EPIGENETICS Epigenetics possible definitions Epigenetics comprises hereditary changes
EPIGENETICS

Epigenetics - possible definition(s) Ø Epigenetics comprises (hereditary) changes in gene expression without altering the DNA sequence Ø Epigenetics is a biological subdiscipline that deals with questions of development, heredity, gene regulation and evolution

1 2 Which pair of twins is identical? Both of them! Young twins hardly differed in their epigenetic code - older twins, however, even more so
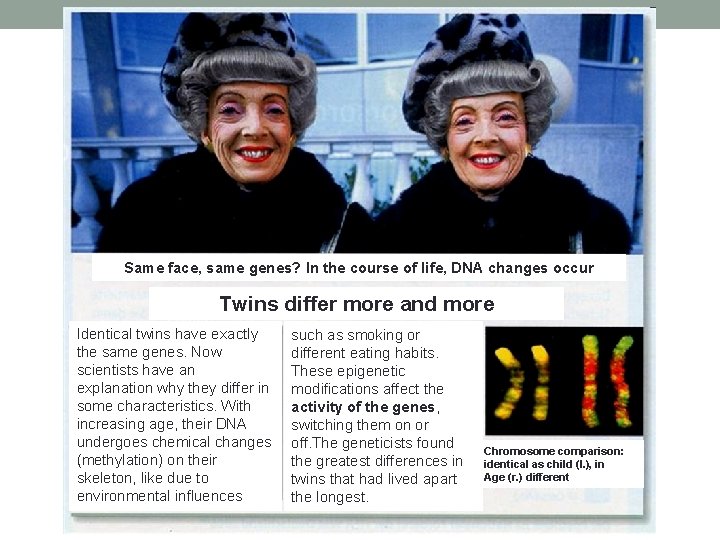
Same face, same genes? In the course of life, DNA changes occur Twins differ more and more Identical twins have exactly the same genes. Now scientists have an explanation why they differ in some characteristics. With increasing age, their DNA undergoes chemical changes (methylation) on their skeleton, like due to environmental influences such as smoking or different eating habits. These epigenetic modifications affect the activity of the genes, switching them on or off. The geneticists found the greatest differences in twins that had lived apart the longest. Chromosome comparison: identical as child (I. ), in Age (r. ) different

Epigenetic mechanisms 1. DNA methylation 2. histone acetylation 3. micro. RNA (mi. RNA)
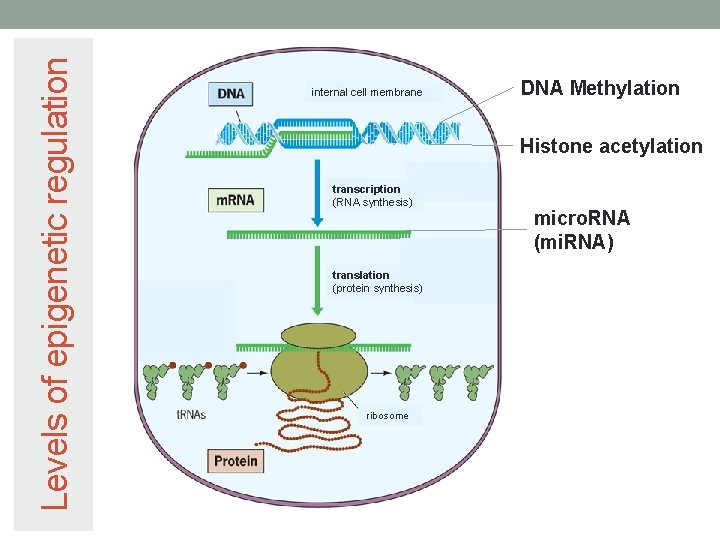
Levels of epigenetic regulation internal cell membrane DNA Methylation Histone acetylation transcription (RNA synthesis) translation (protein synthesis) ribosome micro. RNA (mi. RNA)

