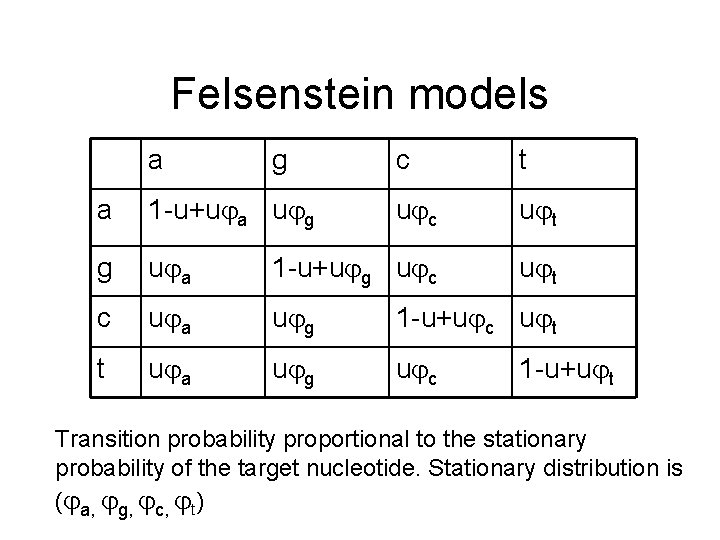
Felsenstein models a g c t u c u t a 1 -u+u a u g g u a 1 -u+u g u c c u a u g 1 -u+u c u t t u a u g u c 1 -u+u t Transition probability proportional to the stationary probability of the target nucleotide. Stationary distribution is ( a, g, c, t)

Reversible models • Many inference procedures require that the evolutionary model be time reversible • What does this mean?

Reversible Markov Chain Looks like time has been reversed. That is, if we can find a such that The models we have seen today all have this property. Source: Wikipedia
- Slides: 25