Establishing a Clinical Genomics Service at an Academic

Establishing a Clinical Genomics Service at an Academic Medical Center Jason Merker, MD, Ph. D Co-Director, Stanford Medicine Clinical Genomics Service Regional APC Meeting, Maui October 25 -28, 2016

Outline I. III. IV. Background on clinical genomics Workflow and case example Four major challenges Update and future directions

Background Clinical Genomics

Wellcome Collection – Medicine Now 3. 4 billion units of DNA code: • 127 volumes • 1, 000 pages per volume

Size of targeted regions in assays Genome – 127 volumes Exome – 2 volumes 400 gene heritable CA panel – 90 pages CF hotspot panel – < ½ page Single gene – 1 -2 lines

Amount of data generated for a single exome at 150 X = 3 bookcases Genome Exome – 300 volumes 400 gene heritable CA panel CF hotspot panel Single gene

Amount of data generated for trio genome analysis ≅ 135 stacked bookcases Honokohau Falls, West Maui 1, 119 ft (341 m) = ~100 bookcases
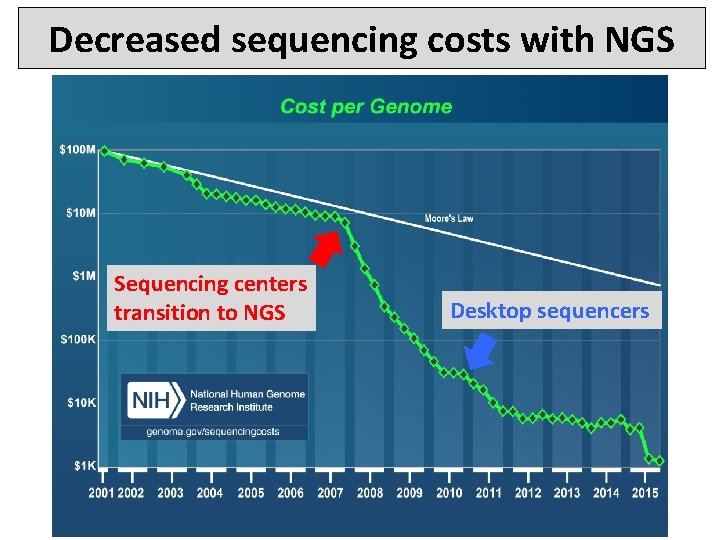
Decreased sequencing costs with NGS Sequencing centers transition to NGS Desktop sequencers
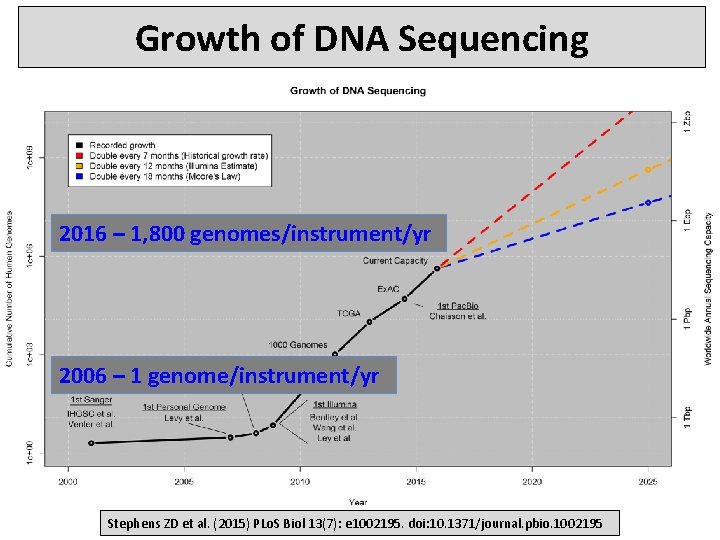
Growth of DNA Sequencing 2016 – 1, 800 genomes/instrument/yr 2006 – 1 genome/instrument/yr Stephens ZD et al. (2015) PLo. S Biol 13(7): e 1002195. doi: 10. 1371/journal. pbio. 1002195

