Evolution of toxinantitoxin systems in Streptococcus bacteria Sarah

Evolution of toxin-antitoxin systems in Streptococcus bacteria Sarah Cook Carnegie Mellon University 03 -727 Phylogenetics 12 December 2017

Toxin-antitoxin (TA) system as abortive infection mechanism • Abortive infection (Abi) mechanism • Bacterial innate immunity • Phage infection resistance • Result in altruistic suicide of infected cell reduce phage progeny release • Discovering Abis that function as TA systems • Plasmid encoded • Antitoxin gene located upstream of toxin gene • Antitoxin necessary for cell survival

Motivation for Phylogenetic Analysis • Dy et al. found type IV TA system in Streptococcus agalactiae • Abi. Ei (antitoxin) – transcriptional regulator • Abi. Eii (toxin) – nucleotidyl transferase • Antic et al. studied strain differences in Streptococcus pneumoniae • Observed two strains with 2 separate toxin genes

Project Aims • Do other Streptococcus bacteria have 2 different toxin (T) genes? • Is there a similar pattern in the antitoxin (A) genes? • Are the phylogenetic distributions of the T and A genes similar? • How does the gene tree compare with the species tree? • Are some species excluded? • Are some groups excluded? • What evolutionary history best explains the distribution of TA genes?

Project Aims • Do other Streptococcus bacteria have 2 different toxin (T) genes? • Is there a similar pattern in the antitoxin (A) genes? • Are the phylogenetic distributions of the T and A genes similar? • How does the gene tree compare with the species tree? • Are some species excluded? • Are some groups excluded? • What evolutionary history best explains the distribution of TA genes?

Amino Acid Sequence Acquisition • To obtain toxin sequences: • BLAST with S. agal sequence from Dy paper • Ref_seq and nr gave essentially the same sequence hits • Find a significant increase in E-value use the “drop off” sequence as query in a second search

Toxin Sequence Alignment • Aligned using Jalview • Decided to use Mafft with accuracy-oriented preset • Trees were the same • Fewer gaps • Better alignment of conserved domains

Two distinct groups in toxin alignment

Project Aims • Do other Streptococcus bacteria have 2 different toxin (T) genes? • Is there a similar pattern in the antitoxin (A) genes? • Are the phylogenetic distributions of the T and A genes similar? • How does the gene tree compare with the species tree? • Are some species excluded? • Are some groups excluded? • What evolutionary history best explains the distribution of TA genes?

Antitoxin Sequence Acquisition and Alignment • BLAST using B 1599 S. pneumoniae “A 1” query sequence • Used ref_seq • Excluded S. pneumoniae • Mafft (accuracy-oriented)
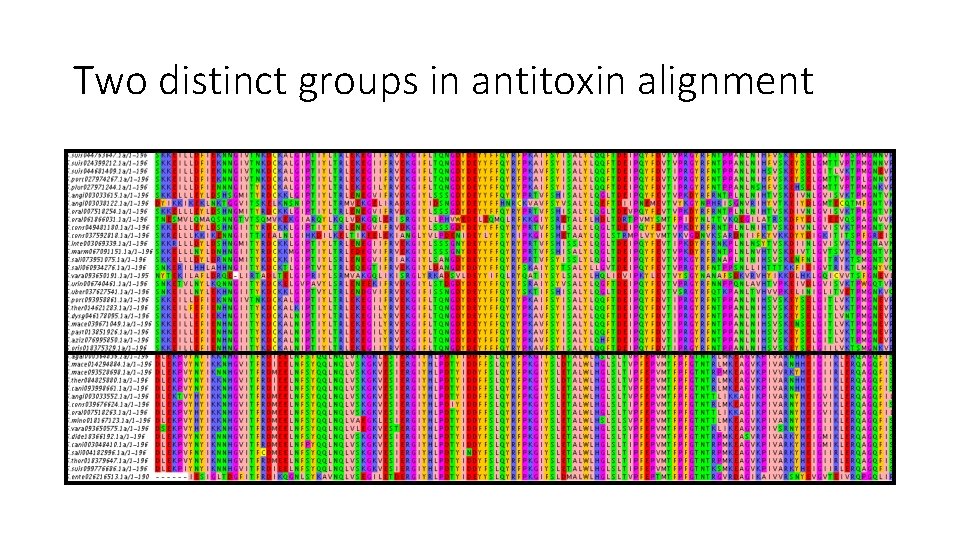
Two distinct groups in antitoxin alignment