DNA METHYLATION AS EPIGENETIC MODIFICATION
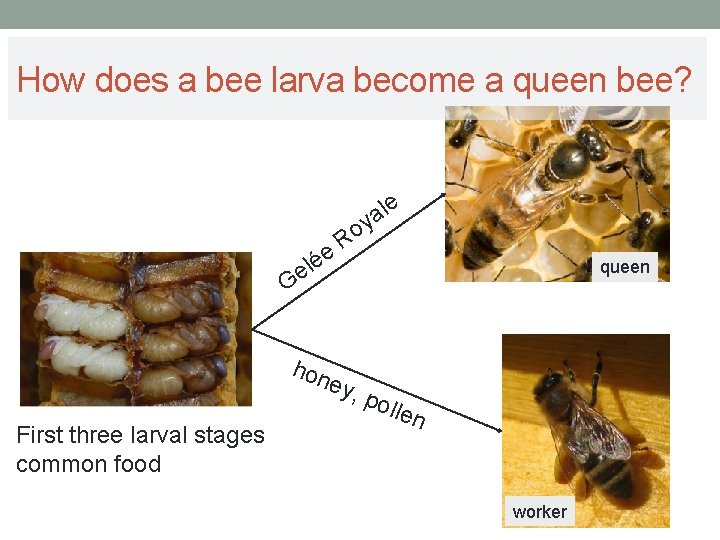
How does a bee larva become a queen bee? le a y e é l e Ro queen G hon ey, First three larval stages common food pol len worker

Characteristics of Gelée Royale Ø Gelée Royale is a mixture of: sugar (10 -23 %), water (60 -70 %), proteins and amino acids (9 -18 %), fats (4 -8 %), thiamine, riboflavin, pyridoxine, niacin, pantothenic acid, biotin, folic acid, sterols, minerals and trace elements. Ø Gelée Royale inhibits the activity of an enzyme called DNMT 3 (DNA methyltransferase 3)

DNA methylation in the bee ØAt least 550 genes are not methylated by the administration of royal jelly (→ Development to queen bee) ØIf the enzyme DNMT 3 is inhibited, queen bees can also be generated from larvae.
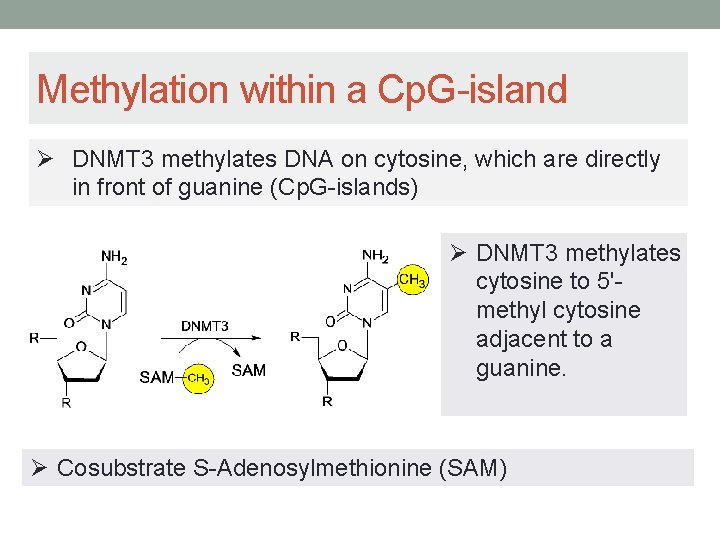
Methylation within a Cp. G-island Ø DNMT 3 methylates DNA on cytosine, which are directly in front of guanine (Cp. G-islands) Ø DNMT 3 methylates cytosine to 5'methyl cytosine adjacent to a guanine. Ø Cosubstrate S-Adenosylmethionine (SAM)
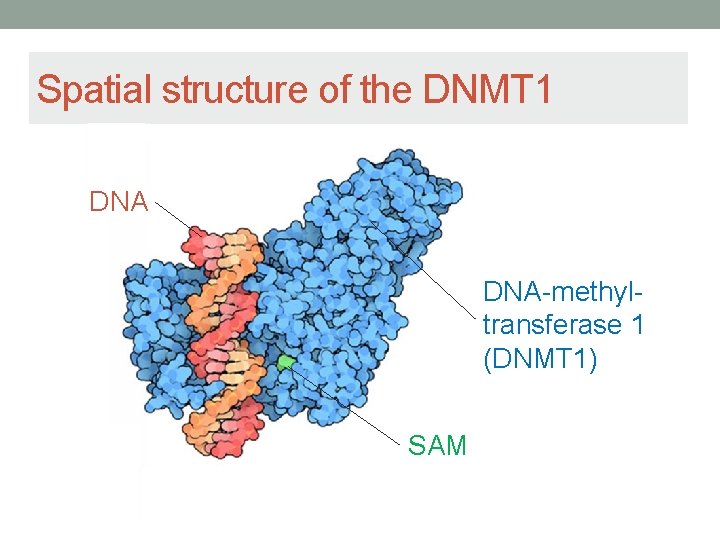
Spatial structure of the DNMT 1 DNA-methyltransferase 1 (DNMT 1) SAM

