Evolutionary Genome Biology Gabor T Marth D Sc

Evolutionary Genome Biology Gabor T. Marth, D. Sc. Department of Biology, Boston College marth@bc. edu Medical Genomics Course – Debrecen, Hungary, May 2006

Lecture overview 1. Inter-species evolution and comparative genomics 2. Intra-species evolution, population genomics, and human origins

1. Inter-species evolution and comparative genomics Initial sequencing and comparative analysis of the mouse genome Mouse Genome Sequencing Consortium Nature 420, 520 -562. 2002
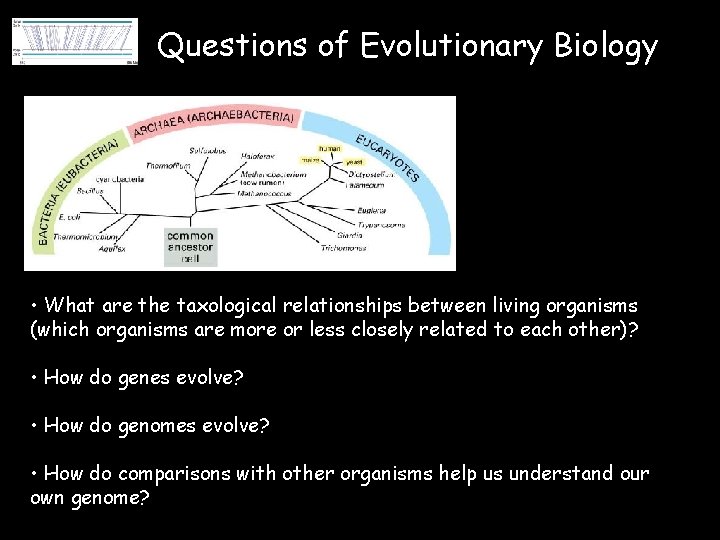
Questions of Evolutionary Biology • What are the taxological relationships between living organisms (which organisms are more or less closely related to each other)? • How do genes evolve? • How do genomes evolve? • How do comparisons with other organisms help us understand our own genome?
