Essential Cell Biology Third Edition Chapter 3 Energy

Essential Cell Biology Third Edition Chapter 3 Energy, Catalysis and Biosynthesis Copyright © Garland Science 2010

Figure 3 -3 Essential Cell Biology (© Garland Science 2010)

Figure 3 -4 Essential Cell Biology (© Garland Science 2010)

The second law of thermodynamics states that the total entropy of an isolated system always increases over time, or remains constant in ideal cases where the system is in a steady state or undergoing a reversible process. The increase in entropy accounts for the irreversibility of natural processes, and the asymmetry between future and past.
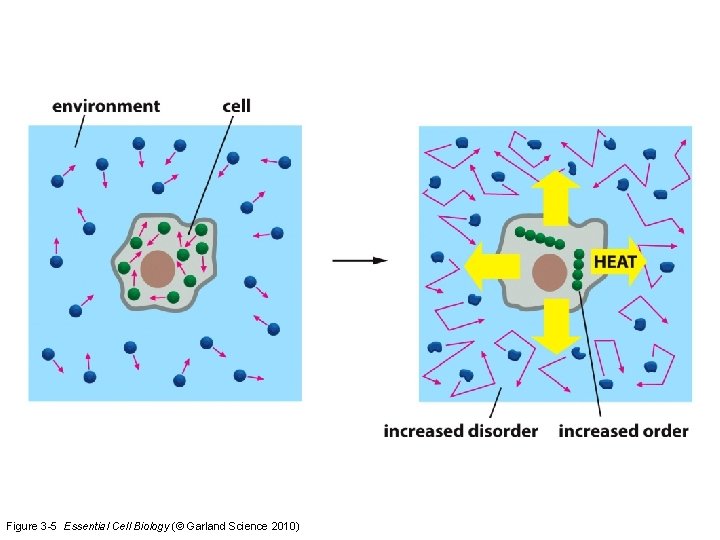
Figure 3 -5 Essential Cell Biology (© Garland Science 2010)
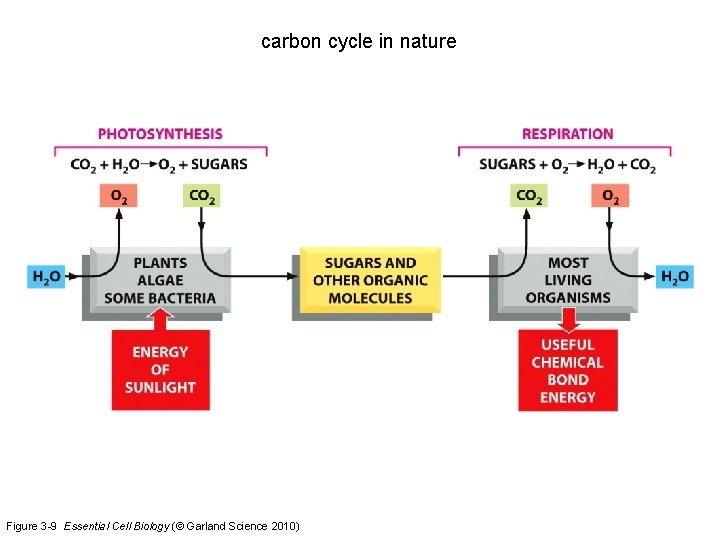
carbon cycle in nature Figure 3 -9 Essential Cell Biology (© Garland Science 2010)
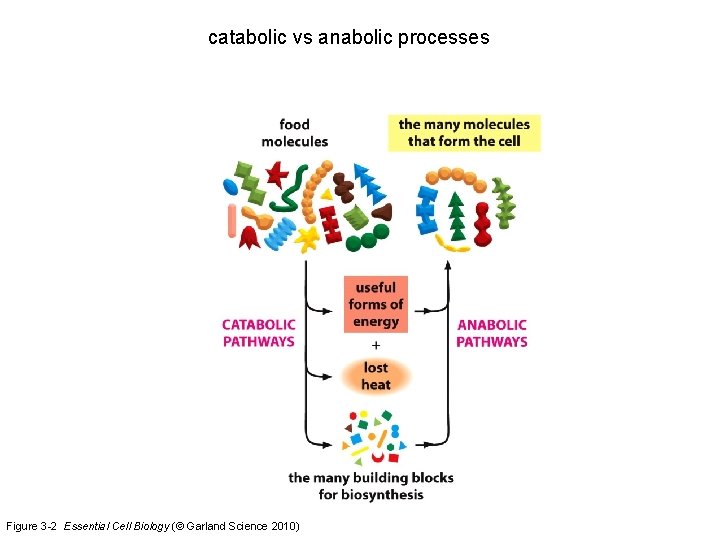
catabolic vs anabolic processes Figure 3 -2 Essential Cell Biology (© Garland Science 2010)
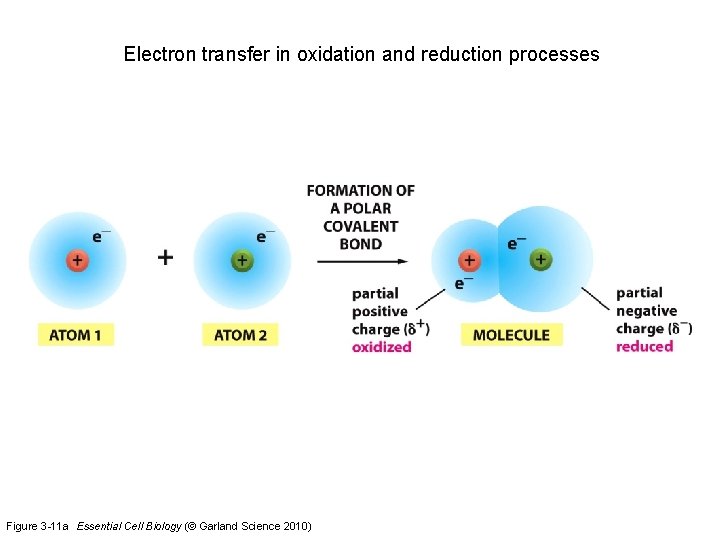
Electron transfer in oxidation and reduction processes Figure 3 -11 a Essential Cell Biology (© Garland Science 2010)
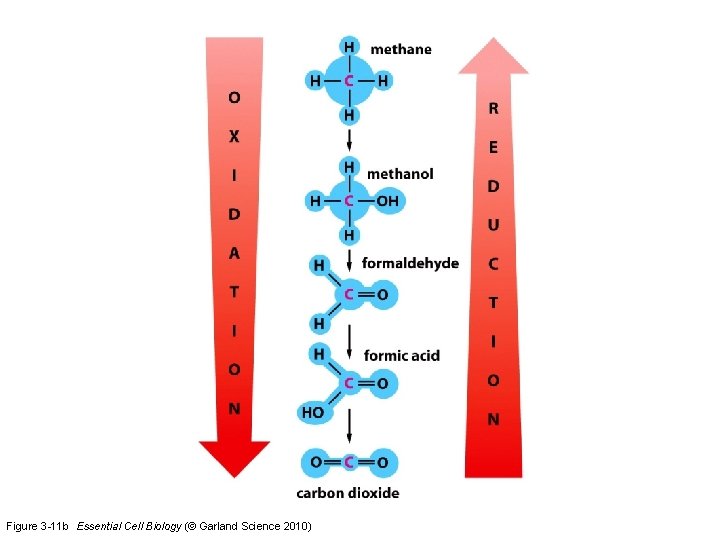