Restriction Enzymes Recognize a specific palindromic DNA sequence and cut it. Methylated DNA couldn`t be cutted. In nature methylation protects DNA from cleavage, foreign non methylated DNA can be recognized and cutted. Cla. I: First restriction enzyme isolated out of Cladosporium spec.
DNA-Methylation and Restriction of λ-DNA Internship Epigenetics
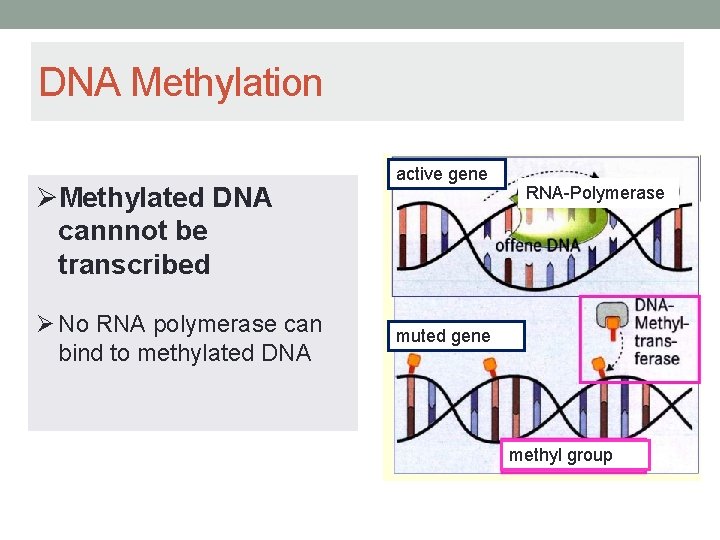
DNA Methylation ØMethylated DNA cannnot be transcribed Ø No RNA polymerase can bind to methylated DNA active gene RNA-Polymerase muted gene methyl group
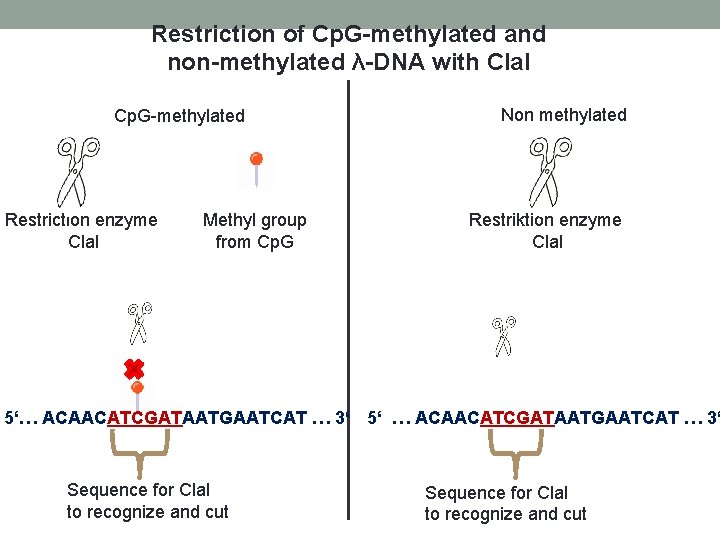
Restriction of Cp. G-methylated and non-methylated λ-DNA with Cla. I Cp. G-methylated Restriction enzyme Cla. I Methyl group from Cp. G Non methylated Restriktion enzyme Cla. I 5‘… ACAACATCGATAATGAATCAT … 3‘ 5‘ … ACAACATCGATAATGAATCAT … 3‘ Sequence for Cla. I to recognize and cut
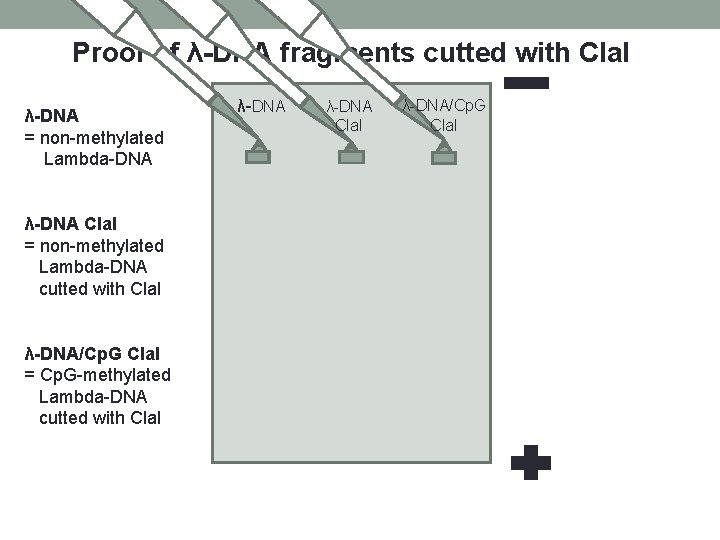
Proof of λ-DNA fragments cutted with Cla. I λ-DNA = non-methylated Lambda-DNA λ-DNA Cla. I = non-methylated Lambda-DNA cutted with Cla. I λ-DNA/Cp. G Cla. I = Cp. G-methylated Lambda-DNA cutted with Cla. I λ-DNA/Cp. G Cla. I

