Evolutionary Informatics Supporting Interoperability in Evolutionary Analysis Working

Evolutionary Informatics: Supporting Interoperability in Evolutionary Analysis Working Group Members Jon Eisen (“phylogenomics”) Joe Felsenstein (PHYLIP) Mark Holder (GARLI) Sergei Kosakovsky Pond (Hy. Phy) Sudhir Kumar (MEGA) Paul Lewis (NCL) Aaron Mackey (Bio. Perl, GMOD) David Maddison (Mesquite) Wayne Maddison (Mesquite) Enrico Pontelli (CDAO) Andrew Rambaut (BEAST) Arlin Stoltzfus (Bio: : NEXUS) David Swofford (PAUP*) Rutger Vos (Bio: : Phylo) Fourth Xuhua Xia (DAMBE) Christian Zmasek (ATV, meeting RIO) NESCent staff Hilmar Lapp Todd Vision WG colleagues Brandon Chisham Brian Devries Gopal Gupta Peter E. Midford William Piel Francisco Prosdocimi Julie Thompson Derrick Zwickl DB Interop Hackathon Jim Balhoff Lucie Chan Dave Clements Karen Cranston Sam Donnelly Vladimir Gapeyev Karla Gendler Vivek Gopalan Roger Hyam Mark Jensen Greg Jordan Matt Kosnik Sheldon Mc. Kay Ryan Scherle Katja Schulz Katja Seltmann Jeet Sukumaran Matt Yoder
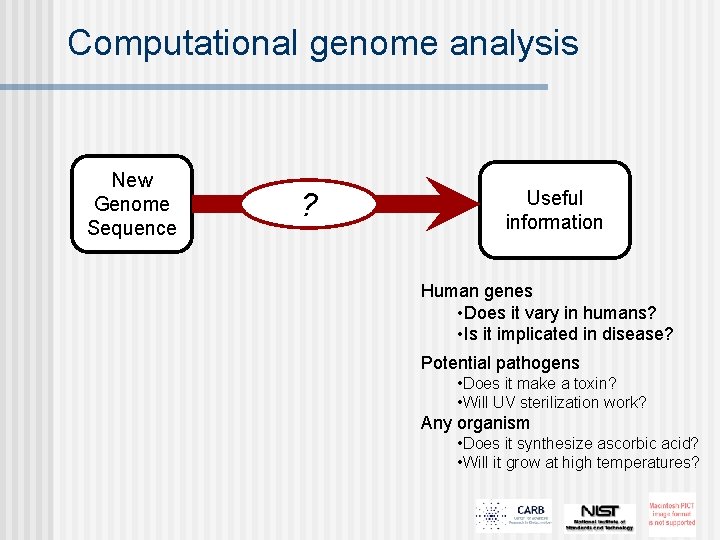
Computational genome analysis New Genome Sequence ? Useful information Human genes • Does it vary in humans? • Is it implicated in disease? Potential pathogens • Does it make a toxin? • Will UV sterilization work? Any organism • Does it synthesize ascorbic acid? • Will it grow at high temperatures?