Project Aims • Do other Streptococcus bacteria have 2 different toxin (T) genes? • Is there a similar pattern in the antitoxin (A) genes? • Are the phylogenetic distributions of the T and A genes similar? • How does the gene tree compare with the species tree? • Are some species excluded? • Are some groups excluded? • What evolutionary history best explains the distribution of TA genes?

Toxin model selection using modelgenerator http: //mcinerneylab. com/software/modelgenerator/
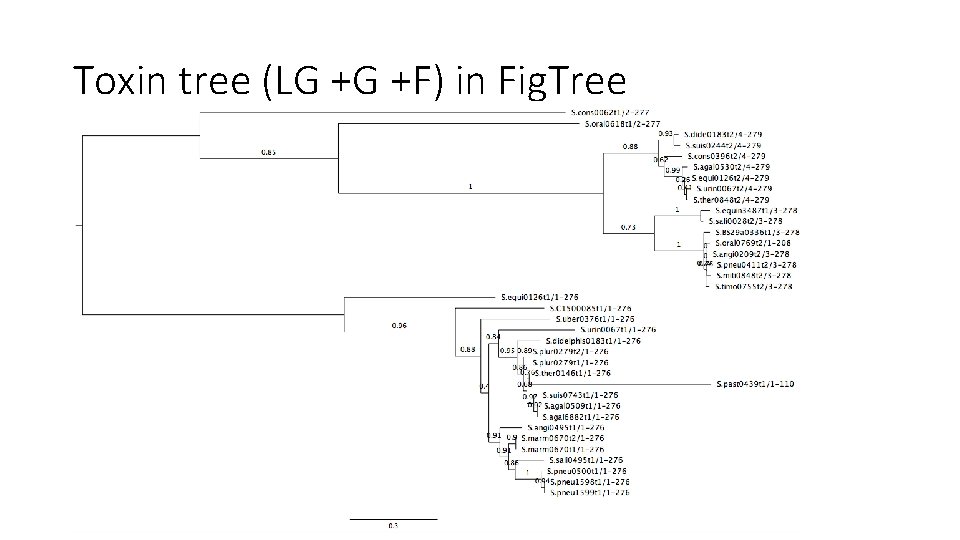
Toxin tree (LG +G +F) in Fig. Tree

Antitoxin model selection using model generator
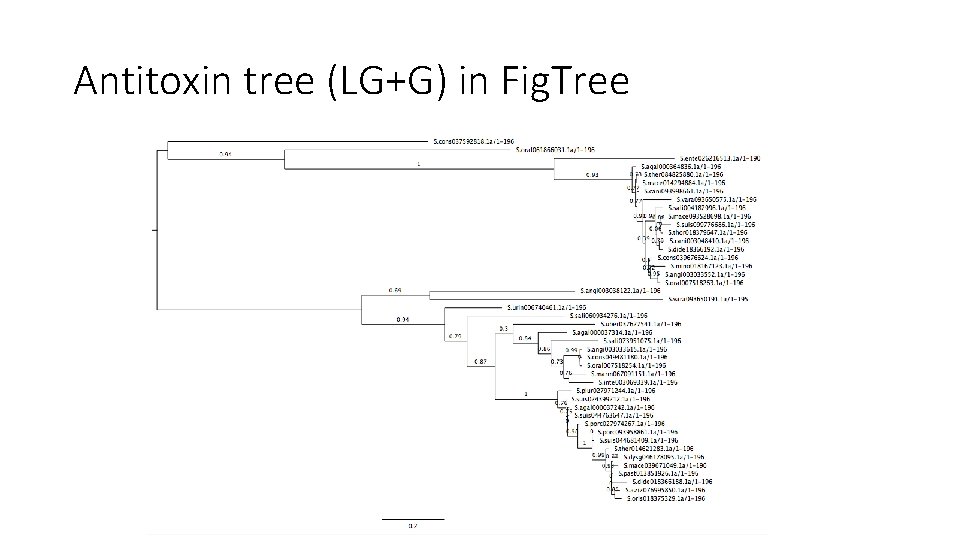
Antitoxin tree (LG+G) in Fig. Tree