HISTONE ACETYLATION
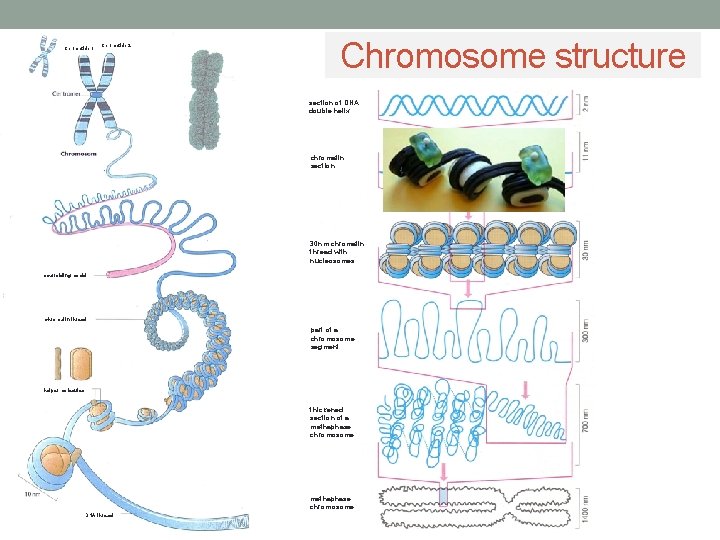
Chromatide 1 Chromatide 2 Chromosome structure section of DNA double helix chromatin section 30 nm chromatin thread with nucleosomes scaffolding model chromatin thread part of a chromosome segment helper molecules thickened section of a methaphase chromosome DNA thread
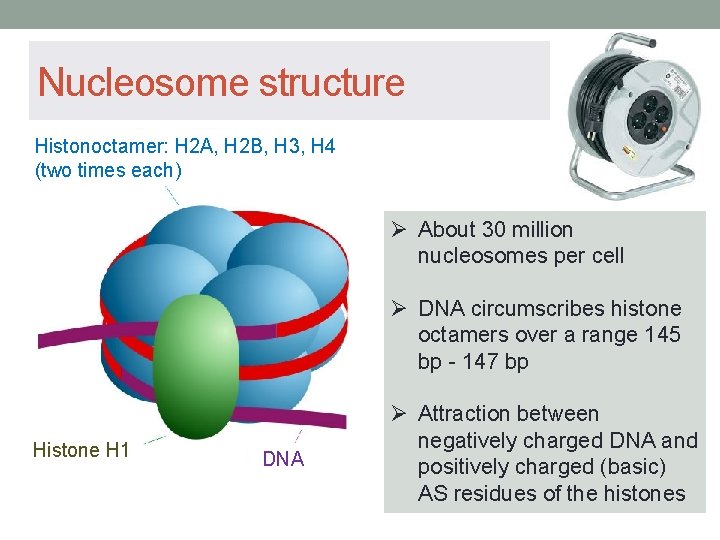
Nucleosome structure Histonoctamer: H 2 A, H 2 B, H 3, H 4 (two times each) Ø About 30 million nucleosomes per cell Ø DNA circumscribes histone octamers over a range 145 bp - 147 bp Histone H 1 DNA Ø Attraction between negatively charged DNA and positively charged (basic) AS residues of the histones
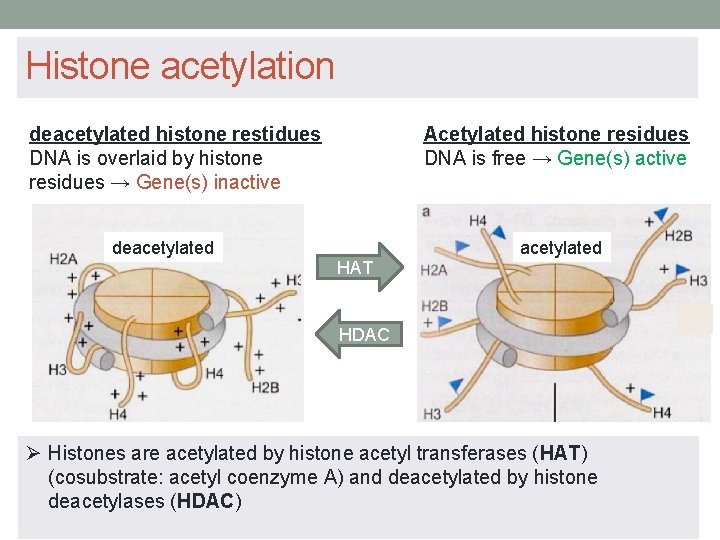
Histone acetylation Acetylated histone residues DNA is free → Gene(s) active deacetylated histone restidues DNA is overlaid by histone residues → Gene(s) inactive deacetylated HAT HDAC Ø Histones are acetylated by histone acetyl transferases (HAT) (cosubstrate: acetyl coenzyme A) and deacetylated by histone deacetylases (HDAC)
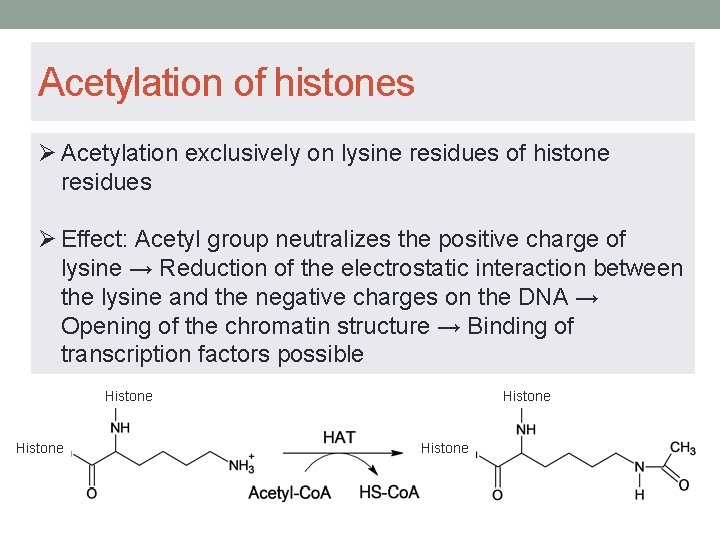
Acetylation of histones Ø Acetylation exclusively on lysine residues of histone residues Ø Effect: Acetyl group neutralizes the positive charge of lysine → Reduction of the electrostatic interaction between the lysine and the negative charges on the DNA → Opening of the chromatin structure → Binding of transcription factors possible Histone