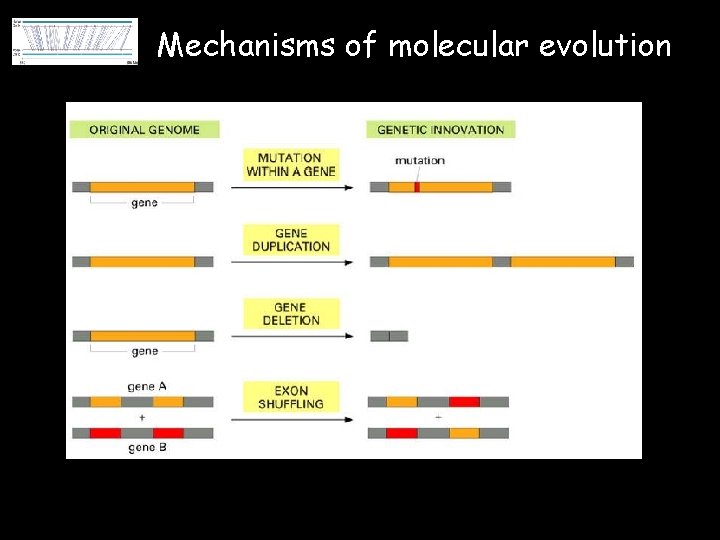
Mechanisms of molecular evolution
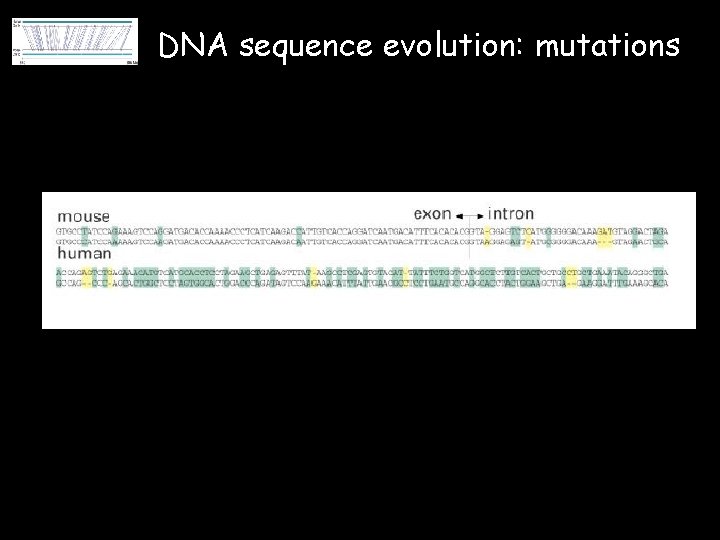
DNA sequence evolution: mutations

Phylogenetic relationships (1) Higgs and Attwood, Bioinformatics and Molecular Evolution, Blackwell Publishing Multiple alignment of mammalian mitochondrial small subunit r. RNA sequences
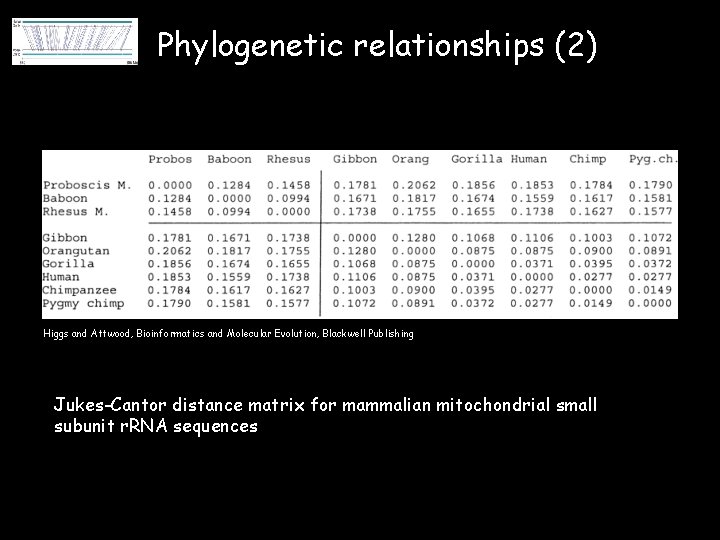
Phylogenetic relationships (2) Higgs and Attwood, Bioinformatics and Molecular Evolution, Blackwell Publishing Jukes-Cantor distance matrix for mammalian mitochondrial small subunit r. RNA sequences

Phylogenetic relationships (3) Higgs and Attwood, Bioinformatics and Molecular Evolution, Blackwell Publishing Phylogenetic tree constructed from mammalian mitochondrial small subunit r. RNA sequences
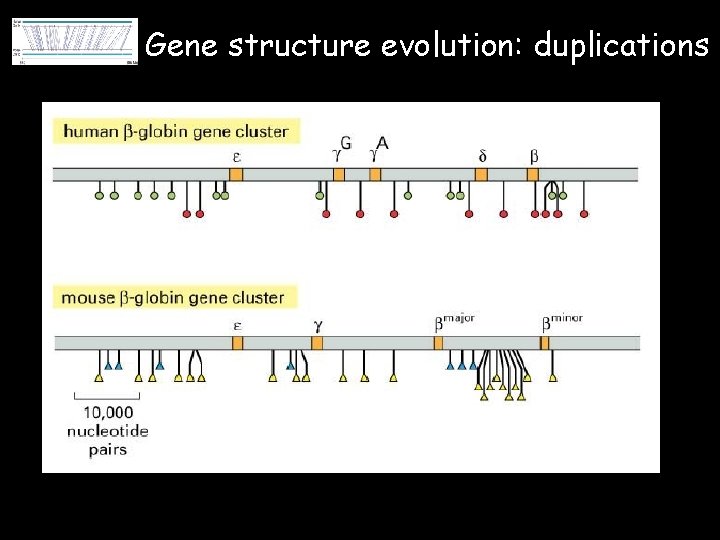
Gene structure evolution: duplications
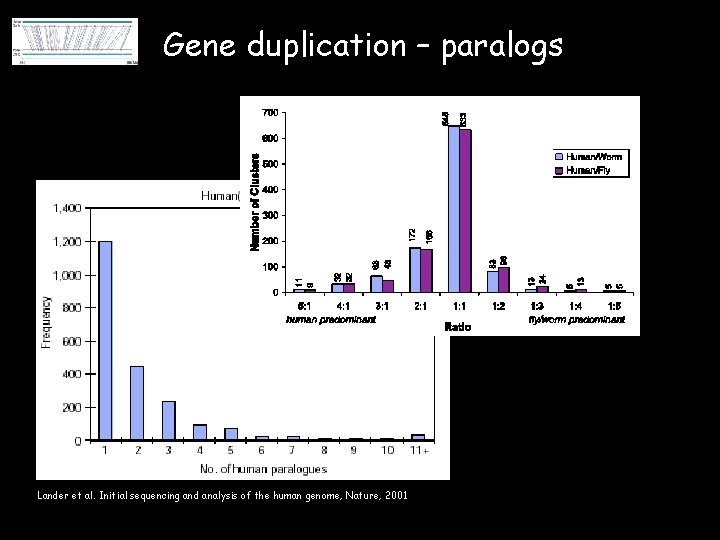