Figure 3 -11 b Essential Cell Biology (© Garland Science 2010)
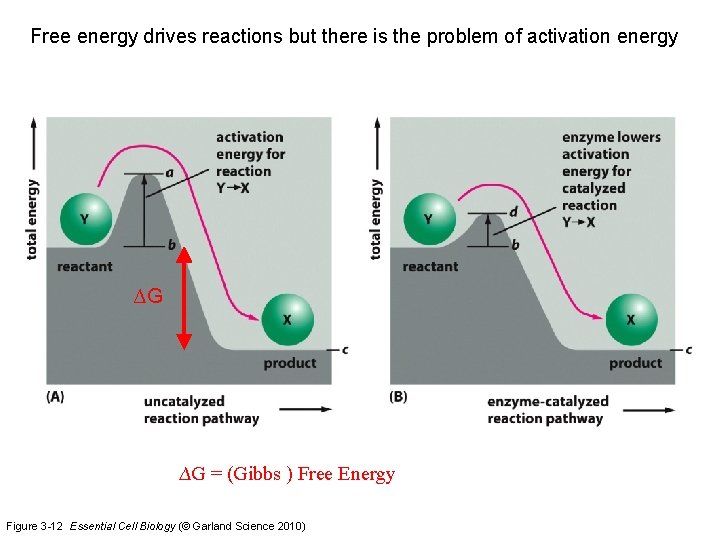
Free energy drives reactions but there is the problem of activation energy ∆G ∆G = (Gibbs ) Free Energy Figure 3 -12 Essential Cell Biology (© Garland Science 2010)
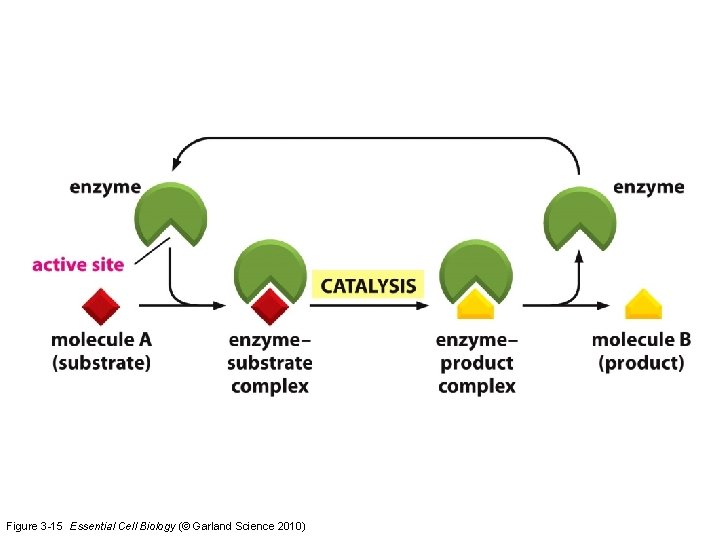
Figure 3 -15 Essential Cell Biology (© Garland Science 2010)
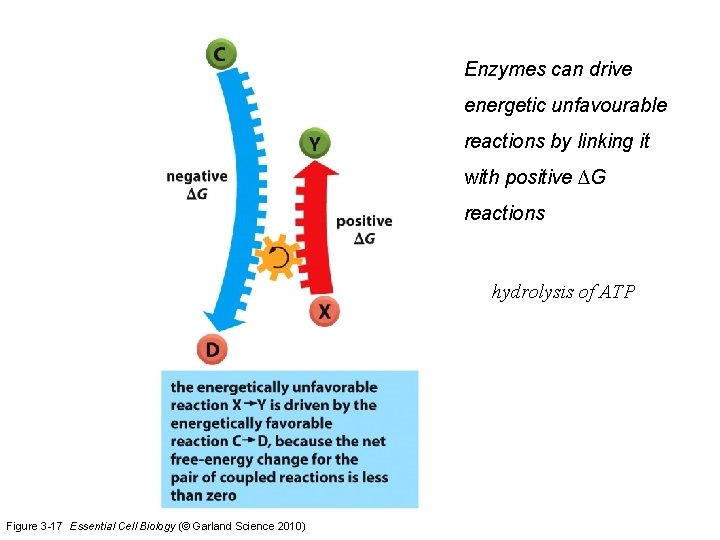
Enzymes can drive energetic unfavourable reactions by linking it with positive ∆G reactions hydrolysis of ATP Figure 3 -17 Essential Cell Biology (© Garland Science 2010)
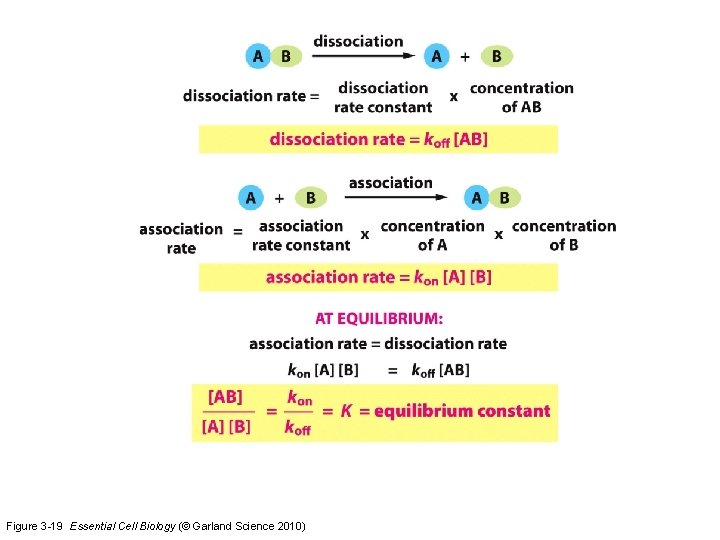
Figure 3 -19 Essential Cell Biology (© Garland Science 2010)
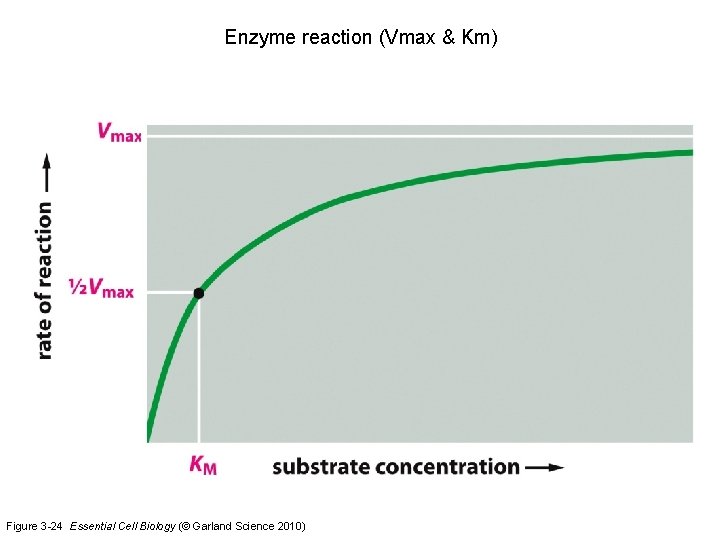
Enzyme reaction (Vmax & Km) Figure 3 -24 Essential Cell Biology (© Garland Science 2010)
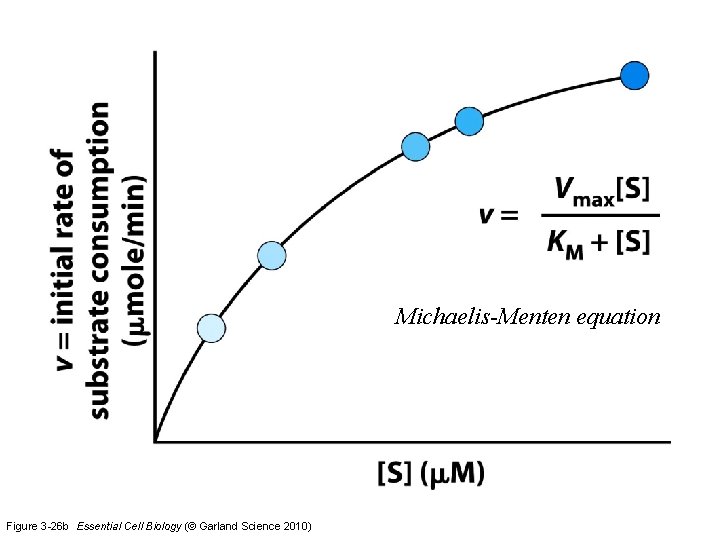
Michaelis-Menten equation Figure 3 -26 b Essential Cell Biology (© Garland Science 2010)
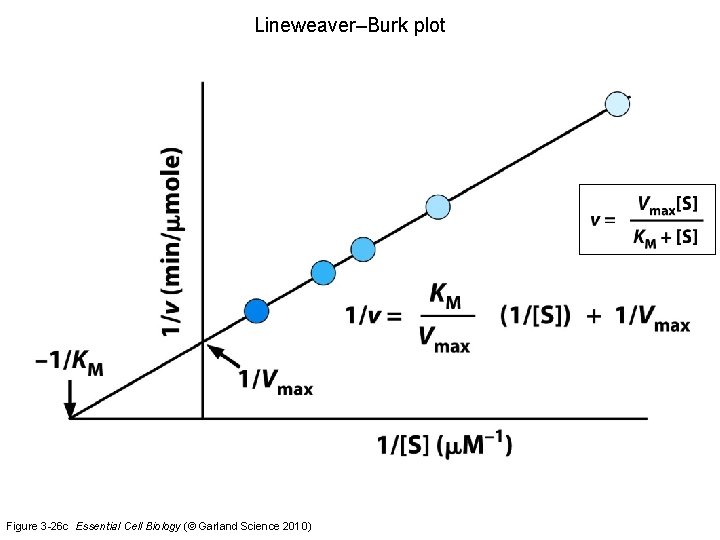
Lineweaver–Burk plot Figure 3 -26 c Essential Cell Biology (© Garland Science 2010)
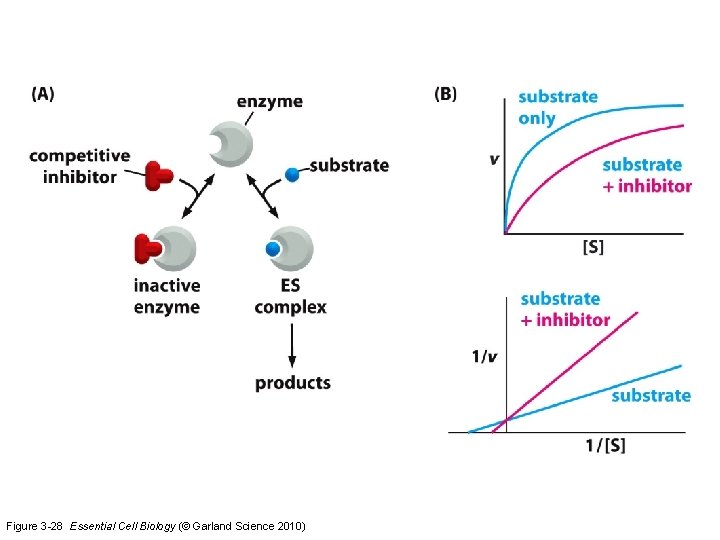
Figure 3 -28 Essential Cell Biology (© Garland Science 2010)
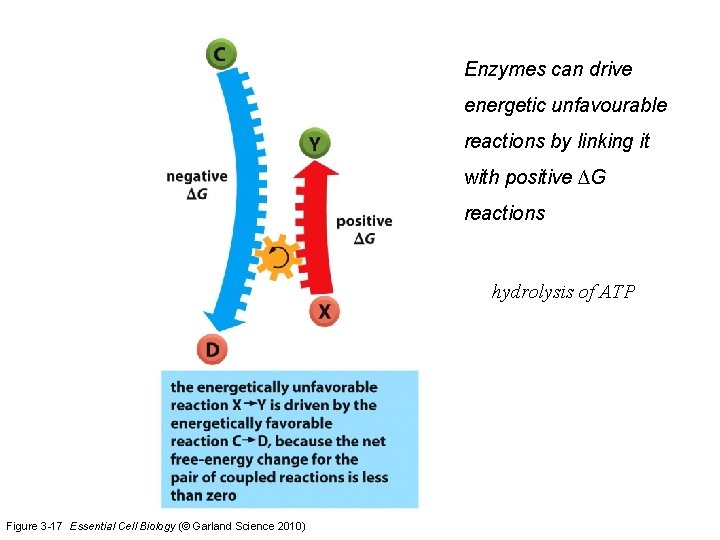