LOCUS AB 060655 4091 bp DNA linear ROD 14 -SEP-2001 DEFINITION Mus musculus Atp 6 f gene for 23 -k. Da subunit of V-ATPase, complete cds. ACCESSION AB 060655 VERSION AB 060655. 1 GI: 14646762 KEYWORDS. SOURCE Mus musculus (house mouse) ORGANISM Mus musculus Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Glires; Rodentia; exon 1128. . 1176 Sciurognathi; Muroidea; Muridae; Murinae; Mus. /gene="Atp 6 f" REFERENCE 1 CDS join(529. . 595, 1128. . 1176, 1407. . 1490, 1621. . 1698, 1893. . 1962, AUTHORS Sun-Wada, G. H. , Murakami, H. , Nakai, H. , Wada, Y. and Futai, M. exon 1407. . 1490 2086. . 2137, 2472. . 2662, 3125. . 3151)/gene="Atp 6 f" TITLE Mouse Atp 6 f, the gene encoding the 23 -k. Da proteolipid of vacuolar proton translocating ATPase 1621. . 1698 /gene="Atp 6 f" exon JOURNAL Gene 274 (1 -2), 93 -99 (2001) /gene="Atp 6 f" /codon_start=1 exon 1893. . 1962 PUBMED 11675001 /gene="Atp 6 f" REFERENCE 2 (bases 1 to 4091) /product="23 -k. Da subunit of V-ATPase" exon 2086. . 2137 AUTHORS Wada, Y. , Sun-Wada, G. , Hideaki, M. and Masamitsu, F. /protein_id="BAB 61955. 1" /gene="Atp 6 f" TITLE Direct Submission exon 2472. . 2662 JOURNAL Submitted (23 -APR-2001) Yoh Wada, ISIR, Osaka University, Division /db_xref="GI: 14646763" /gene="Atp 6 f" of Biological Science; Mihogaoka 8 -1, Ibaraki, Osaka 5670047, Japan /translation="MTGLELLYLGIFVAFWACMVVVGICYTIFDLGFRFDVAWFLTET exon 3125. . 3435 (E-mail: yohwada@sanken. osaka-u. ac. jp, Tel: 81 -6 -6879 -8482, /gene="Atp 6 f" Fax: 81 -6 -6875 -5724) SPFMWSNLGIGLAISLSVVGAAWGIYITGSSIIGGGVKAPRIKTKNLVSIIFCEAVAI ORIGIN FEATURES Location/Qualifiers YGIIMAIVISNMAEPFSATEPKAIGHRNYHAGYSMFGAGLTVGLSNLFCGVCVGIVGS 1 gatcctatag ggcgaattgg agctccccgc ggtggcggcc gctctagaac tagtggatc source 1. . 4091 61 cctggacatc gtgggcgttc gcgtctggca ttccacccta cctctgggtt ggaaaagaca /organism="Mus musculus" GAALADAQNPSLFVKILIVEIFGSAIGLFGVIVAILQTSRVKMGD" 121 acctagaatg acctccgatg aacagcaggc attagctagg caccgcgaaa tcctgct /mol_type="genomic DNA" 181 agcagaagga actaggcagg actagaacag accggaagga tctgcagtga ttggt /strain="129 Sv" 241 aactgggagt ccggtgggaa gttagggaac cagcagcgca ggtggagagc cagta /db_xref="taxon: 10090" 301 cacggagaac gtccgacgaa actacaacca ccacagtgct ccgcggcatg acgtct /chromosome="4". . . /clone="225 b 09" 3901 ttacctaata agtccttttc agtcaacacc tttaggggtc ttacccagca ggcagccctg /clone_lib="Genome Systems" 3961 gttggctgac cttgactcat gctcccagga aagagttggc aaggccctaa ccctctga gene 483. . 3435 4021 tgcccactat ccagaccccg tcccaaatac ctgaagggcc ttagccatcc ggctcctg /gene="Atp 6 f" 4081 ctcttcccat t exon 483. . 595 // /gene="Atp 6 f" CDS join(529. . 595, 1128. . 1176, 1407. . 1490, 1621. . 1698, 1893. . 1962, 2086. . 2137, 2472. . 2662, 3125. . 3151) /gene="Atp 6 f" /codon_start=1 /product="23 -k. Da subunit of V-ATPase" /protein_id="BAB 61955. 1" /db_xref="GI: 14646763" /translation="MTGLELLYLGIFVAFWACMVVVGICYTIFDLGFRFDVAWFLTET SPFMWSNLGIGLAISLSVVGAAWGIYITGSSIIGGGVKAPRIKTKNLVSIIFCEAVAI YGIIMAIVISNMAEPFSATEPKAIGHRNYHAGYSMFGAGLTVGLSNLFCGVCVGIVGS GAALADAQNPSLFVKILIVEIFGSAIGLFGVIVAILQTSRVKMGD" Annotations
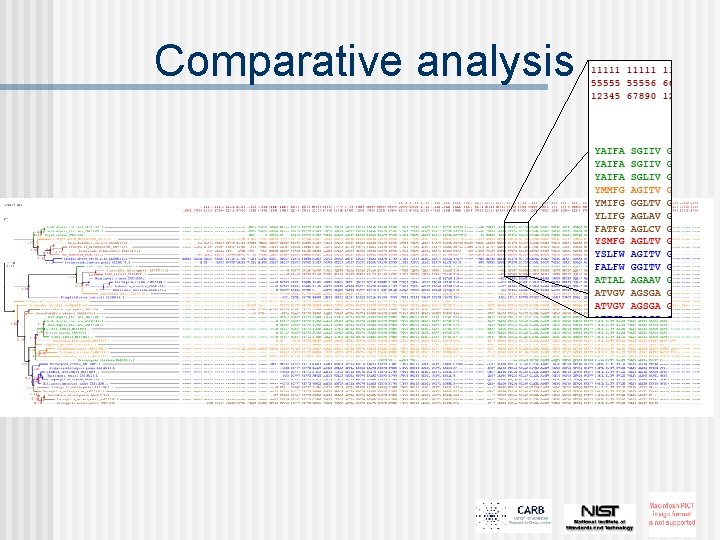
Comparative analysis