Gene duplication – paralogs Lander et al. Initial sequencing and analysis of the human genome, Nature, 2001
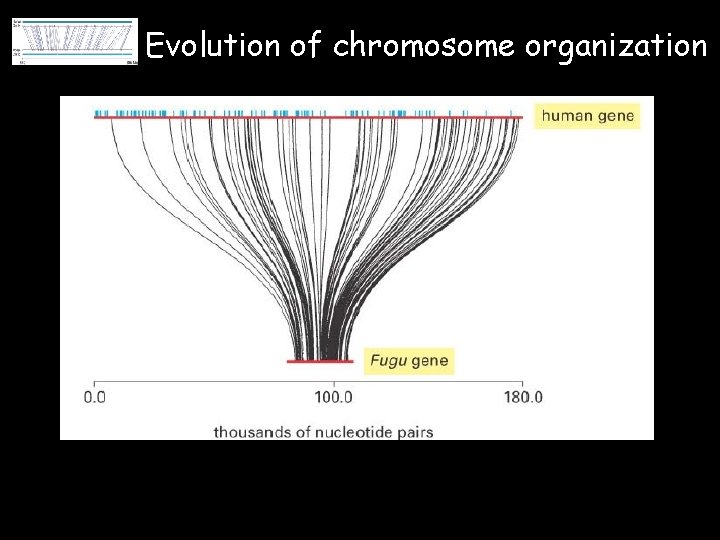
Evolution of chromosome organization
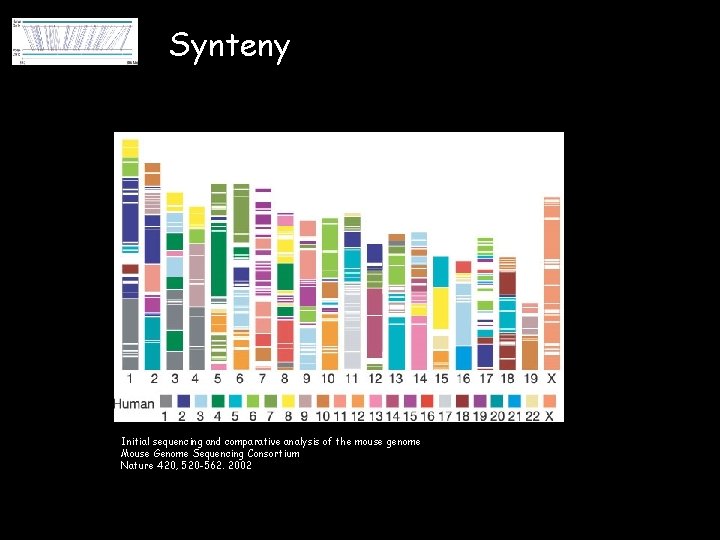
Synteny Initial sequencing and comparative analysis of the mouse genome Mouse Genome Sequencing Consortium Nature 420, 520 -562. 2002
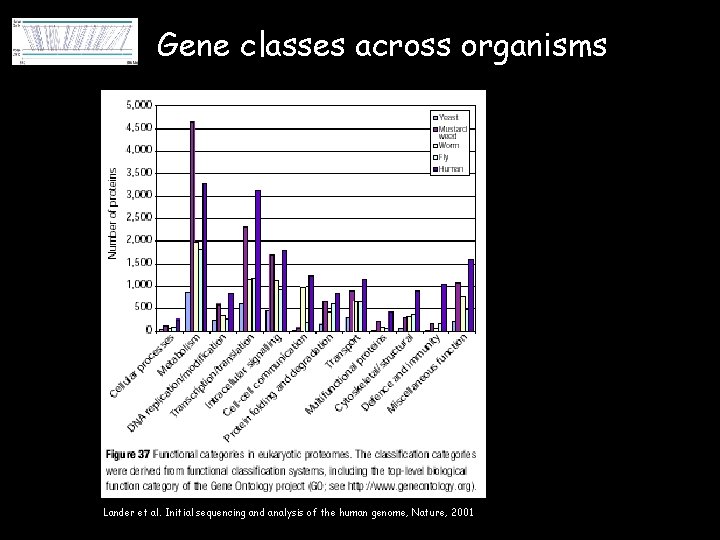
Gene classes across organisms Lander et al. Initial sequencing and analysis of the human genome, Nature, 2001
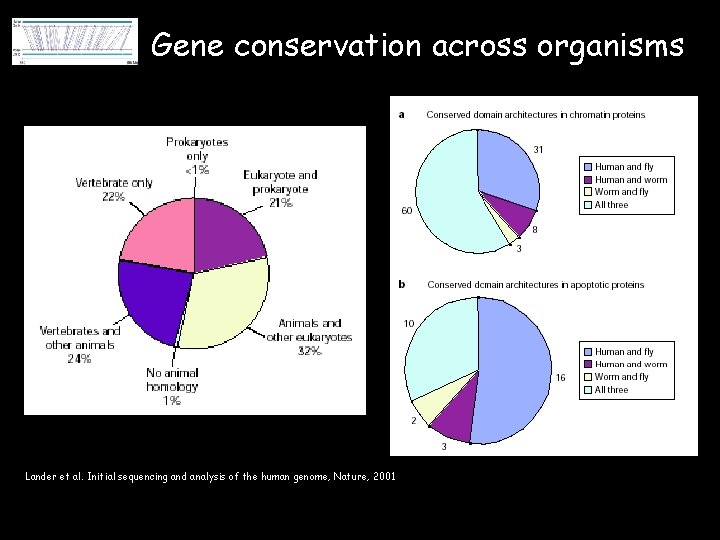
Gene conservation across organisms Lander et al. Initial sequencing and analysis of the human genome, Nature, 2001
Comparative genomics helps gene annotations

2. Intra-species evolution, population genomics, and human origins