Workflow and case example Stanford Medicine Clinical Genomics Service

Stanford Medicine Clinical Genomics Service • Pilot phase started in January 2014 • Use genome and exome sequencing to determine the cause of disease in patients with suspected genetic disorders • Focused on three major disease areas: – Pediatric and adult syndromes – Heritable cancer predisposition – Heritable cardiovascular disease
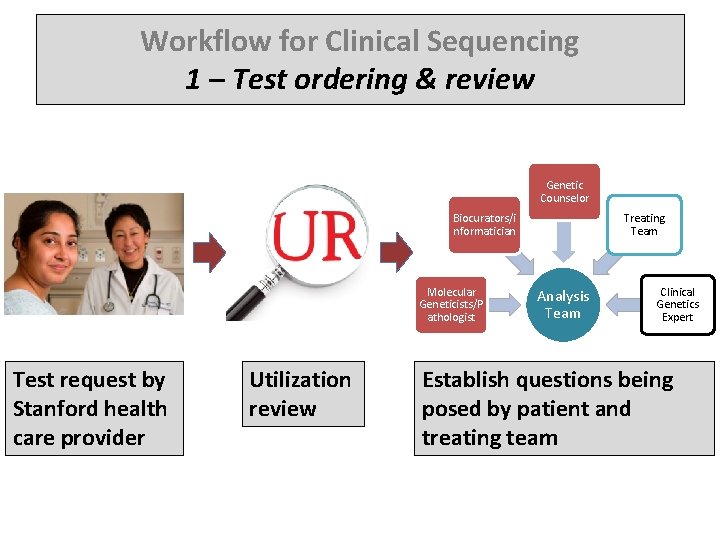
Workflow for Clinical Sequencing 1 – Test ordering & review Genetic Counselor Biocurators/i nformatician Molecular Geneticists/P athologist Test request by Stanford health care provider Utilization review Treating Team Analysis Team Clinical Genetics Expert Establish questions being posed by patient and treating team
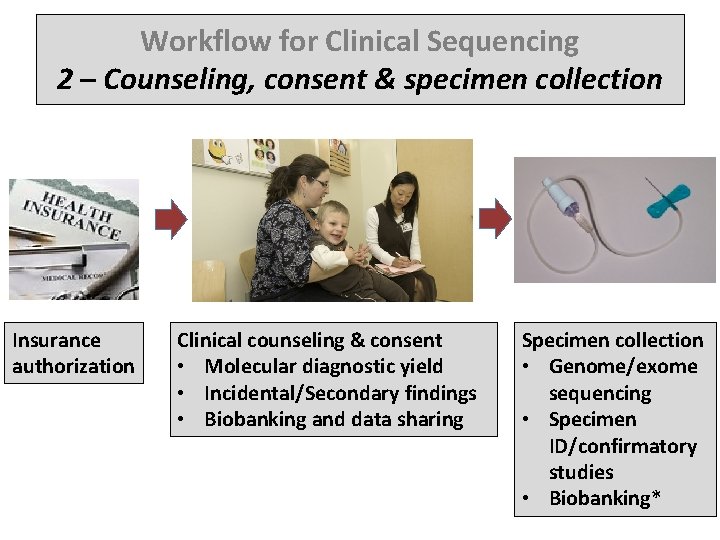
Workflow for Clinical Sequencing 2 – Counseling, consent & specimen collection Insurance authorization Clinical counseling & consent • Molecular diagnostic yield • Incidental/Secondary findings • Biobanking and data sharing Specimen collection • Genome/exome sequencing • Specimen ID/confirmatory studies • Biobanking*
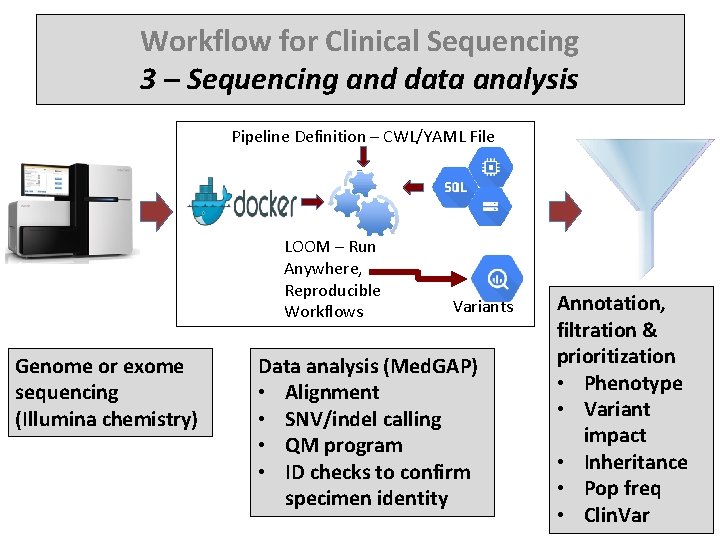