Example: SIFT
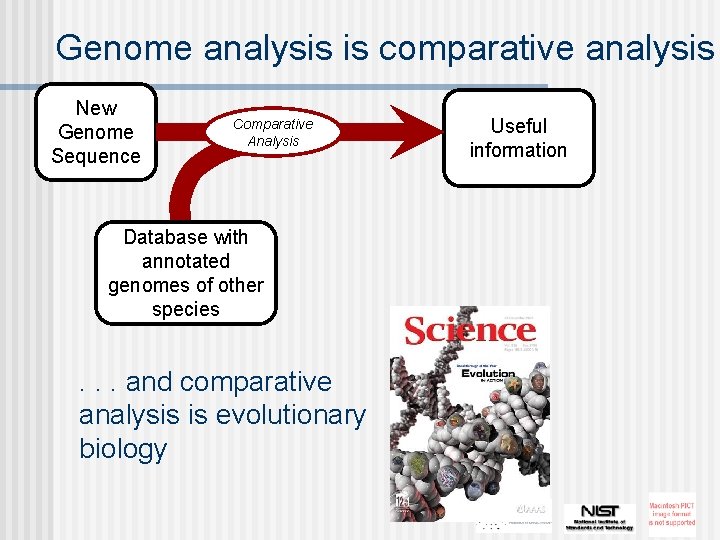
Genome analysis is comparative analysis New Genome Sequence ? Comparative Analysis Database with annotated genomes of other species . . . and comparative analysis is evolutionary biology Useful information