Questions about human evolution • How do we discover / assess genetic variations? • What is the level of diversity across humans? • How can we model the ancestral and mutation processes? • What do phylogenetic analyses of human mitochondrial sequences tell us about human origins and dispersal? • Does mitochondrial DNA give us the full picture? • What do we learn from model-fitting analysis of nuclear DNA? • A single wave of out-of-Africa migration or multiple waves?
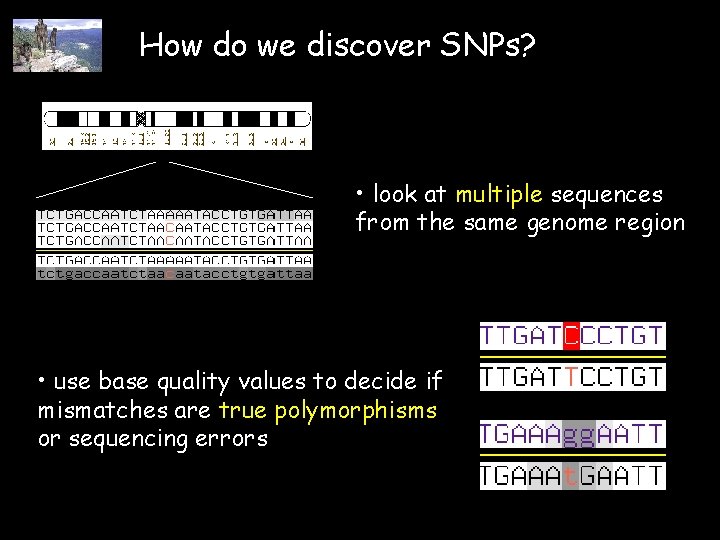
How do we discover SNPs? • look at multiple sequences from the same genome region • use base quality values to decide if mismatches are true polymorphisms or sequencing errors
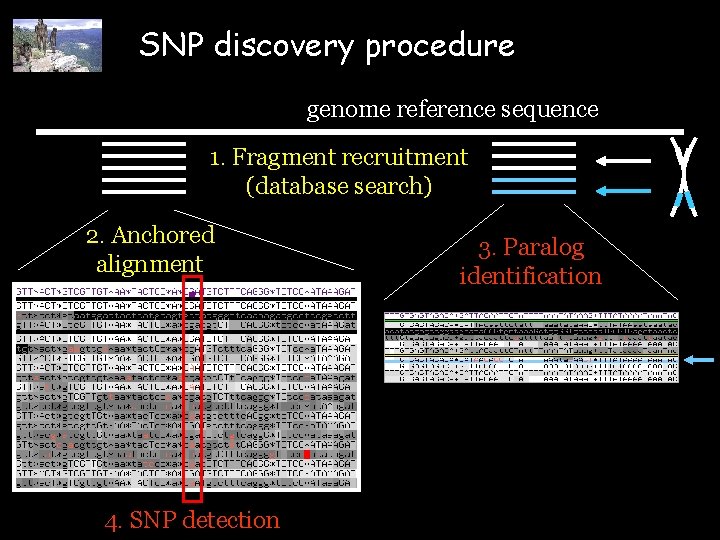
SNP discovery procedure genome reference sequence 1. Fragment recruitment (database search) 2. Anchored alignment 4. SNP detection 3. Paralog identification
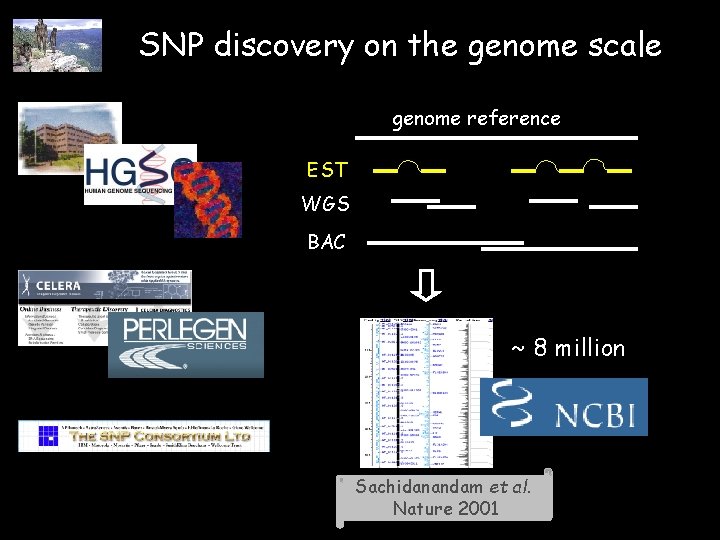