Workflow for Clinical Sequencing 3 – Sequencing and data analysis Pipeline Definition – CWL/YAML File LOOM – Run Anywhere, Reproducible Workflows Genome or exome sequencing (Illumina chemistry) Variants Data analysis (Med. GAP) • Alignment • SNV/indel calling • QM program • ID checks to confirm specimen identity Annotation, filtration & prioritization • Phenotype • Variant impact • Inheritance • Pop freq • Clin. Var
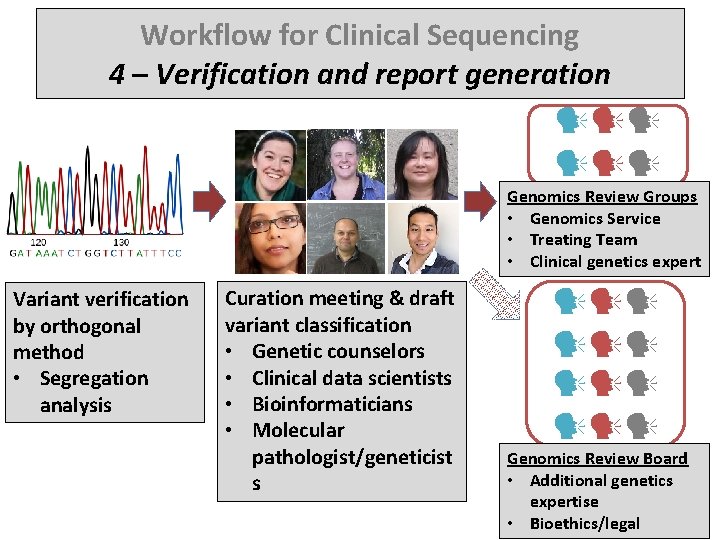
Workflow for Clinical Sequencing 4 – Verification and report generation Genomics Review Groups • Genomics Service • Treating Team • Clinical genetics expert Variant verification by orthogonal method • Segregation analysis Curation meeting & draft variant classification • Genetic counselors • Clinical data scientists • Bioinformaticians • Molecular pathologist/geneticist s Genomics Review Board • Additional genetics expertise • Bioethics/legal
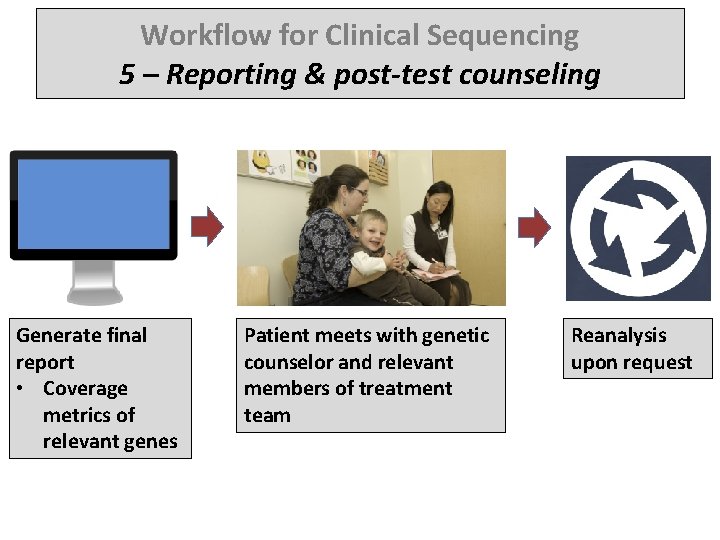
Workflow for Clinical Sequencing 5 – Reporting & post-test counseling Generate final report • Coverage metrics of relevant genes Patient meets with genetic counselor and relevant members of treatment team Reanalysis upon request
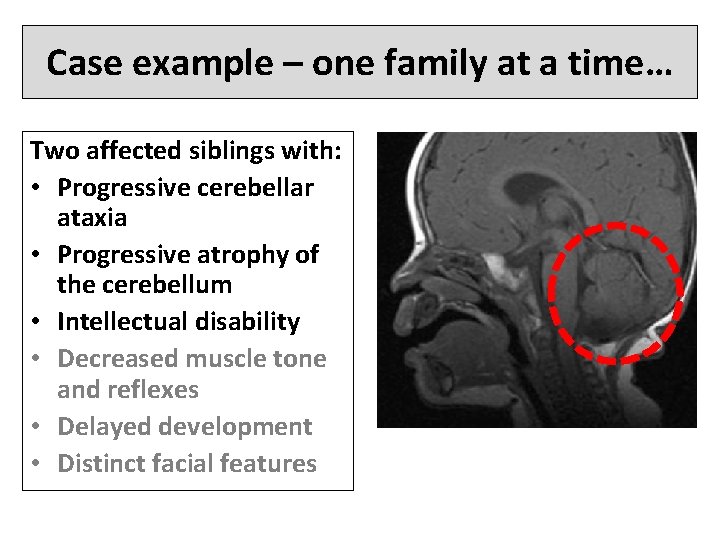
Case example – one family at a time… Two affected siblings with: • Progressive cerebellar ataxia • Progressive atrophy of the cerebellum • Intellectual disability • Decreased muscle tone and reflexes • Delayed development • Distinct facial features