Enzymes can drive energetic unfavourable reactions by linking it with positive ∆G reactions hydrolysis of ATP Figure 3 -17 Essential Cell Biology (© Garland Science 2010)
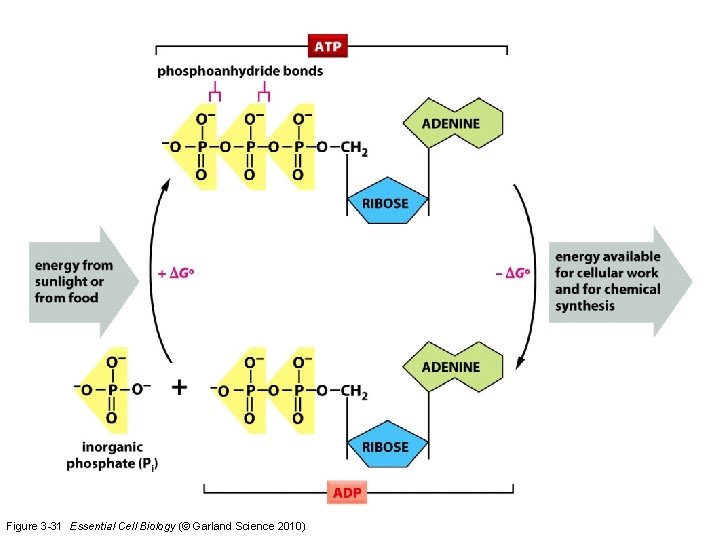
Figure 3 -31 Essential Cell Biology (© Garland Science 2010)
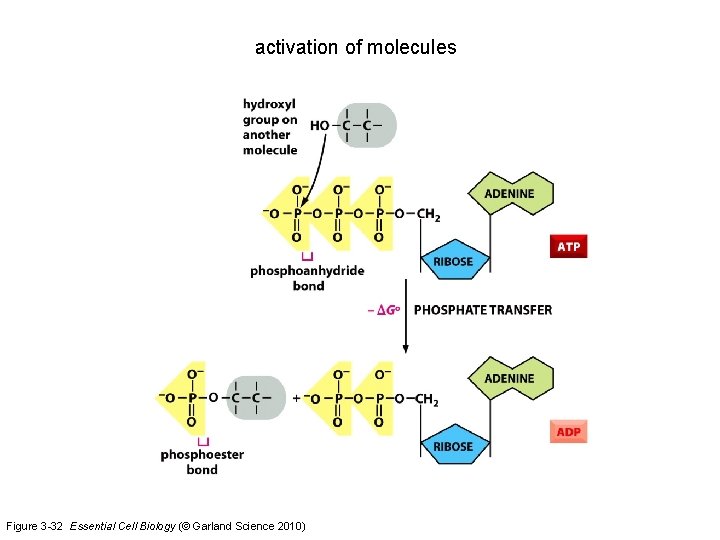
activation of molecules Figure 3 -32 Essential Cell Biology (© Garland Science 2010)
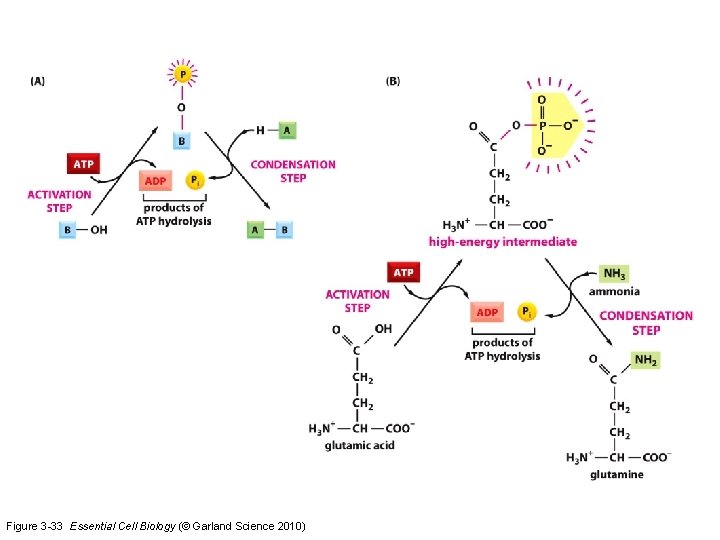
Figure 3 -33 Essential Cell Biology (© Garland Science 2010)
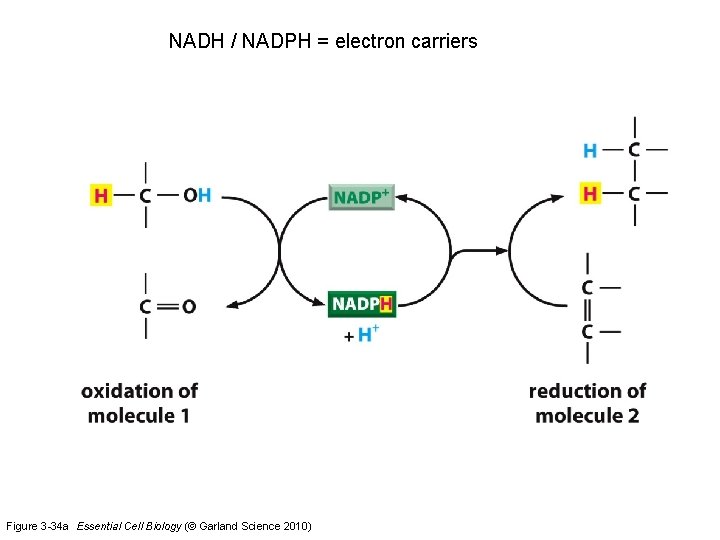
NADH / NADPH = electron carriers Figure 3 -34 a Essential Cell Biology (© Garland Science 2010)
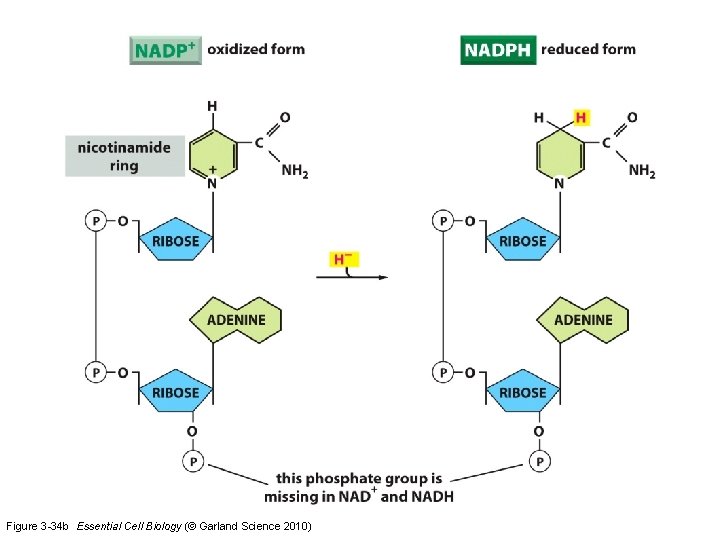
Figure 3 -34 b Essential Cell Biology (© Garland Science 2010)
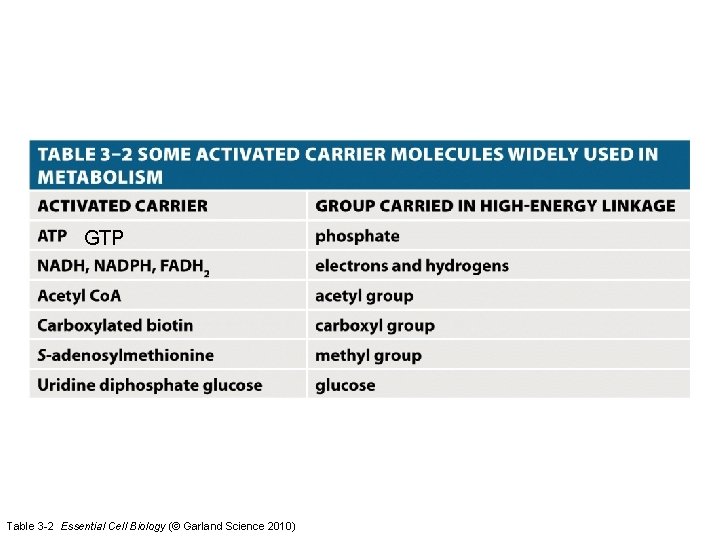
GTP Table 3 -2 Essential Cell Biology (© Garland Science 2010)
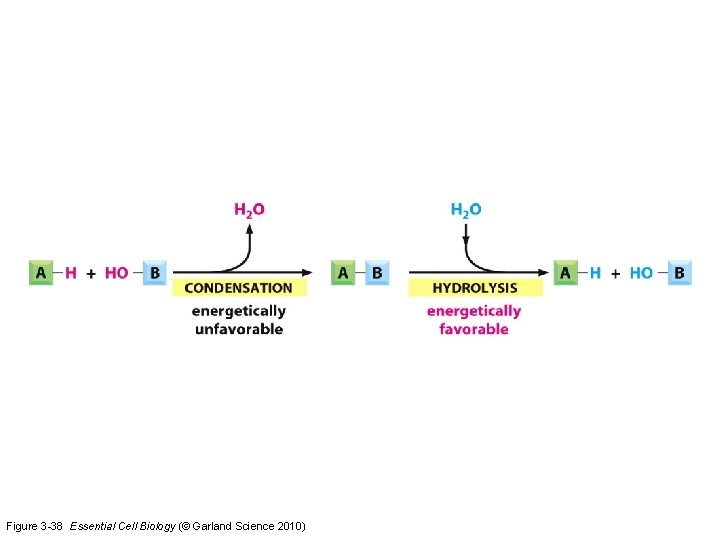
Figure 3 -38 Essential Cell Biology (© Garland Science 2010)
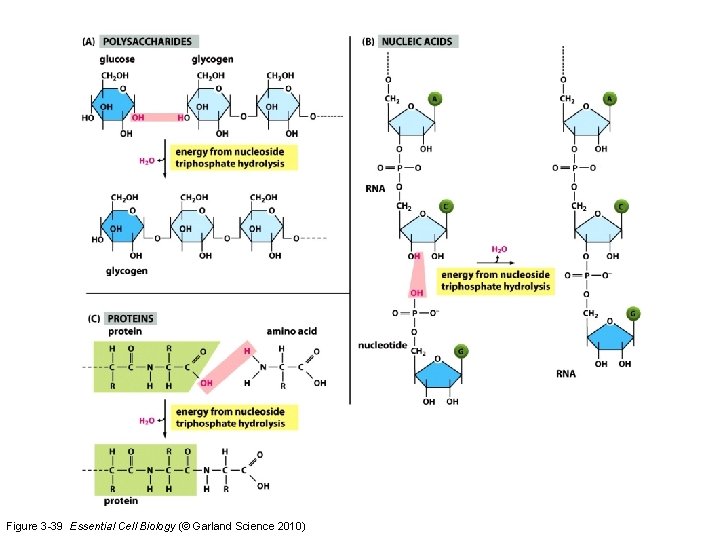
Figure 3 -39 Essential Cell Biology (© Garland Science 2010)

Essential Cell Biology Third Edition Chapter 4 Protein Structure and Function Copyright © Garland Science 2010