SNP discovery on the genome scale genome reference EST WGS BAC ~ 8 million Sachidanandam et al. Nature 2001
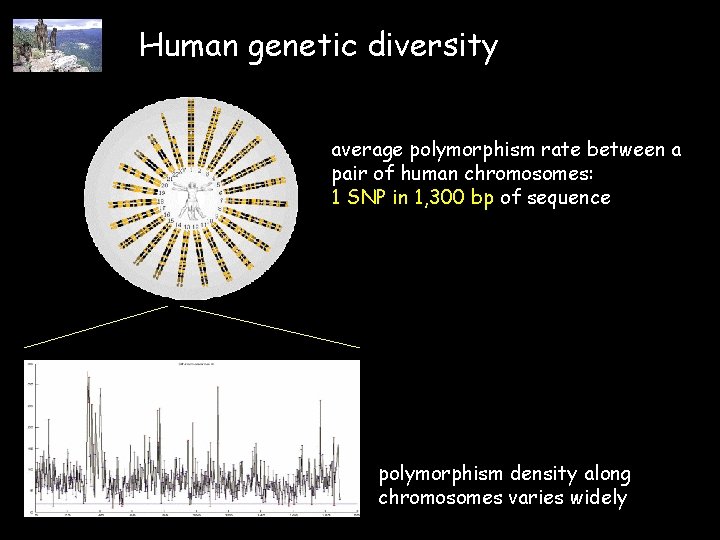
Human genetic diversity average polymorphism rate between a pair of human chromosomes: 1 SNP in 1, 300 bp of sequence polymorphism density along chromosomes varies widely
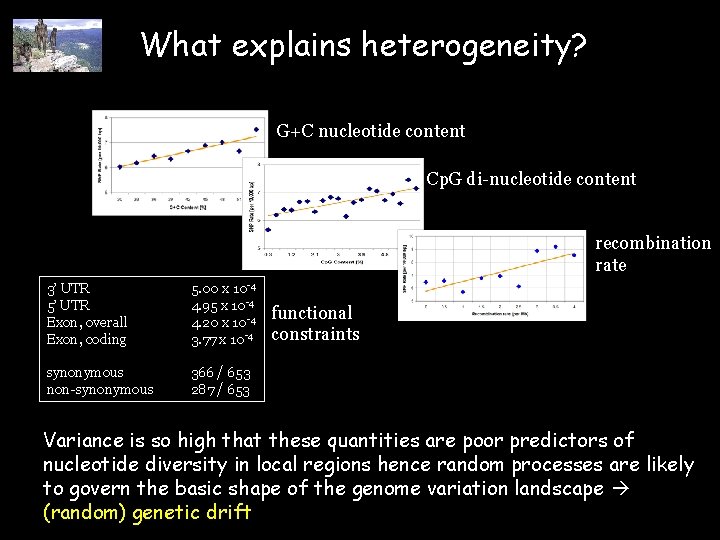
What explains heterogeneity? G+C nucleotide content Cp. G di-nucleotide content recombination rate 3’ UTR 5’ UTR Exon, overall Exon, coding 5. 00 x 10 -4 4. 95 x 10 -4 4. 20 x 10 -4 3. 77 x 10 -4 synonymous non-synonymous 366 / 653 287 / 653 functional constraints Variance is so high that these quantities are poor predictors of nucleotide diversity in local regions hence random processes are likely to govern the basic shape of the genome variation landscape (random) genetic drift
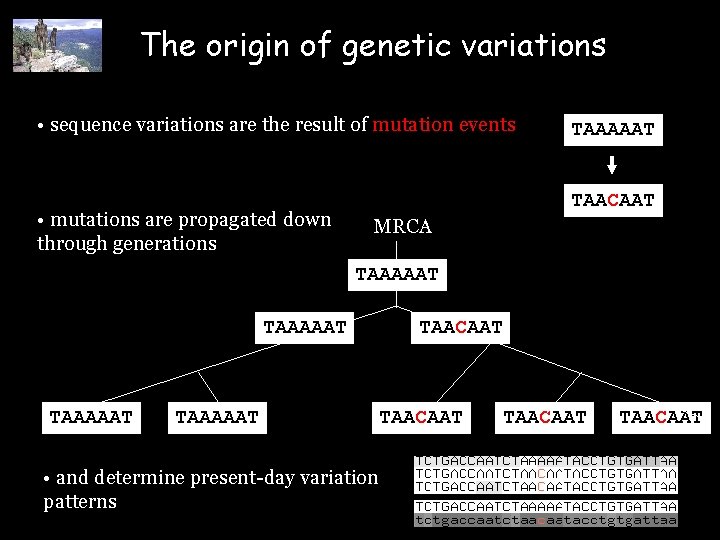