Findings suspicious but not conclusive enough to act on clinically • We performed genome sequencing on this family and found that both copies of a gene (SNX 14) were inactivated in the affected children • Studies in mice indicated that SNX 14 is involved in normal brain and nervous system development • BUT, this gene had not been implicated in human disease


Our service and the treating physicians believe this is the cause of disease in this family • Ends an ~10 year diagnostic odyssey • Provides family information about the disease course and associated management approaches based on the experience of >30 patients • This gene is involved in a cellular process for which there is a therapeutic target that is under investigation in other neurodegenerative disorders
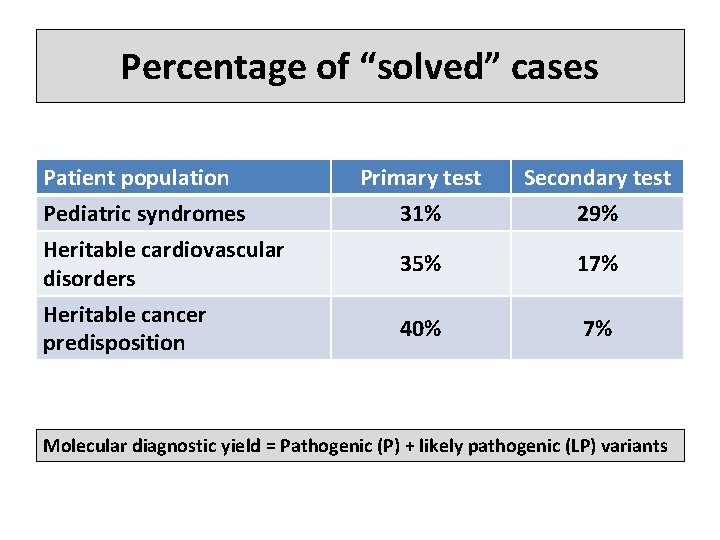
Percentage of “solved” cases Patient population Pediatric syndromes Heritable cardiovascular disorders Heritable cancer predisposition Primary test 31% Secondary test 29% 35% 17% 40% 7% Molecular diagnostic yield = Pathogenic (P) + likely pathogenic (LP) variants