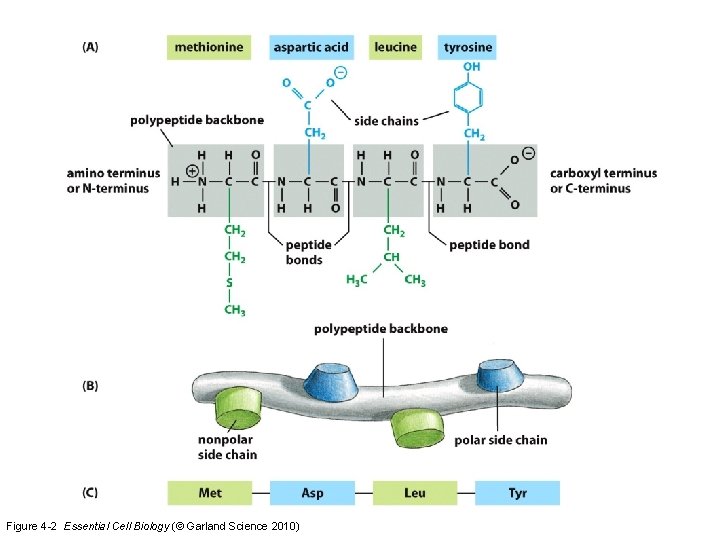
Figure 4 -2 Essential Cell Biology (© Garland Science 2010)
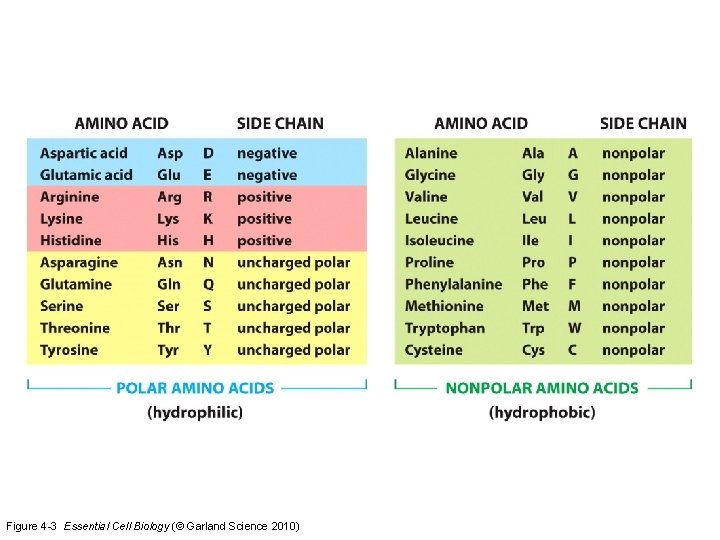
Figure 4 -3 Essential Cell Biology (© Garland Science 2010)
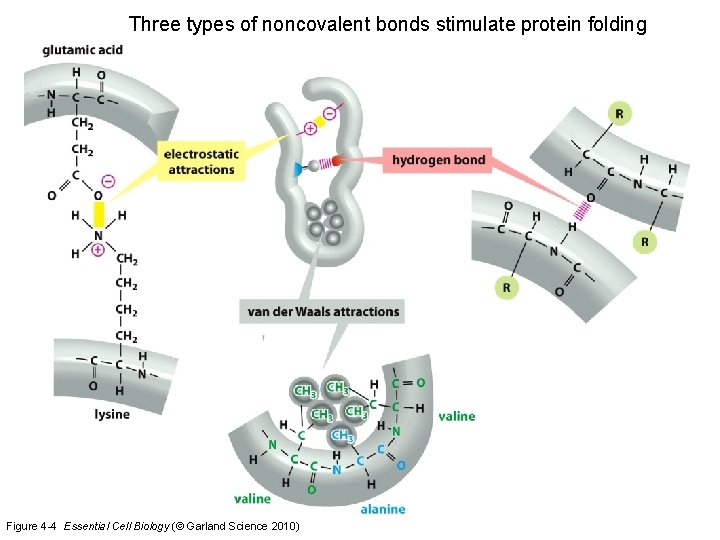
Three types of noncovalent bonds stimulate protein folding Figure 4 -4 Essential Cell Biology (© Garland Science 2010)
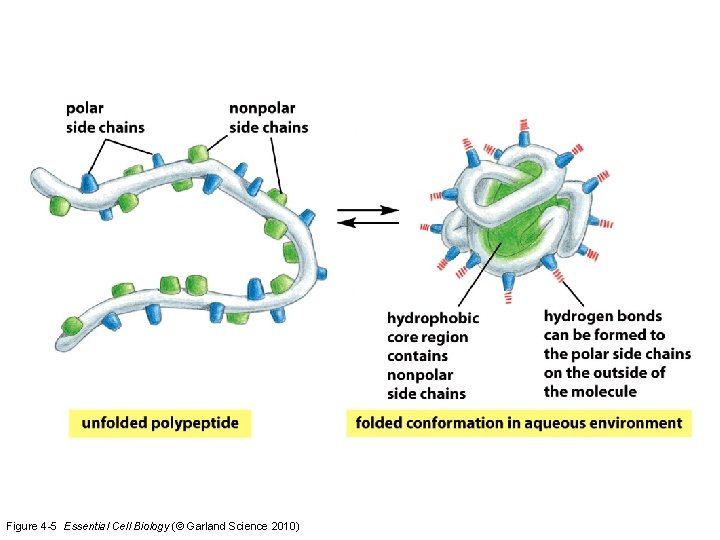
Figure 4 -5 Essential Cell Biology (© Garland Science 2010)
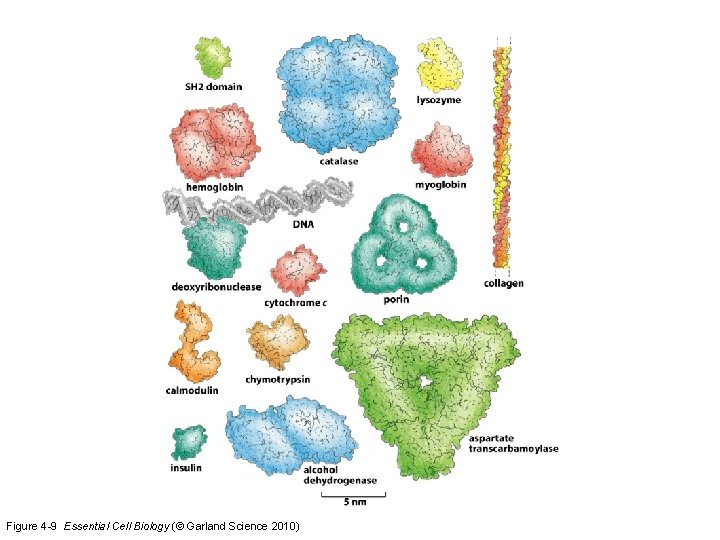
Figure 4 -9 Essential Cell Biology (© Garland Science 2010)
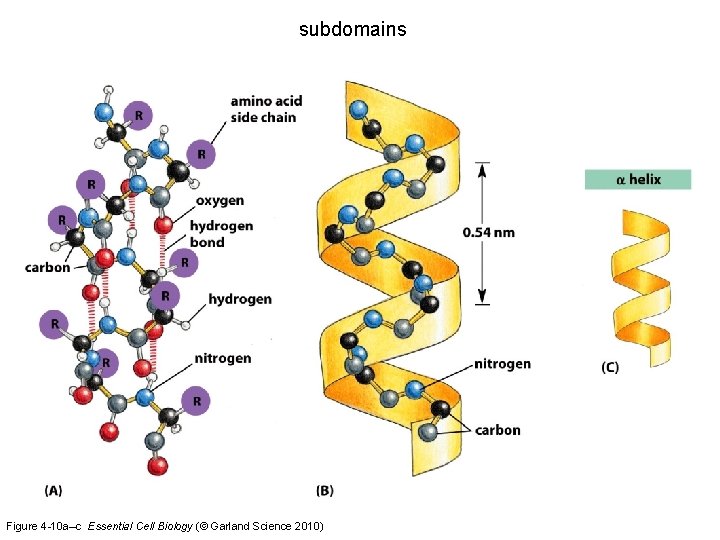
subdomains Figure 4 -10 a–c Essential Cell Biology (© Garland Science 2010)
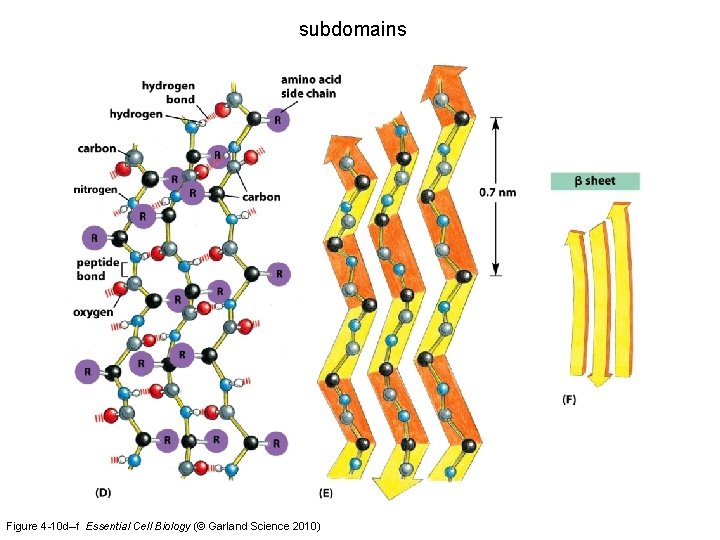
subdomains Figure 4 -10 d–f Essential Cell Biology (© Garland Science 2010)
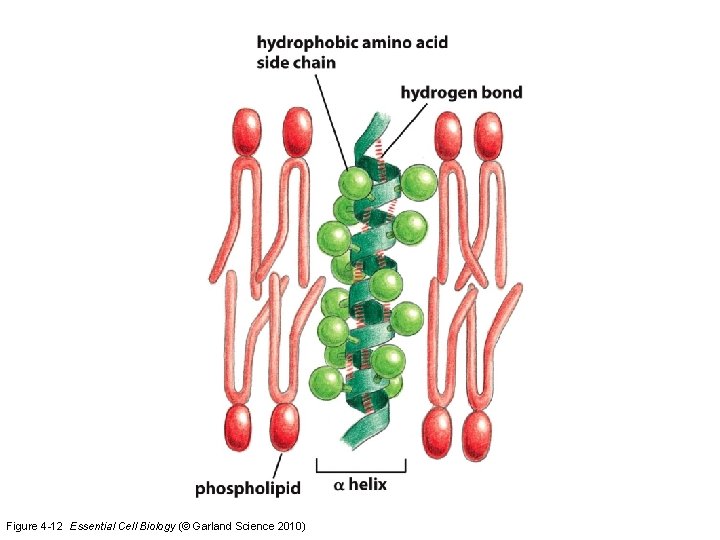
Figure 4 -12 Essential Cell Biology (© Garland Science 2010)
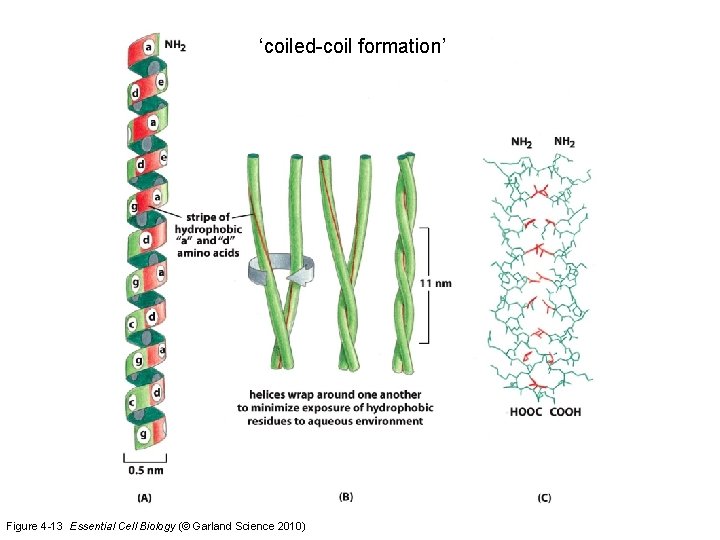
‘coiled-coil formation’ Figure 4 -13 Essential Cell Biology (© Garland Science 2010)
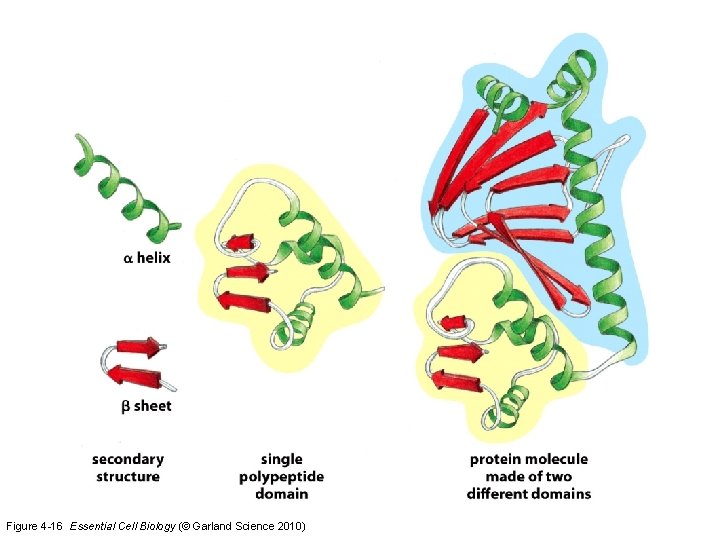
Figure 4 -16 Essential Cell Biology (© Garland Science 2010)
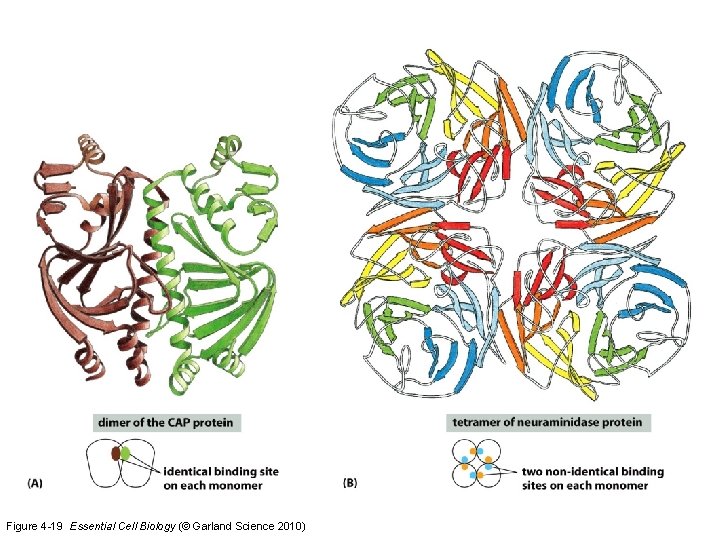
Figure 4 -19 Essential Cell Biology (© Garland Science 2010)