Project Aims • Do other Streptococcus bacteria have 2 different toxin (T) genes? • Is there a similar pattern in the antitoxin (A) genes? • Are the phylogenetic distributions of the T and A genes similar? • How does the gene tree compare with the species tree? • Are some species excluded? • Are some groups excluded? • What evolutionary history best explains the distribution of TA genes?
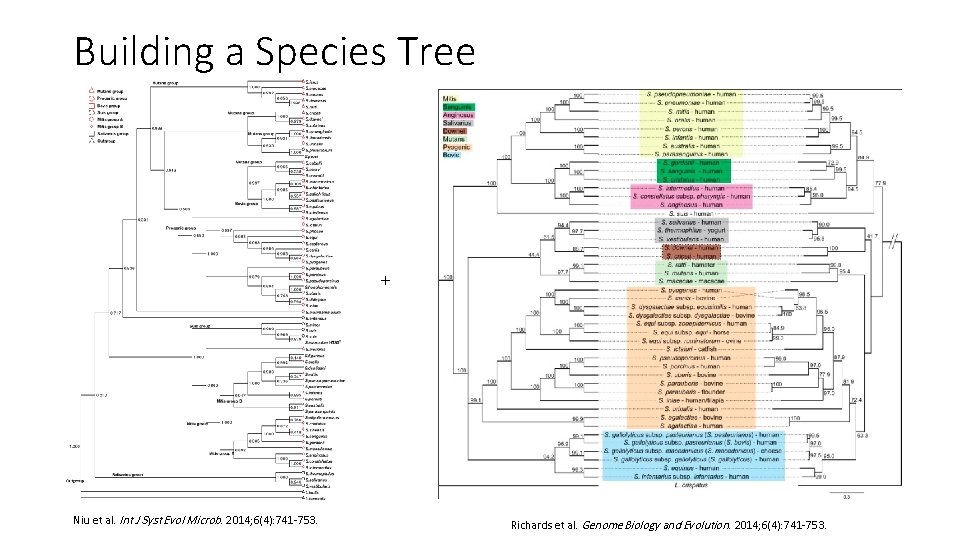