A bold generalization "It matters not at all whether you work with genetic elements, with viruses, bacteria, fungi, animals, or plants. The same principles apply if your subject is molecular evolution, the diversity of genetic systems, comparative morphology, physiology, ecology, or behaviour. " (p. 7) Harvey, P. H. , and M. D. Pagel. 1991. The Comparative Method in Evolutionary Biology. Oxford University Press, Oxford. What are these principles?
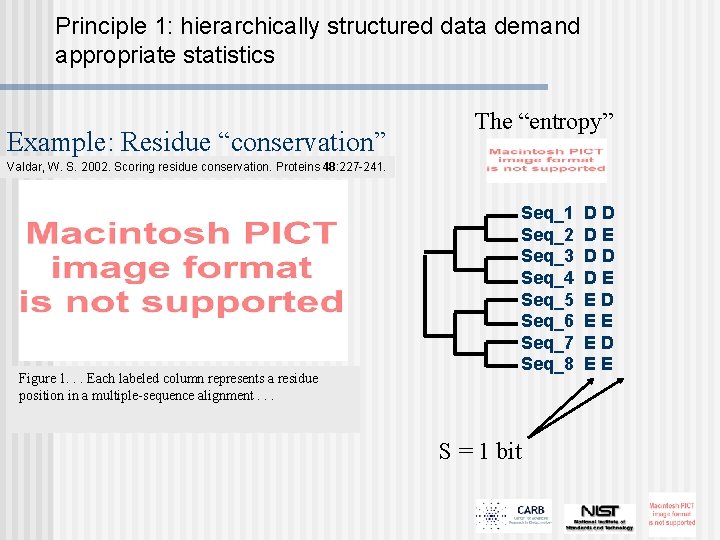
Principle 1: hierarchically structured data demand appropriate statistics Example: Residue “conservation” The “entropy” Valdar, W. S. 2002. Scoring residue conservation. Proteins 48: 227 -241. Figure 1. . . Each labeled column represents a residue position in a multiple-sequence alignment. . . Seq_1 Seq_2 Seq_3 Seq_4 Seq_5 Seq_6 Seq_7 Seq_8 S = 1 bit DD DE ED EE
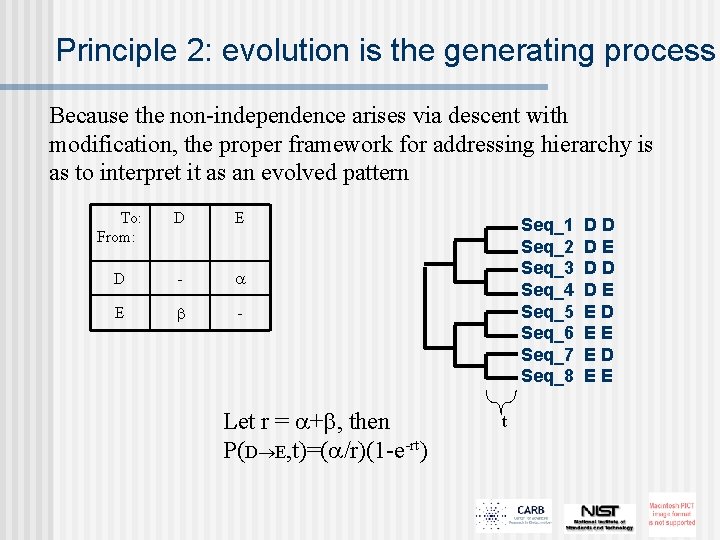
Principle 2: evolution is the generating process Because the non-independence arises via descent with modification, the proper framework for addressing hierarchy is as to interpret it as an evolved pattern To: From: D E D - E - Let r = + , then P(D E, t)=( /r)(1 -e-rt) Seq_1 Seq_2 Seq_3 Seq_4 Seq_5 Seq_6 Seq_7 Seq_8 t DD DE ED EE

Example: intron “loss vs. gain” problem gain intron A 1 B 1 C 0 D 0 E 0 F 0 max 0 Distance from root gain loss max 0 present loss max 0 Distance from root intron A 1 B 1 C 0 D 0 E 0 F 0 1 0 - 1 - Probabilities intron A 1 B 1 C 0 D 0 E 0 F 0 max 0 Distance from root Probability of presence Possibilities 0 AB 1 (Prob) 0 F 0 E CD Distance from root max

Example: functional inference presence A 1 B 1 C 0 D 0 E 0 F 0 t Let r = + , then P(0 1, t)=( /r)(1 -e-rt) functional attribute A 1 B 1 C ? D ? E 0 F 0