Histone code Ø Acetylation of the histone residues can be formulated using a code active Gene: inactive Gene: H 3 K 14 ac H 3 K 14 Histone 3 Lysine (K) at position 14 in the histone residue acetylated (ac)
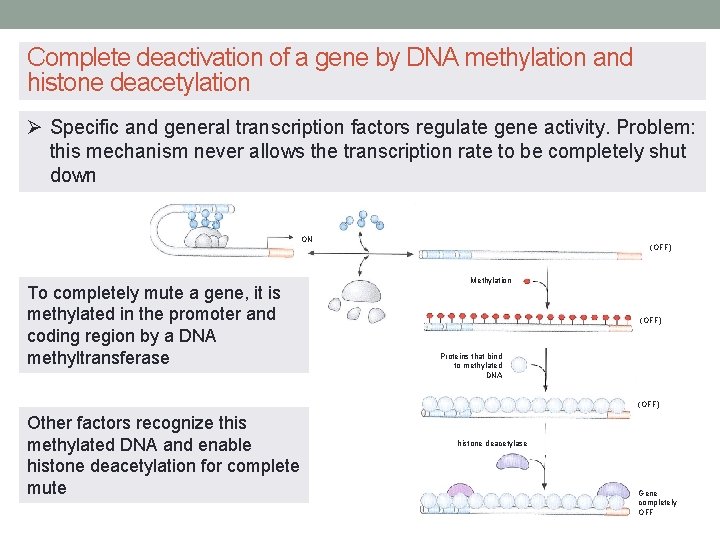
Complete deactivation of a gene by DNA methylation and histone deacetylation Ø Specific and general transcription factors regulate gene activity. Problem: this mechanism never allows the transcription rate to be completely shut down ON To completely mute a gene, it is methylated in the promoter and coding region by a DNA methyltransferase (OFF) Methylation (OFF) Proteins that bind to methylated DNA (OFF) Other factors recognize this methylated DNA and enable histone deacetylation for complete mute histone deacetylase Gene completely OFF