Building a Species Tree + Niu et al. Int J Syst Evol Microb. 2014; 6(4): 741 -753. Richards et al. Genome Biology and Evolution. 2014; 6(4): 741 -753.
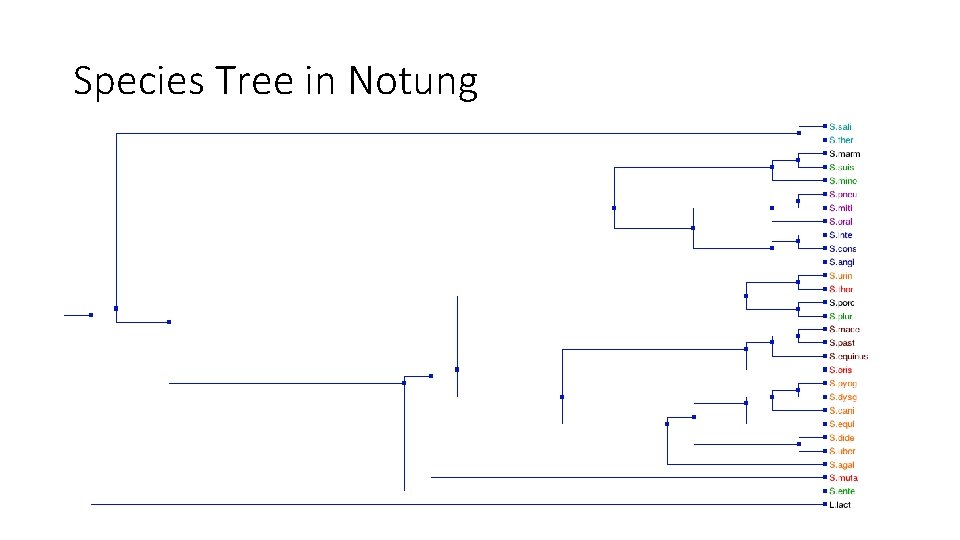
Species Tree in Notung
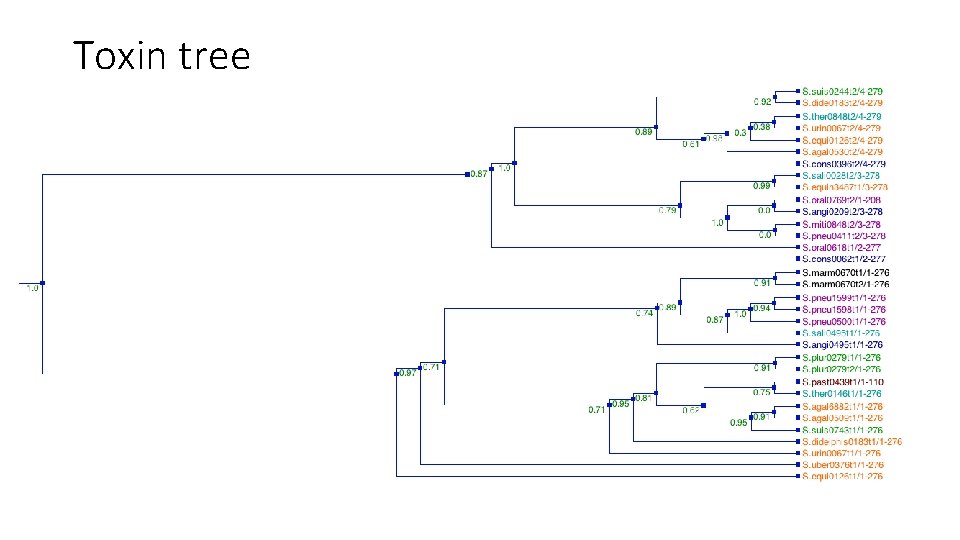
Toxin tree
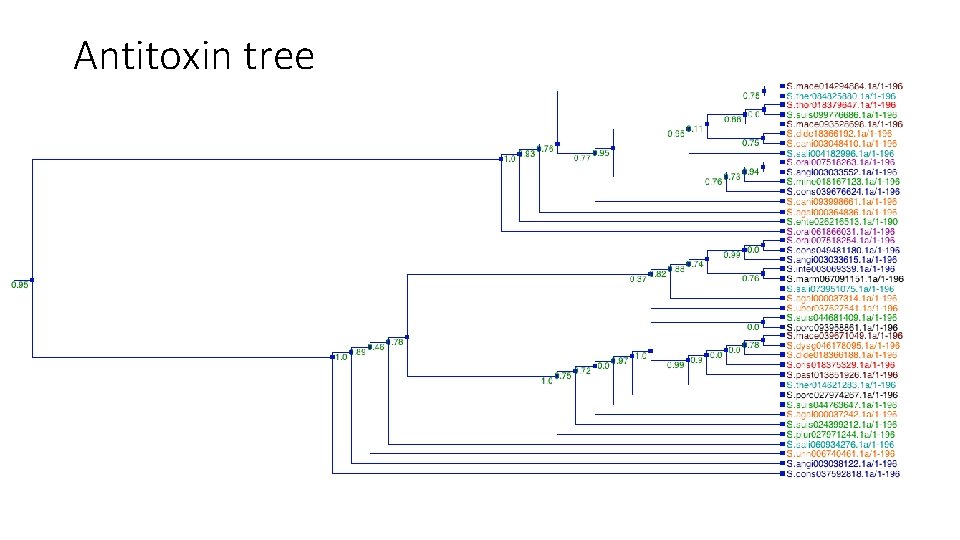
Antitoxin tree
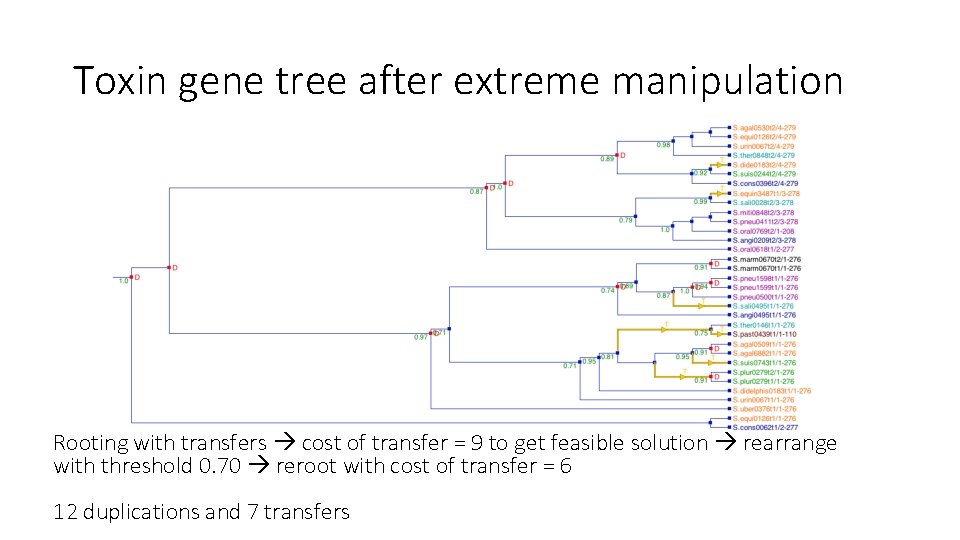