Four major challenges Stanford Medicine Clinical Genomics Service

Challenge 1: Personnel recruitment/retention Faculty • Directors (Ashley/Merker) • Clinical Molecular Geneticist Bioinformatics • Director of Bioinformatics • Bioinformaticians (3) • Software Engineers (2) • Testing Engineer Laboratory • Supervisor • Clinical Laboratory Scientists (2) Interpretation • Clinical Data Scientists (2) • Genetic Counselors (2) Administration • Administrative Director • Administrative Assistant
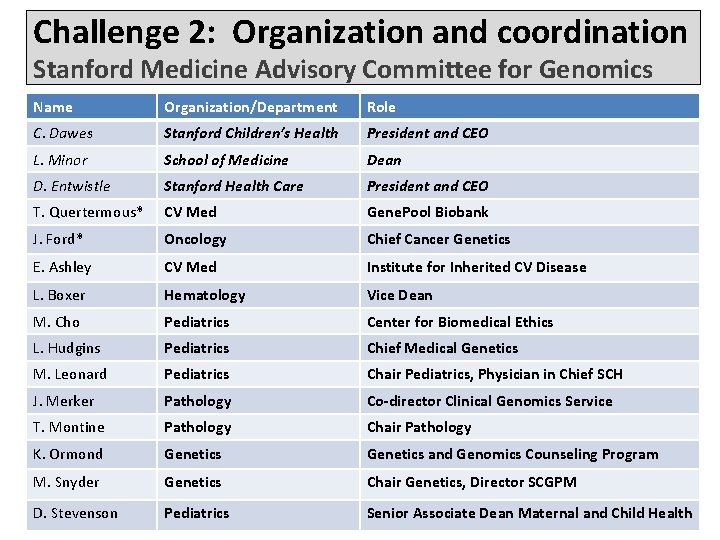
Challenge 2: Organization and coordination Stanford Medicine Advisory Committee for Genomics Name Organization/Department Role C. Dawes Stanford Children’s Health President and CEO L. Minor School of Medicine Dean D. Entwistle Stanford Health Care President and CEO T. Quertermous* CV Med Gene. Pool Biobank J. Ford* Oncology Chief Cancer Genetics E. Ashley CV Med Institute for Inherited CV Disease L. Boxer Hematology Vice Dean M. Cho Pediatrics Center for Biomedical Ethics L. Hudgins Pediatrics Chief Medical Genetics M. Leonard Pediatrics Chair Pediatrics, Physician in Chief SCH J. Merker Pathology Co-director Clinical Genomics Service T. Montine Pathology Chair Pathology K. Ormond Genetics and Genomics Counseling Program M. Snyder Genetics Chair Genetics, Director SCGPM D. Stevenson Pediatrics Senior Associate Dean Maternal and Child Health