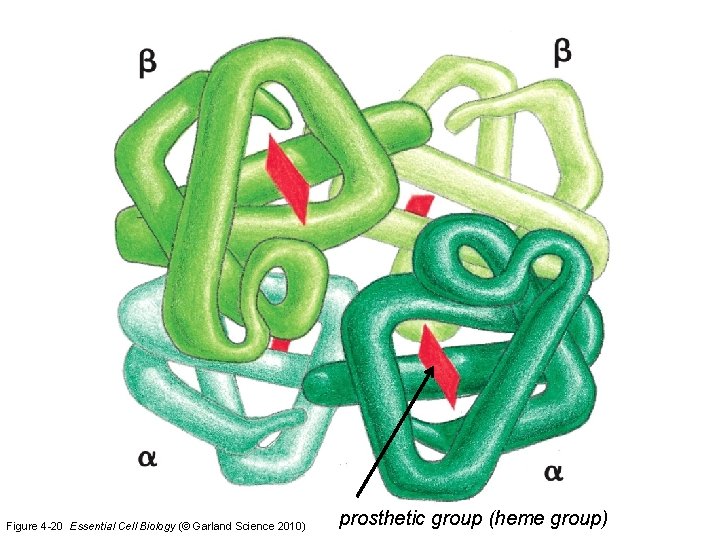
Figure 4 -20 Essential Cell Biology (© Garland Science 2010) prosthetic group (heme group)
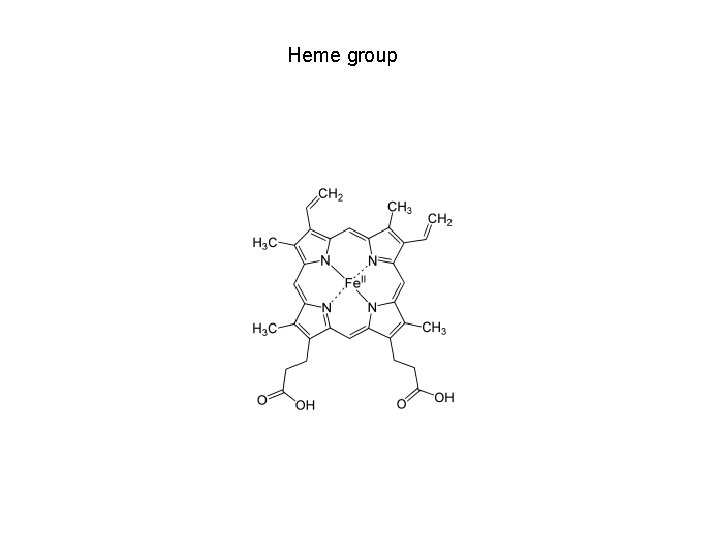
Heme group
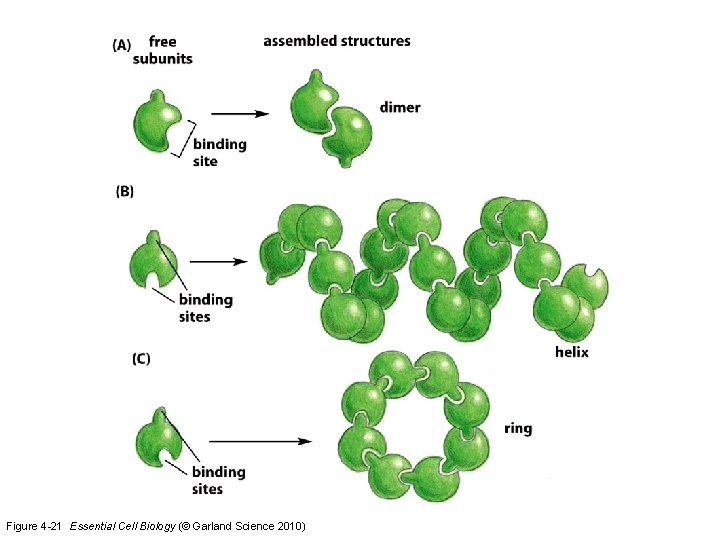
Figure 4 -21 Essential Cell Biology (© Garland Science 2010)
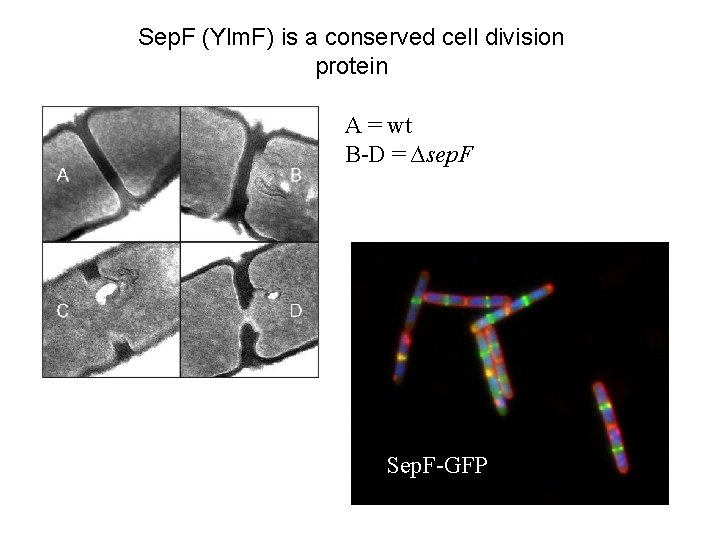
Sep. F (Ylm. F) is a conserved cell division protein A = wt B-D = ∆sep. F Sep. F-GFP
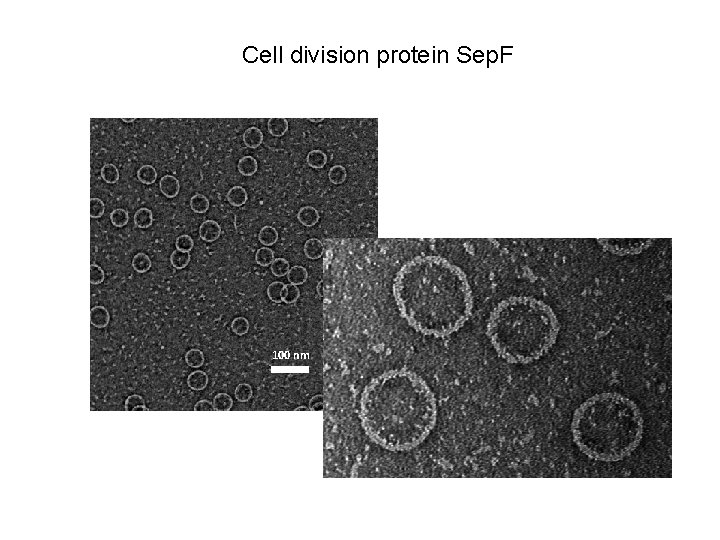
Cell division protein Sep. F 100 nm

Effect of Sep. F on Fts. Z bundling Fts. Z +GTP+ Sep. F 100 nm
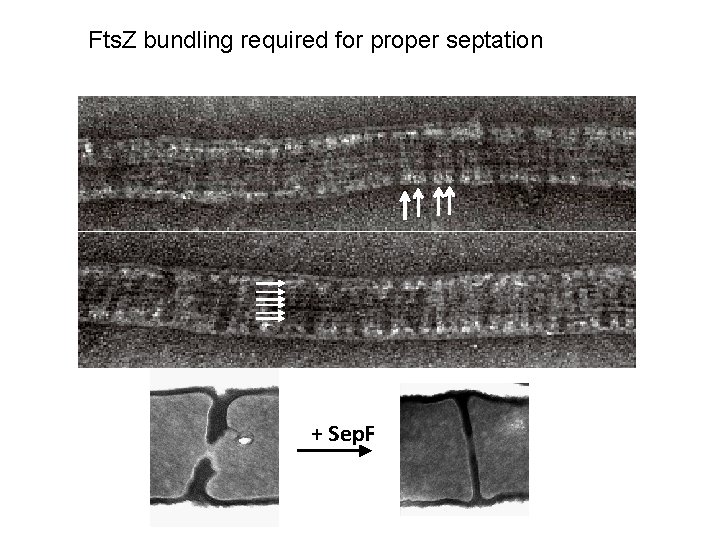
Fts. Z bundling required for proper septation + Sep. F
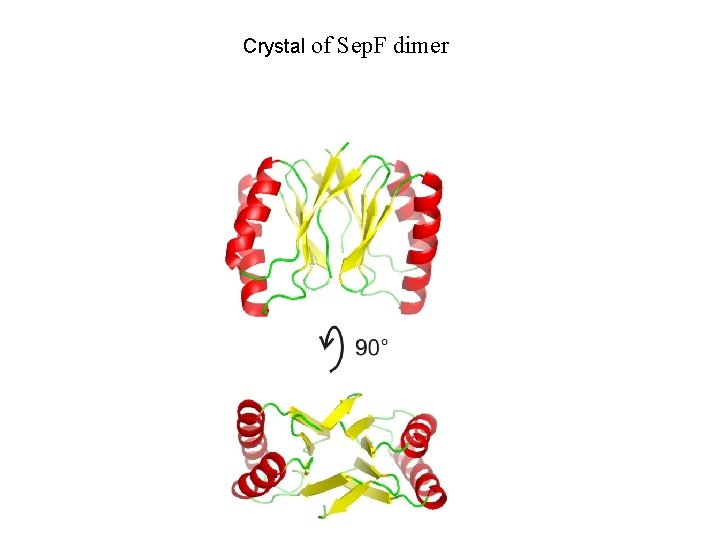
Crystal of Sep. F dimer
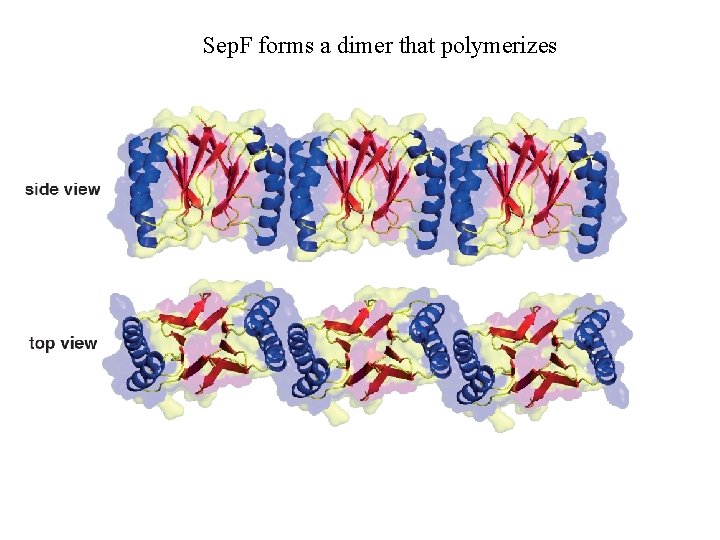
Sep. F forms a dimer that polymerizes
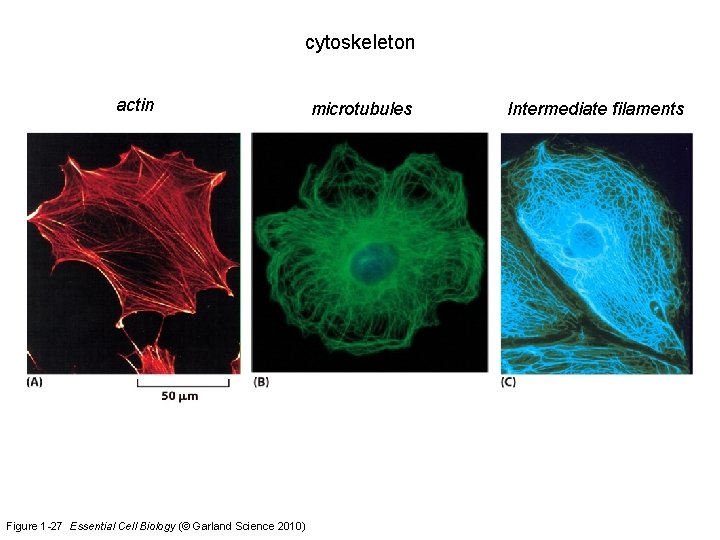
cytoskeleton actin Figure 1 -27 Essential Cell Biology (© Garland Science 2010) microtubules Intermediate filaments
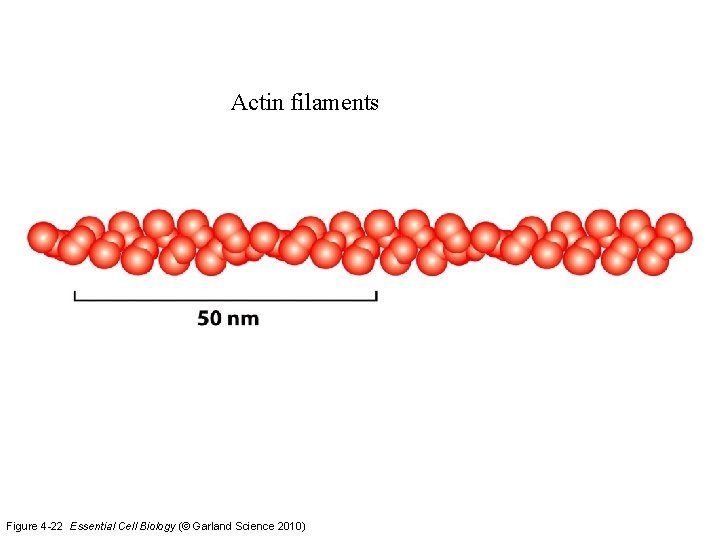
Actin filaments Figure 4 -22 Essential Cell Biology (© Garland Science 2010)
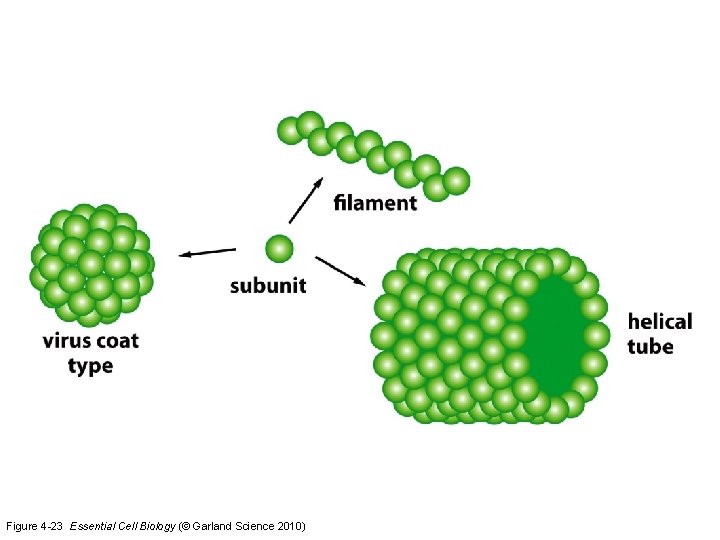
Figure 4 -23 Essential Cell Biology (© Garland Science 2010)