Toxin gene tree after extreme manipulation Rooting with transfers cost of transfer = 9 to get feasible solution rearrange with threshold 0. 70 reroot with cost of transfer = 6 12 duplications and 7 transfers
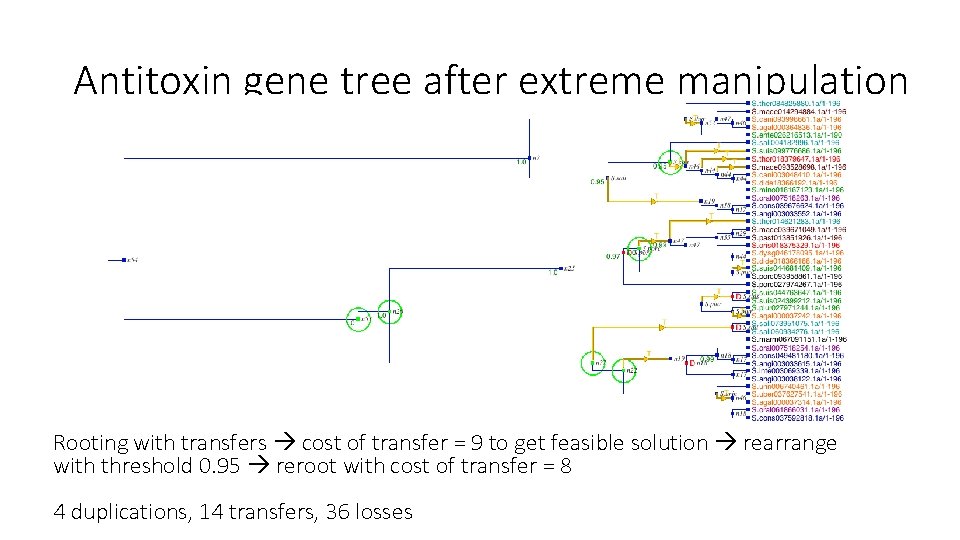
Antitoxin gene tree after extreme manipulation Rooting with transfers cost of transfer = 9 to get feasible solution rearrange with threshold 0. 95 reroot with cost of transfer = 8 4 duplications, 14 transfers, 36 losses