The origin of genetic variations • sequence variations are the result of mutation events • mutations are propagated down through generations TAAAAAT TAACAAT MRCA TAAAAAT • and determine present-day variation patterns TAACAAT

Recombination messes up phylogenies accgttatgtaga acggttatgtaga acggttatgtaga accgttatgtaga • because of recombination, DNA sequences may not have a unique common ancestor, hence phylogenetic analysis may not apply
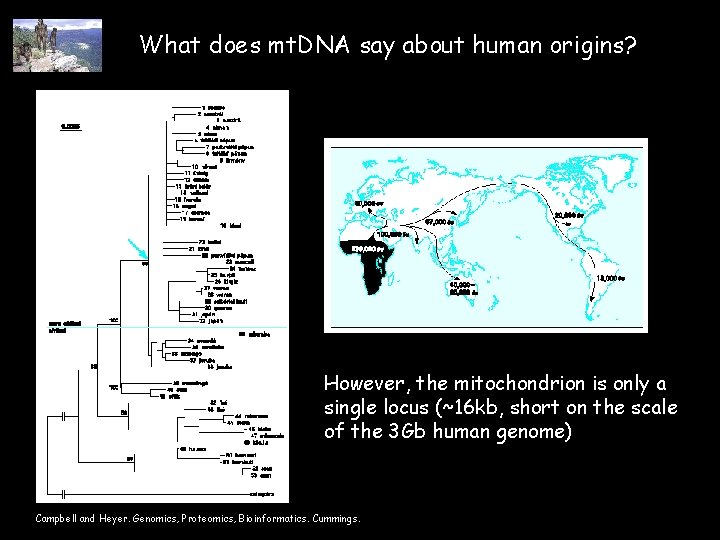
What does mt. DNA say about human origins? However, the mitochondrion is only a single locus (~16 kb, short on the scale of the 3 Gb human genome) Campbell and Heyer. Genomics, Proteomics, Bioinformatics. Cummings.
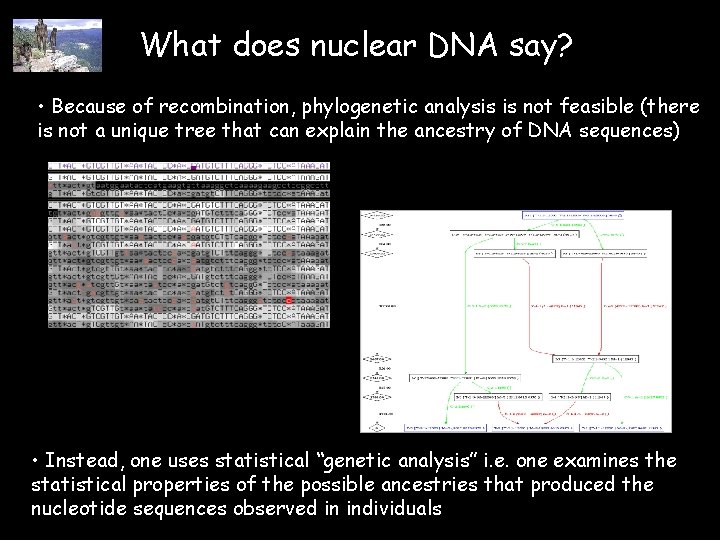