Figure 4 -24 Essential Cell Biology (© Garland Science 2010)
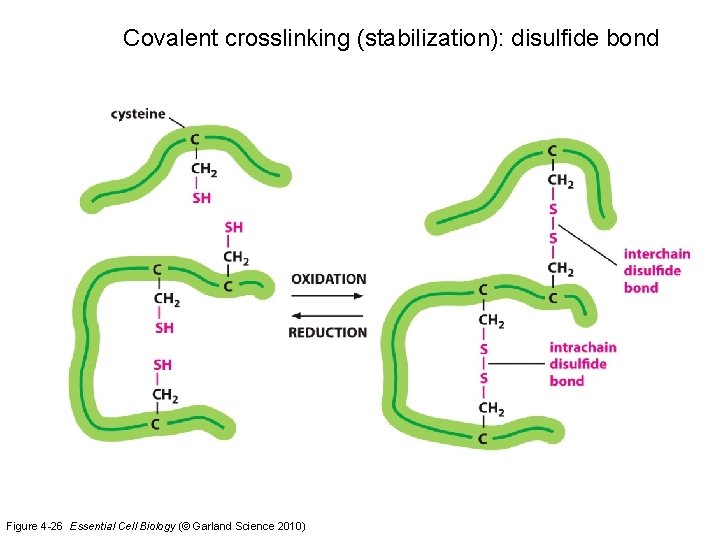
Covalent crosslinking (stabilization): disulfide bond Figure 4 -26 Essential Cell Biology (© Garland Science 2010)
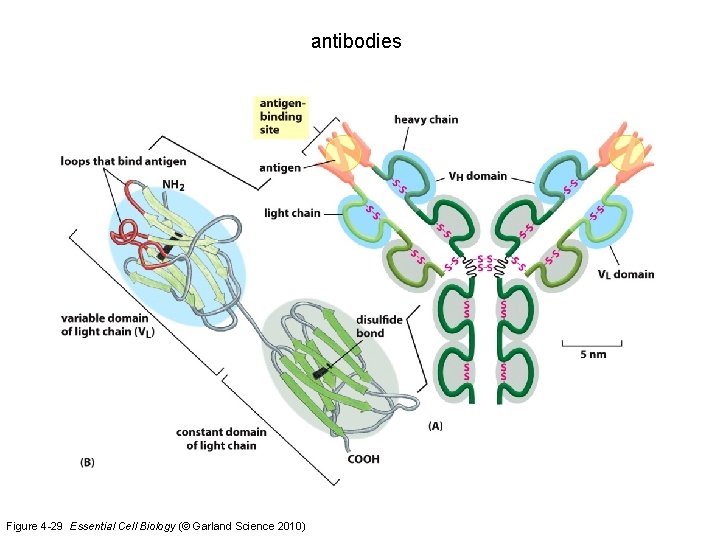
antibodies Figure 4 -29 Essential Cell Biology (© Garland Science 2010)
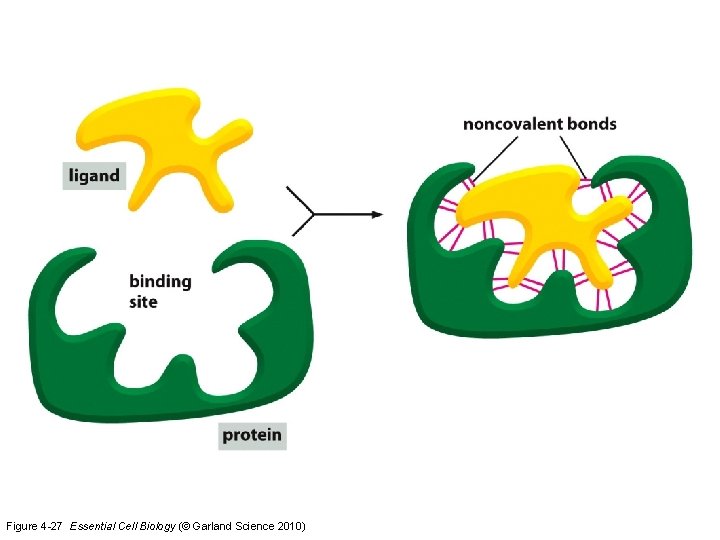
Figure 4 -27 Essential Cell Biology (© Garland Science 2010)
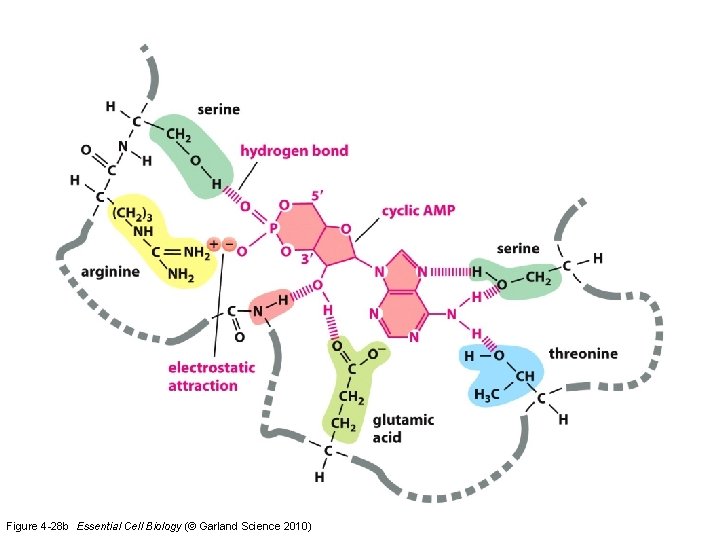
Figure 4 -28 b Essential Cell Biology (© Garland Science 2010)
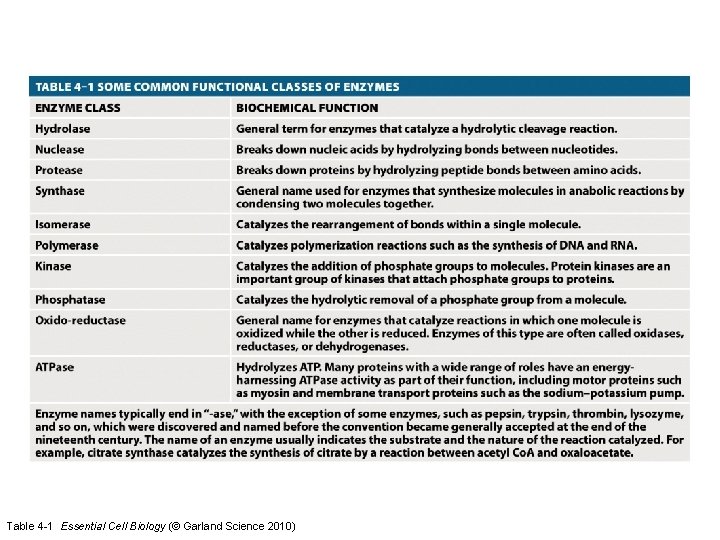
Table 4 -1 Essential Cell Biology (© Garland Science 2010)
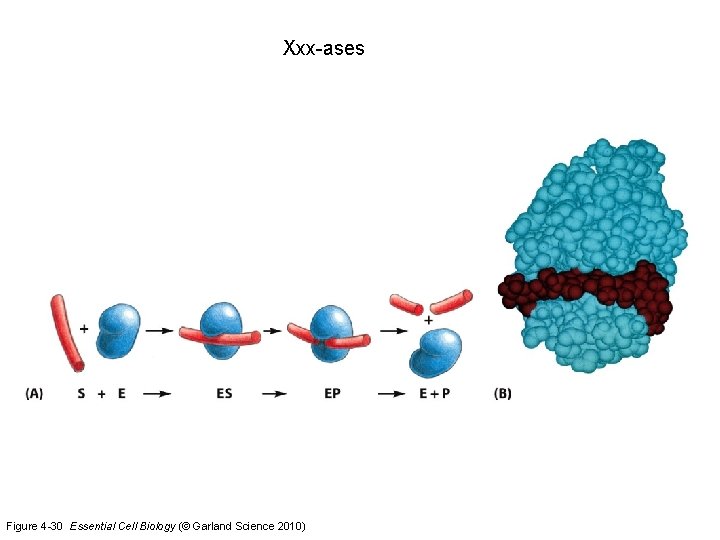
Xxx-ases Figure 4 -30 Essential Cell Biology (© Garland Science 2010)
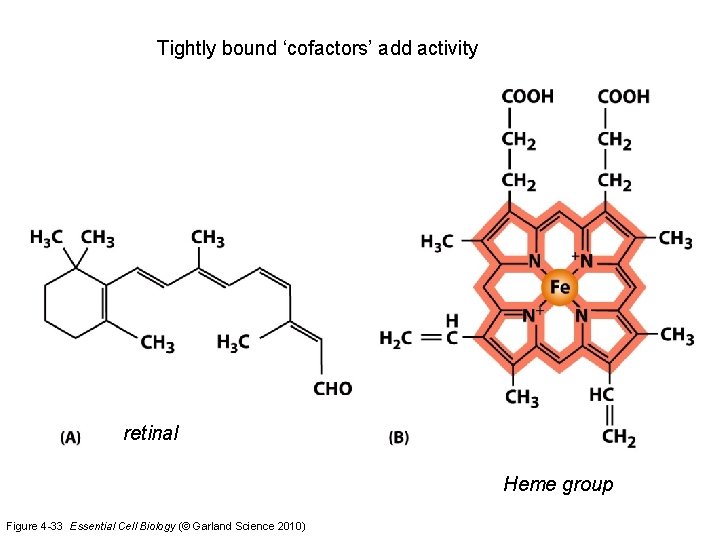
Tightly bound ‘cofactors’ add activity retinal Heme group Figure 4 -33 Essential Cell Biology (© Garland Science 2010)
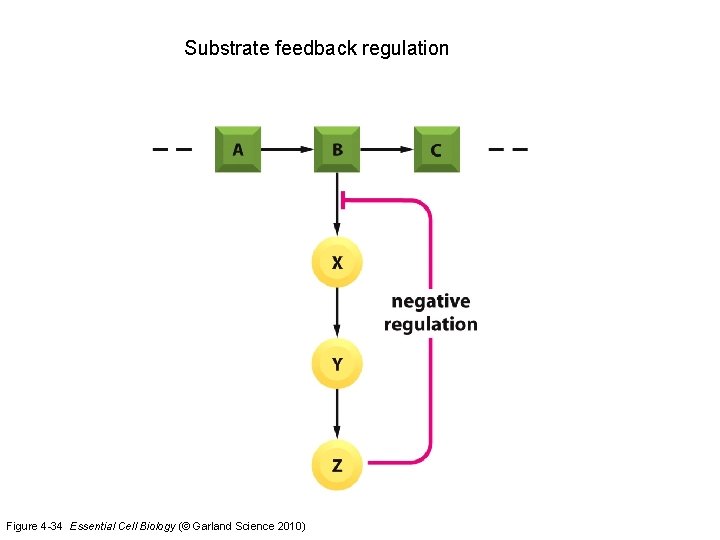
Substrate feedback regulation Figure 4 -34 Essential Cell Biology (© Garland Science 2010)
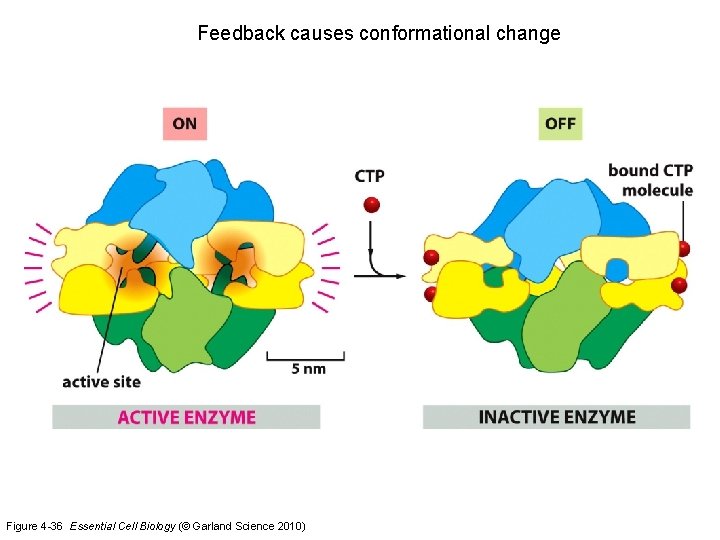
Feedback causes conformational change Figure 4 -36 Essential Cell Biology (© Garland Science 2010)
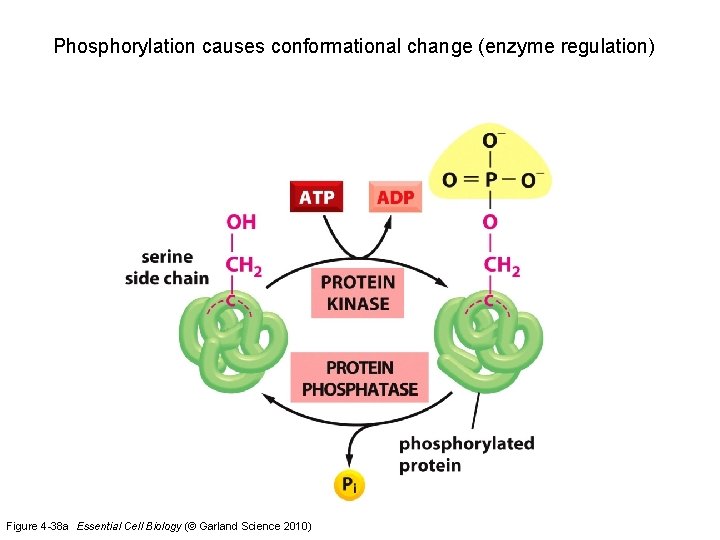