Future Directions • Different toxin sequence acquisition method • Include B 1599/1598 sequences in alignment • Use tblastn • Include S. pneu in analysis • Find T 1/A 1, T 2/A 2 from B 1599 S. pneu • Are they always located next to each other? Constraint on gene tree

References Richards VP, Palmer SR, Pavinski Bitar PD, et al. Phylogenomics and the Dynamic Genome Evolution of the Genus Streptococcus. Genome Biology and Evolution. 2014; 6(4): 741 -753. doi: 10. 1093/gbe/evu 048. Dy RL, Przybilski R, Semeijn K, Salmond GPC, Fineran PC. A widespread bacteriophage abortive infection system functions through a Type IV toxin–antitoxin mechanism. Nucleic Acids Research. 2014; 42(7): 4590 -4605. doi: 10. 1093/nar/gkt 1419. Antic I, Brothers KM, Stolzer M, et al. Gene Acquisition by a Distinct Phyletic Group within Streptococcus pneumoniae Promotes Adhesion to the Ocular Epithelium. Limbago BM, ed. m. Sphere. 2017; 2(5): e 00213 -17. doi: 10. 1128/m. Sphere. 00213 -17.

Supplementary Information
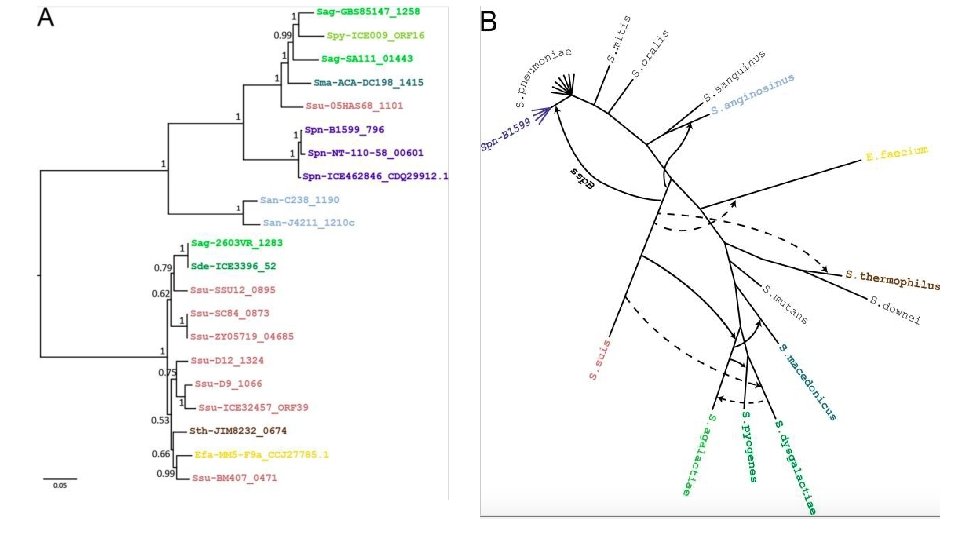
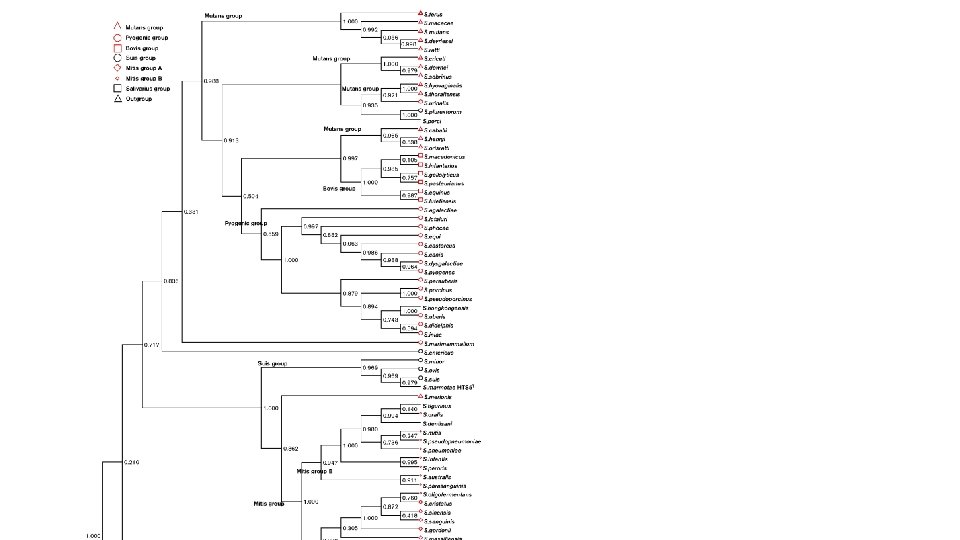
- Slides: 28