Influence of histone modifications on behaviour Ø Mothers of rats lick their young Ø These boys are more relaxed and less aggressive Ø The leak decondens a region of DNA that codes for the GR gene (GR = glucocorticoid receptor) Ø The receptor is more strongly expressed and the individuals concerned are less susceptible to stress (a mutation in the GR gene is associated in humans with depression and/or post-traumatic stress disorder) http: //learn. genetics. utah. edu/content/epigenetics/rats/

RNA INTERFERENCE BY MICRO-RNAS m. RNA ↓

The human genome Ø the human genome comprises about 20000 genes for proteins Ø 1. 5 % of the genome codes for proteins, 98. 5 % is nc. DNA (non-coding DNA) Ø 70 % of the genome codes for regulatory RNA = nc. RNA (non-coding RNA), located in introns Ø 80000 genes for nc. RNA in humans Ø about 15 different RNA types in humans
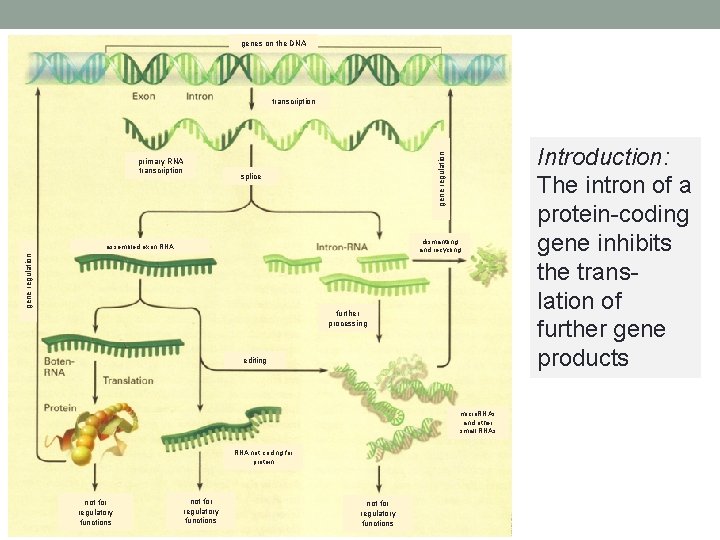
genes on the DNA primary RNA transcription gene regulation transcription splice dismantling and recycling gene regulation assembled exon RNA further processing editing micro. RNAs and other small RNAs RNA not coding for protein not for regulatory functions Introduction: The intron of a protein-coding gene inhibits the translation of further gene products