Phosphorylation causes conformational change (enzyme regulation) Figure 4 -38 a Essential Cell Biology (© Garland Science 2010)
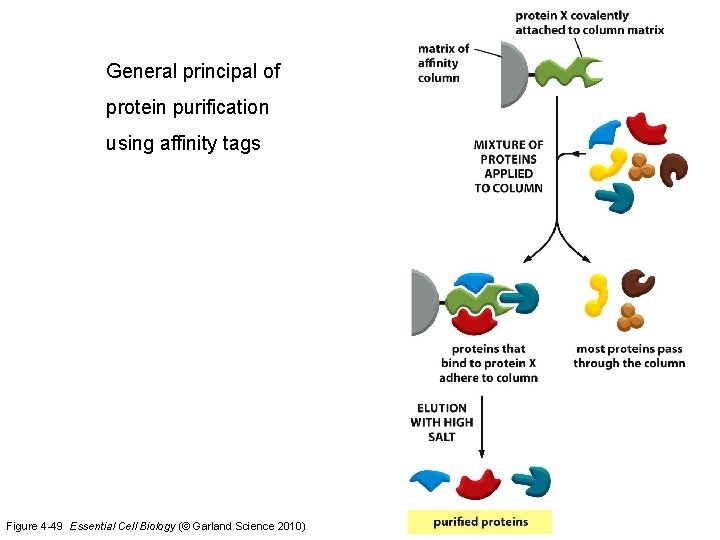
General principal of protein purification using affinity tags Figure 4 -49 Essential Cell Biology (© Garland Science 2010)
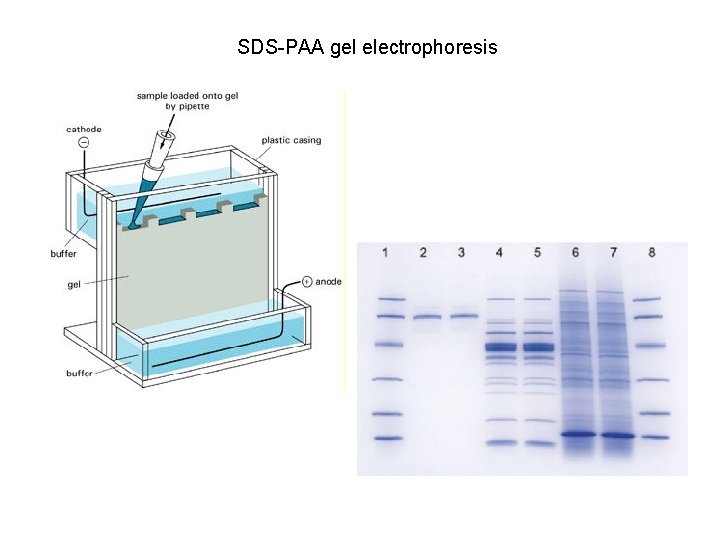
SDS-PAA gel electrophoresis
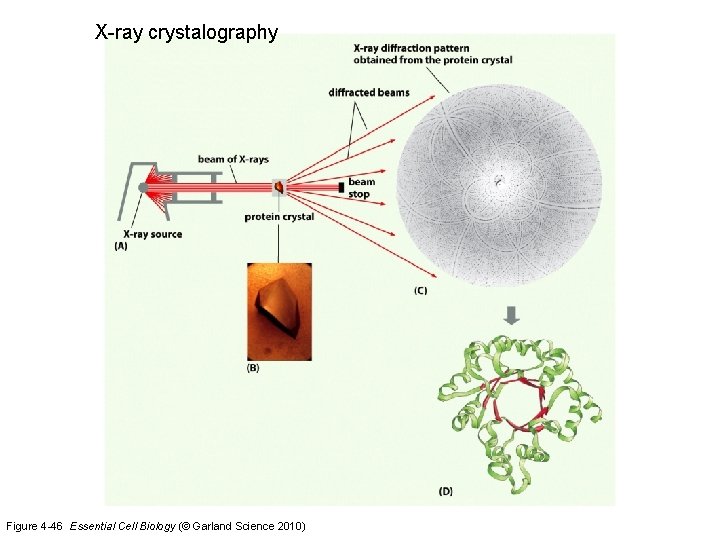
X-ray crystalography Figure 4 -46 Essential Cell Biology (© Garland Science 2010)
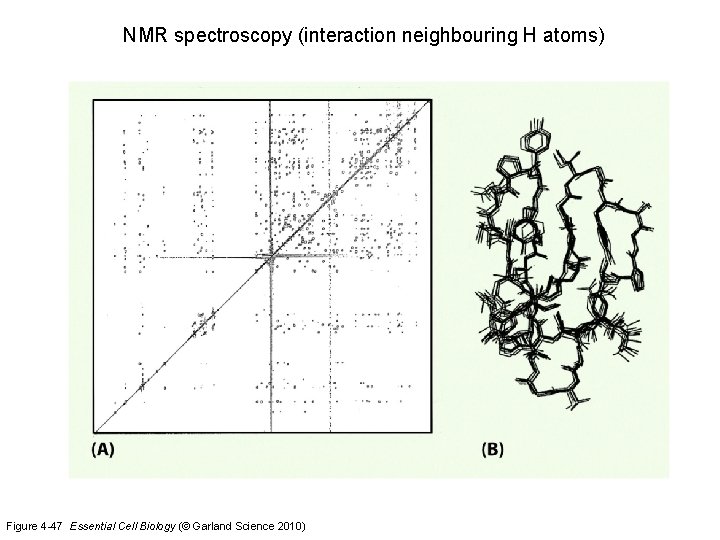
NMR spectroscopy (interaction neighbouring H atoms) Figure 4 -47 Essential Cell Biology (© Garland Science 2010)

Keywords Chapter 3 : entropy carbon cycle catabolic/anabolic oxidation/reduction (methane/CO 2/O 2) activation energy (enzymes) standard free energy ∆G equilibrium constant, Vmax, Km (effect substrate inhibitor) activation of bond by phosphorylation oxidation/reduction main energy carriers (ATP, GTP, NAD(P)H, FADH 2, acetyl-Co. A Keywords Chapter 4: amino acids (polar (neutral, positive, negative) hydrophobic) electrostatic interaction, hydrogen bond, hydrophobic (vd Waals) alpha-helix (e. g. trans-membrane helix), beta-sheet, coiled-coil formation cysteine bridges monomer, dimer, oligomer, polymer ligand - enzyme hydrolase, nuclease, protease, synthase, isomerase, polymerase, kinase, phosphatase, ATPase cofactor (e. g. heme) substrate feedback regulation SDS-PAA gel electrophoresis

Examples Exam questions 5: What process in the carbon cycle produces CO 2 ? (A) respiration (B) anabolism (C) catabolism (D) photosynthesis 6: What is the Km in enzyme reactions ? (A) reaction constant that determines Vmax (B) substrate concentration that determines Vmax (C) reaction constant that determines the enzyme concentration (D) substrate concentration that results in 50% of the maximum reaction rate 7: Which statement is correct ? (A) condensation produces energy and hydrolysis produces energy (B) condensation produces energy and hydrolysis requires energy (C) condensation requires energy and hydrolysis requires energy (D) condensation requires energy and hydrolysis produces energy 8: Which noncovalent interactions help to fold a protein ? (A) electrostatic interactions, hydrogen bonds, hydrophobic interactions (B) electrostatic interactions and cysteine bridges (C) cysteine bridges (D) condensation interactions
- Slides: 67