Principle 3: the result is an inference with uncertainty that should be treated explicitly • assign uncertainties to inferences • provide explicit probability distribution Example from Huelsenbeck “The phylogeny is usually treated as known without error; this assumption is problematic because inferred phylogenies are subject to both stochastic and systematic errors. ” Huelsenbeck, J. P. , B. Rannala, and J. P. Masly. 2000. Science 288: 2349 -2350. 0 1 0 - 1 -

Character-state data model Tree OTU: Operational Taxonomic Unit Character Data 13 Q Q E The “state” is Q (Glutamine) for “character” 13 (column 13) of “OTU” H_sapiens_4826964

#NEXUS [!Data and tree from: Schluter, D. 1989. Pp. 79 -95 in D. B. Wake and G. Roth, eds. , Complex organismal functions: Integration and evolution in vertebrates. Wiley, N. Y. ] BEGIN DATA; DIMENSIONS NTAX=14 NCHAR=5; FORMAT MISSING=? GAP=- ; CHARLABELS [1] Maxillary_tomia [2] lateral_groove [3] posterolateral_teeth [4] intercalary_ridge [5] maxillary_tomia; STATELABELS 1 thick thin, 2 deep shallow, 3 sharp reduced, 4 absent present, 5 'round-edged' 'sharp-edged'; MATRIX presumed_ancestor 00000 Geospiza_difficilis 00000 Geospiza_scandens 00000 Geospiza_conirostris 00000 Geospiza_magnirostris 00000 Geospiza_fortis 00000 Geospiza_fuliginosa 00000 Camarhynchus_pallidus 11101 Camarhynchus_heliobates 11101 Camarhynchus_psittacula 11101 Camarhynchus_pauper 11101 Camarhynchus_parvulus 11101 Platyspiza_crassirostris 11010 Certhidea_olivacea 11101; END; Character BEGIN DATA; MATRIXNTAX=14 NCHAR=5; DIMENSIONS State Data presumed_ancestor 00000 FORMAT MISSING=? GAP=- ; (example from Mac. Clade Geospiza_difficilis 00000 CHARLABELS documentation) Geospiza_scandens 00000 [1] Maxillary_tomia Geospiza_conirostris 00000 [2] lateral_groove Geospiza_magnirostris [3] posterolateral_teeth 00000 Geospiza_fortis [4] intercalary_ridge 00000 Geospiza_fuliginosa 00000 [5] maxillary_tomia; Camarhynchus_pallidus 11101 STATELABELS Camarhynchus_heliobates 11101 1 thick thin, Camarhynchus_psittacula 11101 2 deep shallow, Camarhynchus_pauper 11101 3 sharp reduced, BEGIN ASSUMPTIONS; Camarhynchus_parvulus 11101 OPTIONS DEFTYPE=unord Poly. Tcount=MINSTEPS ; 4 absent present, END; Platyspiza_crassirostris 11010 5 'round-edged' 'sharp-edged'; BEGIN TREES; TRANSLATE Certhidea_olivacea 11101; MATRIX 1 presumed_ancestor, 2 Geospiza_difficilis, 3 Geospiza_scandens, 4 Geospiza_conirostris, 5 Geospiza_magnirostris, 6 Geospiza_fortis, 7 Geospiza_fuliginosa, 8 Camarhynchus_pallidus, END; presumed_ancestor 00000 9 Camarhynchus_heliobates, 10 Camarhynchus_psittacula, 11 Camarhynchus_pauper, 12 Camarhynchus_parvulus, 13 Platyspiza_crassirostris, 14 Certhidea_olivacea; . . . TREE * UNTITLED = [&R] (1, (((2, (3, 4), ((5, 6), 7)), (((8, 9), ((10, 11), 12)), 13)), 14)); END;

The problem, restated Genome sequences Comparative analysis 99. 99 % accurate Useful inferences Far less accurate Power comes from comparative analysis Comparative analysis is an evolutionary problem § § § Depends on a tree describing relationships Depends on representing dynamics of evolution Requires attention to uncertainty • • • Facilitating tree-based analysis with better informatics Improving models of evolutionary change Incorporating prior knowledge How to improve evolutionary analysis?