What does nuclear DNA say? • Because of recombination, phylogenetic analysis is not feasible (there is not a unique tree that can explain the ancestry of DNA sequences) • Instead, one uses statistical “genetic analysis” i. e. one examines the statistical properties of the possible ancestries that produced the nucleotide sequences observed in individuals
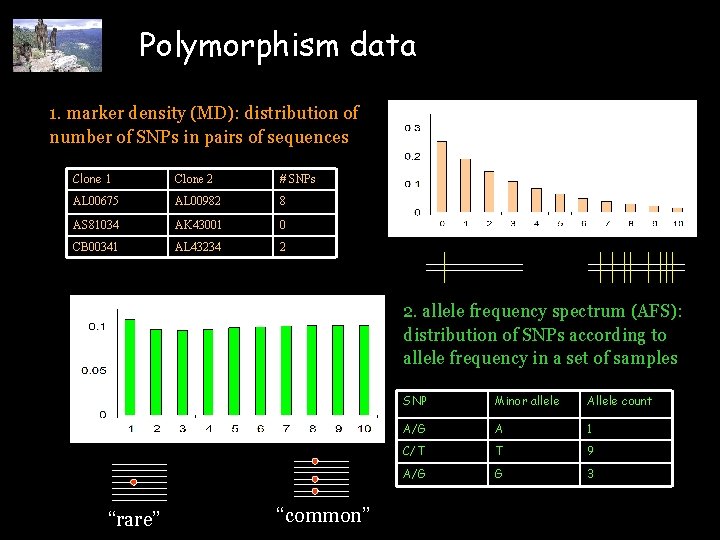
Polymorphism data 1. marker density (MD): distribution of number of SNPs in pairs of sequences Clone 1 Clone 2 # SNPs AL 00675 AL 00982 8 AS 81034 AK 43001 0 CB 00341 AL 43234 2 2. allele frequency spectrum (AFS): distribution of SNPs according to allele frequency in a set of samples “rare” “common” SNP Minor allele Allele count A/G A 1 C/T T 9 A/G G 3
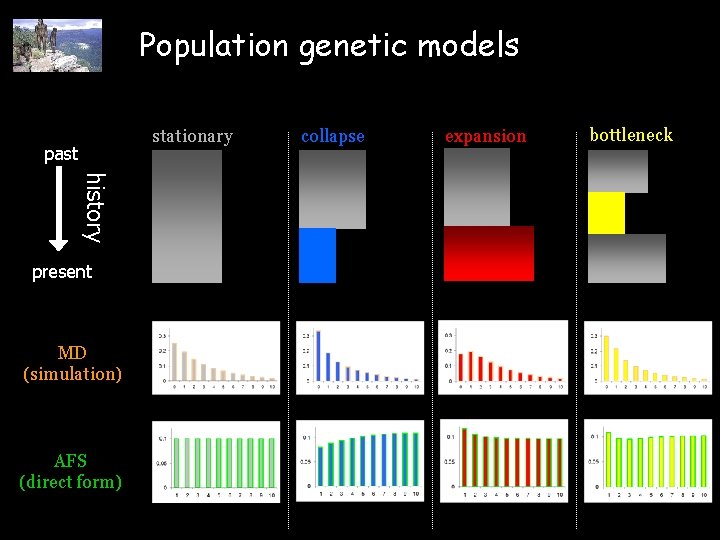
Population genetic models stationary past history present MD (simulation) AFS (direct form) collapse expansion bottleneck
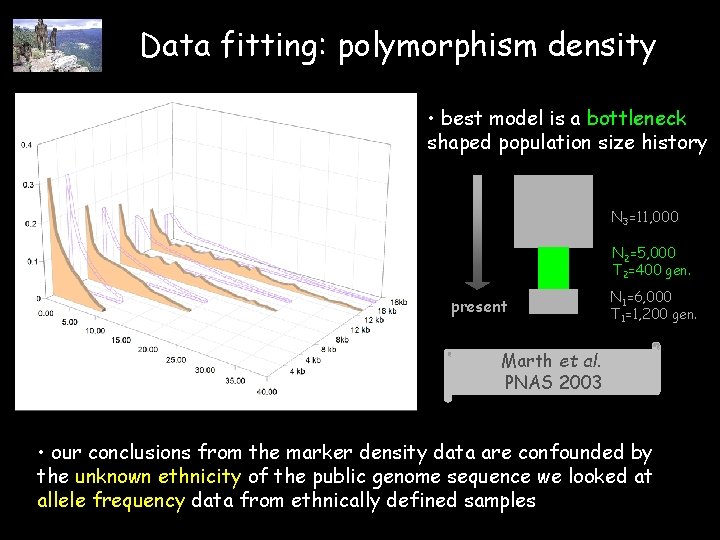