RNA interference (RNAi) Mechanism for downregulation of translation Medium: small RNA molecules interfering with m. RNA Possibility: Downregulation by micro-RNAs (mi. RNAs)

Micro-RNAs (mi. RNA) Ø Form of regulatory RNA Ø micro. RNA is found in plants, worms, flies, mice and humans Ø Translation of > 30% of human genes regulated by mi. RNAs Ø over 3000 different mi. RNAs in humans
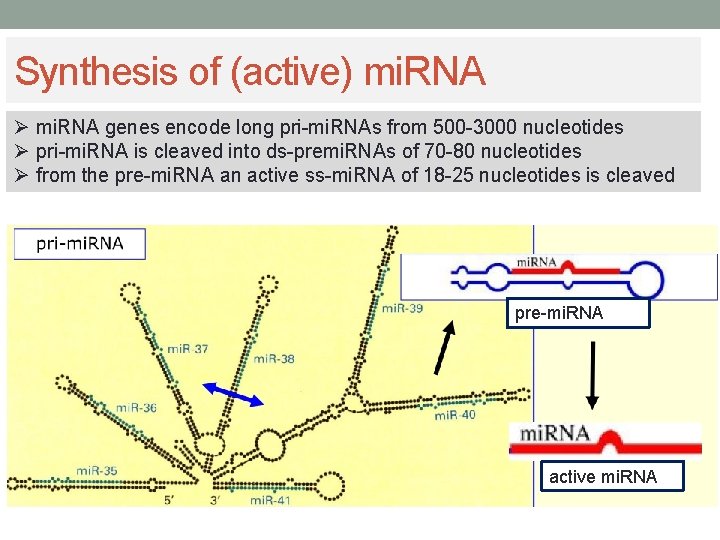
Synthesis of (active) mi. RNA Ø mi. RNA genes encode long pri-mi. RNAs from 500 -3000 nucleotides Ø pri-mi. RNA is cleaved into ds-premi. RNAs of 70 -80 nucleotides Ø from the pre-mi. RNA an active ss-mi. RNA of 18 -25 nucleotides is cleaved pre-mi. RNA active mi. RNA
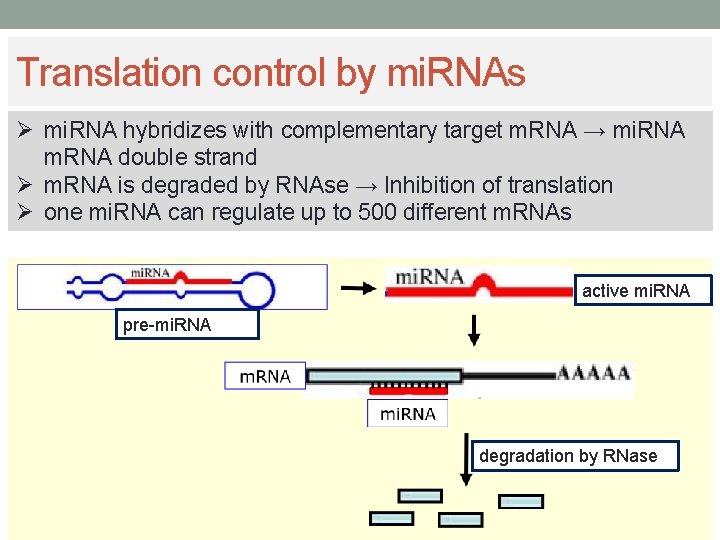
Translation control by mi. RNAs Ø mi. RNA hybridizes with complementary target m. RNA → mi. RNA m. RNA double strand Ø m. RNA is degraded by RNAse → Inhibition of translation Ø one mi. RNA can regulate up to 500 different m. RNAs active mi. RNA pre-mi. RNA degradation by RNase

Sources Lectures by - Prof. Dr. Diethard Baron - Prof. Dr. Thomas Jenuwein Literature:

Thank you for your attention! We make more of it. . .
- Slides: 34