Epigenetics RNAi and Heterochromatin formation RNAi pathways RNAi







- Slides: 10

Epigenetics: RNAi and Heterochromatin formation

RNAi pathways • RNAi (RNA interference) -> silencing by ds. RNA • ds. RNA -> si. RNA -> target m. RNA -> Post. Transcriptional Gene Silencing (PTGS) • ds. RNA -> si. RNA -> target chromatin -> Transcriptional Gene Silencing (TGS) • Dicer: RNase III class ribonuclease -> convert ds. RNA -> si. RNA • Argonaute/PIWI family: RNAse H -> cleaving target m. RNA

Three main RNAi complexes 1 RISC (RNA-Induced Silencing Complex): si. RNA, Argonaute, -> remove target m. RNAs (PTGS) 2 RITS (RNA-Induced Transcriptional Silencing Complex): si. RNA, target chromatins (TGS) 3 mi. RNA-mediated silencing mechanism: endogenous non-coding gene -> mi. RNA by Dicer and Argonaute -> suppress translation

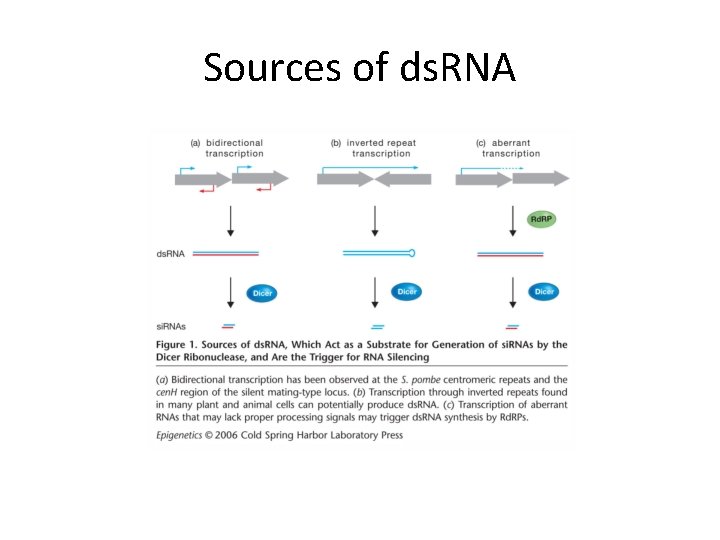
Sources of ds. RNA

Early indications • PSTV (potato spindle tuber viroid): a 359 -nt RNA virus triggers DNA methylation on any transgenic PSTV sequences in the host genome, and this requires the viroid transcription • In plants, aberrant transcripts trigger DNA methylation of similar sequences

Connection of RNAi with Heterochromatin formation Mutations in RNAi complex (dcr 1, ago 1, rdp 1) in S. pombe caused loss of heterochromatin and accumulation of repeat-derived transcripts Sequencing of a small RNA library (~22 nt long) derived many hit from centromeric repeats

ds. RNA and si. RNA generation • Bi-directional transcription of repeats • Transcription of inverted repeats • In S. pombe, RDRC (RNA-directed RNA polymerase Complex) is required for si. RNA production: Rdp 1, Cid 12, Hrr 1

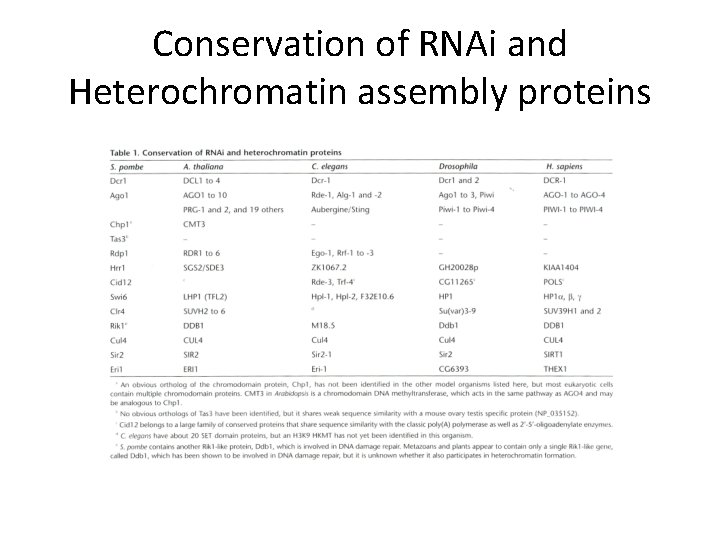
Conservation of RNAi and Heterochromatin assembly proteins

si. RNA target via RNA-RNA models Two models for targeting RITS RNA-DNA model Vs RNA-RNA model Disturbance in transcription causes failure in PTGS Localizing RITS complex to centromeric regions, which is also si. RNA dependent

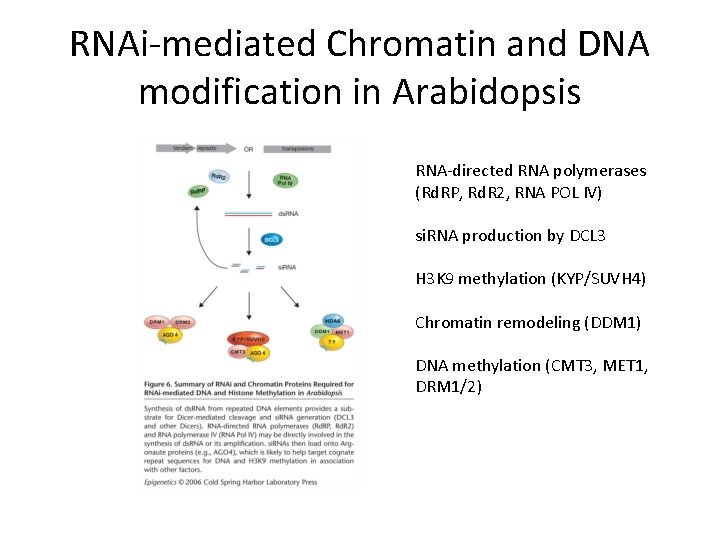
RNAi-mediated Chromatin and DNA modification in Arabidopsis RNA-directed RNA polymerases (Rd. RP, Rd. R 2, RNA POL IV) si. RNA production by DCL 3 H 3 K 9 methylation (KYP/SUVH 4) Chromatin remodeling (DDM 1) DNA methylation (CMT 3, MET 1, DRM 1/2)