How to advance evolutionary analysis? • Automate • Improve current models • Add more parameters • Expand universe of problem • Include more prior knowledge • Improve methods of numerical analysis • Demonstrate benefits of evolutionary analysis convincingly • Improve informatics support • standards (e. g. , NEXUS, Ne. XML) • libraries (Bio: : Phylo, Bio: : NEXUS) • applications (Mesquite, Nexplorer) • ontologies (CDAO) • intelligent user-oriented systems (e. g. , Galaxy)

Working group report outline n n n Development and evolution of goals Activities Products and other outcomes n n n n Ne. XML standard and implementations CDAO standard, publication Phylo. WS standard other Impacts Lessons learned Follow-ups

Informal meeting Activities Philly, June 2006 Evolutionary Informatics Working Group Phylohackathon 2007 Phylo. WS (Tokyo) WG 1 WG 2 2008 CDAO WG 3 DBH 1 2009 Ontology session at Evolution 2008 NESCent Phyloinformatics course Google Summer-of-Code projects Evol. Ontology RCN meeting at NESCent 2010 2011 Timespan of NIH project if funded • Comparative Data Analysis Ontology • Domain-specific language • Workflow construction using reasoning • Services infrastructure

Working group report outline n n n Development and evolution of goals Activities Products and other outcomes n n n n Ne. XML standard and implementations CDAO standard, publication Phylo. WS standard other Impacts Lessons learned Follow-ups

Prioritization exercise In spring of 2007, participants ranked 11 proposed items leaders devised coherent plan with suggested tactics

First meeting May 21 -23, 2007, NESCent n Priorities and activities n Supporting current file formats n Substitution model language n Central unifying artefact n New data exchange format n Outreach (funding, community needs) n

Tangible Outcomes, period 1 n New data exchange format (wiki) n n Current formats (wiki) n n Assessment Initial results on related technologies Central Unifying Artefact (wiki, docs, online demos) n n n n Use assessment (incomplete) Examples (incomplete) Transition Model Language (wiki) n n Detailed proposal Ne. XML draft Ontology development strategy (CDAO) Concept glossary Ontology-based semantic transformation demos Project proposal (4 -year, ~1. 2 M$ NIH RO 1) International team of collaborators Outreach: not much (broader awareness)

Tangible Outcomes, period 2 n New data exchange format (wiki, nexml. org) n n Current formats (wiki) n n NA Comparative data analysis ontology (wiki, docs, demos) n n n NA Transition Model Language (wiki) n n Ne. XML (Rutger’s talk) Analyzed related artefacts Expanded concept glossary Developed first draft of CDAO Started evaluation Outreach: not much (broader awareness)

Tangible Outcomes, period 3 n New data exchange format (Ne. XML) n n n Current formats (wiki) n n NA Comparative data analysis ontology (wiki, docs, demos) n n NA Transition Model Language (wiki) n n More support from apps developers Broader support in libraries Completed first evaluation cycle Manuscript written, submitted, accepted Started v. 2 with more terms; support for protocols Outreach: n Meetings and workshops

Working group report outline n n n Development and evolution of goals Activities Products and other outcomes n n n n Ne. XML standard and implementations CDAO standard, publication Phylo. WS standard other Impacts Lessons learned Follow-ups

CDAO: key concepts & relations character state data matrix Annotation: taxonomic_link has … part_of Annotation: has Alignment procedure… part_of character TU belongs_to character state datum represents_TU has is_transformation_of state transformation topology node rooted tree is_a tree has Annotation: Tree Procedure Model… has child has part_of directed child_node ancest or edge has descendant has parent node parent_node is_a state has left_stat transformation e is_a unrooted part_ofedge has tree has Annotation: Length… has node has left_node connects_to node has right_sta te state


Summary of outputs n Ne. XML Standard n APIs n Supporting apps and resources n n CDAO Standard, evaluation n publication n n Phylo. WS Standard n Demo implementations n