Challenge 3: Compute and storage Google Cloud Platform • Limited ability to expand on premise • Build for scale with an uncertain ramp • Find solution that meets security, legal and other business requirement of three distinct entities • HIPAA-compliant implementation
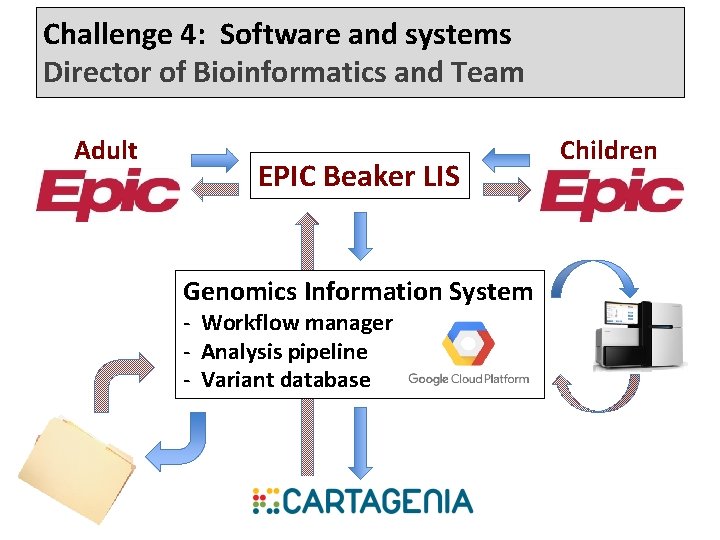
Challenge 4: Software and systems Director of Bioinformatics and Team Adult EPIC Beaker LIS Genomics Information System - Workflow manager - Analysis pipeline - Variant database Children

Update and future directions Stanford Medicine Clinical Genomics Service

Transition from translational pilot to clinical service around September 2017 Dry Laboratory Moved in September 2016 Cloud-based Compute & Storage Dev. environment - ETC November 2016 Prod. environment – ETC September 2017 Wet Laboratory ETC December 2016 Other (in silico) panels: • Heritable cardiovascular • Heritable CA predisposition • Neurologic disease

What’s next… • Improved sequencing approaches – Fill in coverage gaps – Longer read length to identify difficult to call variants – Incorporation of ancillary data (e. g. , RNA-Seq) • Improved data analysis – Identification of difficult to call variants (larger indels, CNVs, SVs) – Analysis of non-coding variants • Increased gene-disease and variant-disease associations – Clinical findings coded with standard terminology – Data sharing efforts • More efficient and improved counseling, consent and result return • Better incorporation of genetic data into clinical practice – Provider education – EMR integration

Acknowledgements Stanford Health Care Clinical Laboratories & David Entwistle* Pathology Stanford Medicine Advisory Thomas Montine* Amir Rubin Clinical Genomics Service Committee on Genomics* James Hereford Stephen Galli James Ford Euan Ashley* Pravene Nath Merrie Bass Thomas Quertermous Somalee Datta Stephen Ayers Tena Cherry Linda Boxer Dianna Fisk Tom Bruynell John Christopher Mildred Cho Jim Ford David Connor Manijeh Danaye-Elmi Louanne Hudgins José González Gary Fritz Shirley Weber Mary Leonard Megan Grove Monica Gupta James Zehnder Kelly Ormond Nathan Hammond Tarun Mahendroo Michael Snyder Louanne Hudgins Stanford Children’s Health Jean-Raymond X. David Stevenson Ruchi Joshi Pierre Christopher Dawes* Isaac Liao Balaji Ramadoss Anne Mc. Cune Stanford University SOM Zena Ng Suzanne Roosevelt Ed Kopetsky Dean Lloyd Minor* Ranjan Muthumalai Neil Shah José González Michael Halaas Suzanne Roosevelt Christopher Sharp Mary Leonard Carlos Suarez Mohan Vasan Harriet Nibbelin Tam Sneddon Yohan Vetteth Natalie Pageler Sowmi Utiramerur Steven Wright Mark O'Connor Shana White Melody Zhang https: //stanfordhealthcare. org/clinicalgenomics Amin Zia SHC and SCH patients and health care teams
- Slides: 30