Data fitting: polymorphism density • best model is a bottleneck shaped population size history N 3=11, 000 N 2=5, 000 T 2=400 gen. present N 1=6, 000 T 1=1, 200 gen. Marth et al. PNAS 2003 • our conclusions from the marker density data are confounded by the unknown ethnicity of the public genome sequence we looked at allele frequency data from ethnically defined samples
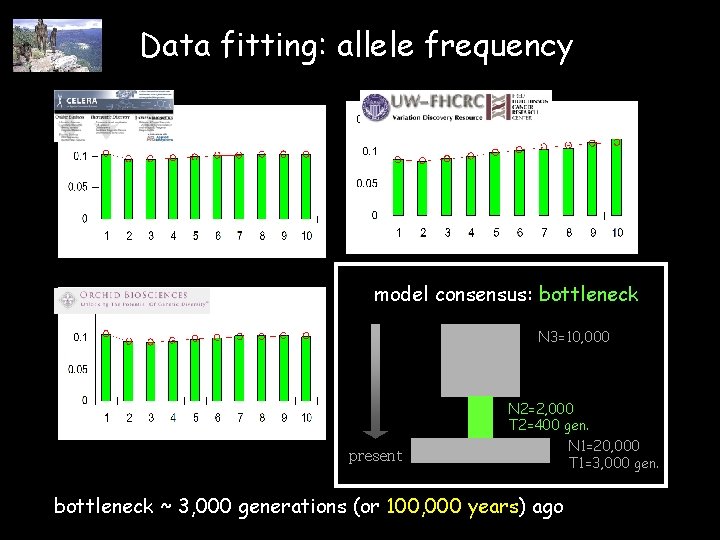
Data fitting: allele frequency model consensus: bottleneck N 3=10, 000 present N 2=2, 000 T 2=400 gen. N 1=20, 000 T 1=3, 000 gen. bottleneck ~ 3, 000 generations (or 100, 000 years) ago
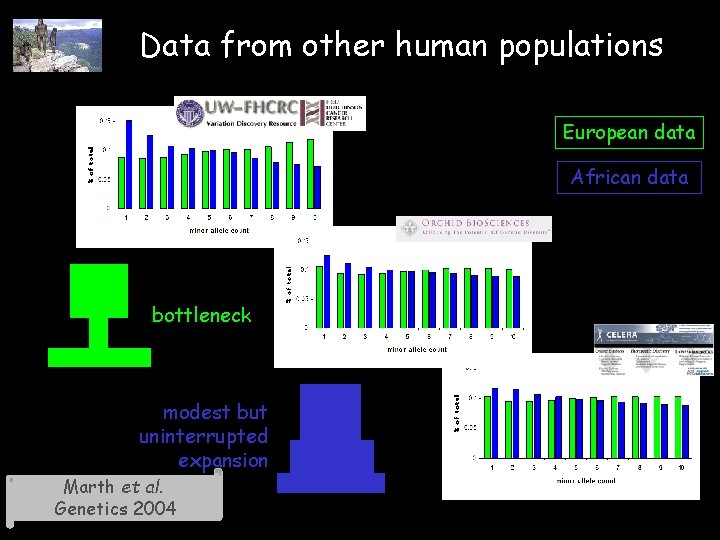
Data from other human populations European data African data bottleneck modest but uninterrupted expansion Marth et al. Genetics 2004
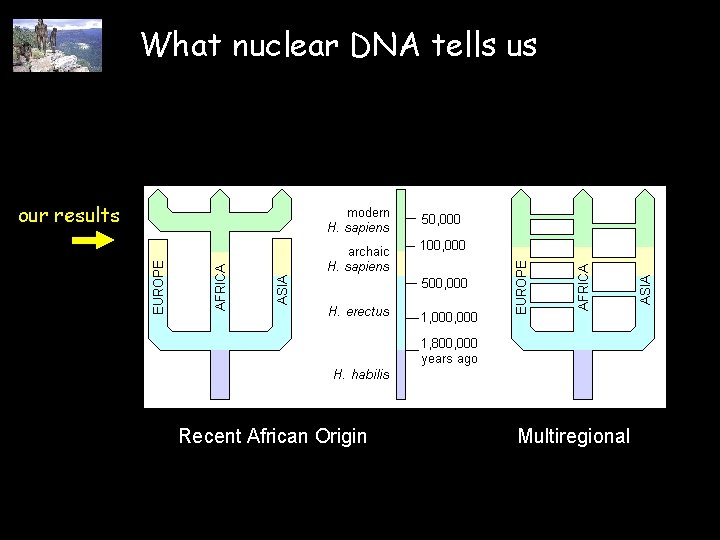
What nuclear DNA tells us our results Recent African Origin Multiregional
- Slides: 33