Working group report outline n n n Development and evolution of goals Activities Products and other outcomes n n n n Ne. XML standard and implementations CDAO standard, publication Phylo. WS standard other Impacts Lessons learned Follow-ups

Impacts Cohesion & awareness n Interactions, spin-offs, related projects n Penetration of standards n Use of implementations n

This week’s hackathon Takes place of 4 th wg meeting n Aims to rely on wg technologies n Opportunity to n assess useability and scope of wg artefacts n assess prospects for technology “push” n assess potential gains from interoperability n expose weakness in wg artefacts n expose further interop needs n

Tangible Outcomes, this week n Semantics — accessing content via an RDF triple store n n n Phylr — UI for phylo. WS access to combined data n n n Process, translate, load from nexml into triplet store Java API and SPARQL query interface Dendro. Py nexml; bio. SQL; http: //dbhack 1. nescent. org: 8080/SRW/search/treebase? Java API for nexml — lightweight IO in Java n n Extensively implemented DOM approach (31 classes, 28 interfaces, 7 test classes live demo of test classes on last day!

Tangible Outcomes, this week n Visualization — UI to overlay on a tree data repository n n n Taxonomic intelligence — access to data via taxo. info n n http: //iptol. iplantcollaborative. org/hackathon/ — live demo Integration with Morphbank image collection Tree. Base REST API — live demo of local implementation Rogue projects and other outcomes n n n Ne. XML test files (metadata representation and visualization) mx improvements (tree viewing, nexml IO) Tentative metadata standard for nexml (meta, RDFa)

Introduction Current status (2/4) Parsers and writers • Nexml parsers and writers: o mesquite, java, using xmlbeans o Bio: : Phylo, perl o py. Nexml, python o DAMBE, Visual Basic o stubs for c++ xmlbeans o plans for ruby? The problem Evo. Info interests This subproject Nexus issues Parsing Extensibility XML goodies Design Principles Re-use Patterns Inheritance References Implementation Approach ERD Inheritance Anatomy Characters Trees Current status Schema blocks Parsers & writers Experiments To do Resources

Lessons learned — successes Choosing participants n Preparation for meetings n Use of collaboration tools n Identifying targets n

Lessons learned — challenges Preparation for meetings n Dissemination n Evaluation n Identifying targets n

Possible follow-ups Another hackathon this summer n Google So. C projects n MIAPA project n Renew evoinfo working group n

Evolutionary Informatics: Supporting Interoperability in Evolutionary Analysis NESCent Evolutionary Informatics Working Group Jon Eisen (“phylogenomics”) Joe Felsenstein (PHYLIP) Mark Holder (GARLI) Sergei Kosakovsky Pond (Hy. Phy) Sudhir Kumar (MEGA) Paul Lewis (NCL) Aaron Mackey (Bio. Perl, GMOD) David Maddison (Mesquite) Wayne Maddison (Mesquite) Enrico Pontelli (CDAO) Andrew Rambaut (BEAST) Arlin Stoltzfus (Bio: : NEXUS) David Swofford (PAUP*) Rutger Vos (Bio: : Phylo) Fourth Xuhua Xia (DAMBE) meeting Christian Zmasek (ATV, RIO) NESCent staff Hilmar Lapp Todd Vision WG colleagues Brandon Chisham Brian Devries Gopal Gupta Peter E. Midford William Piel Francisco Prosdocimi Julie Thompson Derrick Zwickl DB Interop Hackathon Jim Balhoff Lucie Chan Dave Clements Karen Cranston Sam Donnelly Vladimir Gapeyev Karla Gendler Vivek Gopalan Roger Hyam Mark Jensen Greg Jordan Matt Kosnik Sheldon Mc. Kay Ryan Scherle Katja Schulz Katja Seltmann Jeet Sukumaran Matt Yoder
- Slides: 38