Epidemiology 719 Quantitative methods in genetic epidemiology Bhramar

Epidemiology 719 Quantitative methods in genetic epidemiology Bhramar Mukherjee and Sebastian Zoellner bhramar@umich. edu szoellne@umich. edu

Acknowledgements • • Peter Kraft (HSPH) Ken Rice (UW) Nilanjan Chatterjee (NCI) Stephen Channock (NCI) Lu Wang (UM) Nan Laird (HSPH) Goncalo Abecasis (UM)

A brave new world Course Overview


Reverse Effects

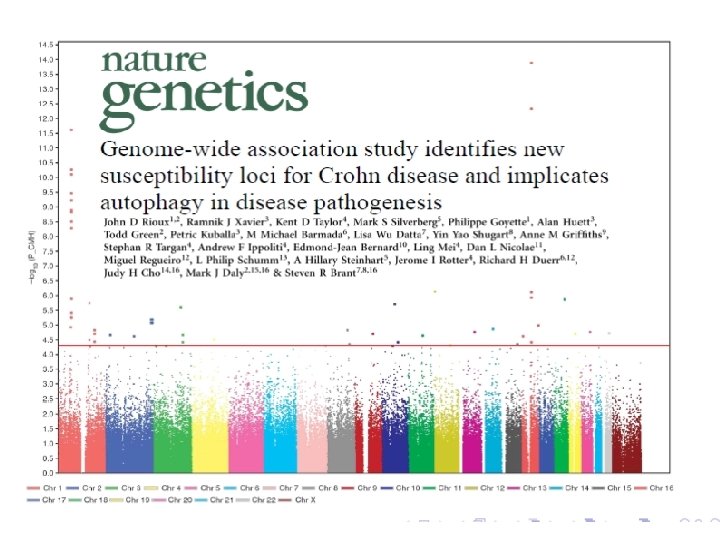

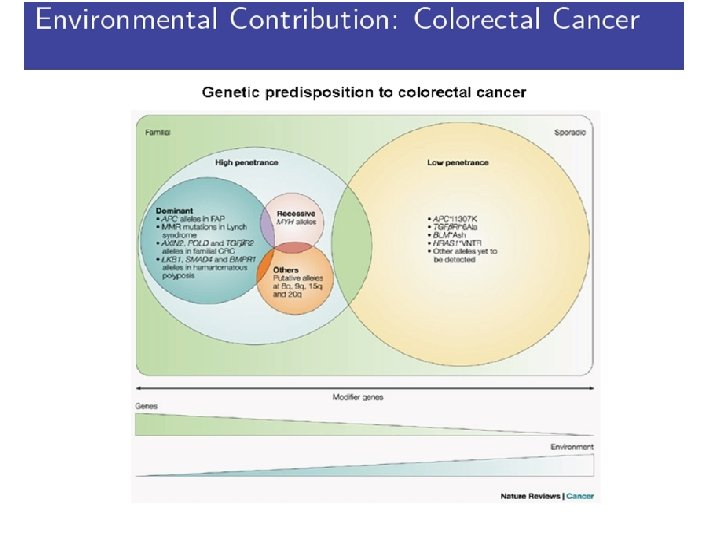










Central Course Theme Genetic Association and Gene-Environment Interaction

Course Advice for You:

Assigned Paper 1

Assigned Paper 1 • GWAS of Age-related macular degeneration • Initial GWAS identified four loci explaining one-half of the heritability. Appreciable predictive power. • Additional GWAS to explain remaining heritability. Combined scan vs replication. Meta-Analysis.

Assigned Paper 2

Assigned Paper 2 • Collaborative Association Study of Psoriasis • Examined ~1, 500 cases / ~1, 500 controls at ~500, 000 SNPs • • Examined 20 promising SNPs in extra ~5, 000 cases / ~5, 000 controls • Outcome: 7 regions of confirmed association with psoriasis

Assigned Paper 3

Assigned Paper 3 • Meta-analysis of colorectal cancer (COGENT study). • A thorough evaluation of ten confirmed loci for colorectal cancer. Very detailed. Supplementary material also available online. • Interesting combination of various study design.

Tests for Association

Basic principle of GWAS

Depends on study design • Case-control study • Family-based study: case-parent triad, case-sib pairs being popular choices • Longitudinal Cohort Study • Looking at a secondary outcome under case-control sampling


The GWAS Mantra!

Primary Analysis • Single marker association tests • Genetic susceptibility model - Dominant, recessive, co-dominant • Which test to use • Multiple testing correction









Case-Control Study: Standard Analysis


Pros and Cons • Simple, Complete. • Robust to misspecification of the true dominance pattern • Less powerful. • Unreliable for sparse table


Pros and Cons • Test statistic has single df, so more powerful. • Simple to report. • Not robust to true mode of dominance • Does not present entire information in the data.

Armitage’s trend test • Test linear trend in log(OR) with # A allele • Test statistic still has single d. f. • Simplicity, use information from the 2 x 3 array • More robust than 2 x 2 tests, but less robust than the 2 d. f. test.

Allelic test • Previous tests were based on genotype • Can also treat allele as the unit of observation. • You have doubled the sample size!!

But… • Serious impact on Type 1 error under departures from HWE • Interpretation becomes trickier.
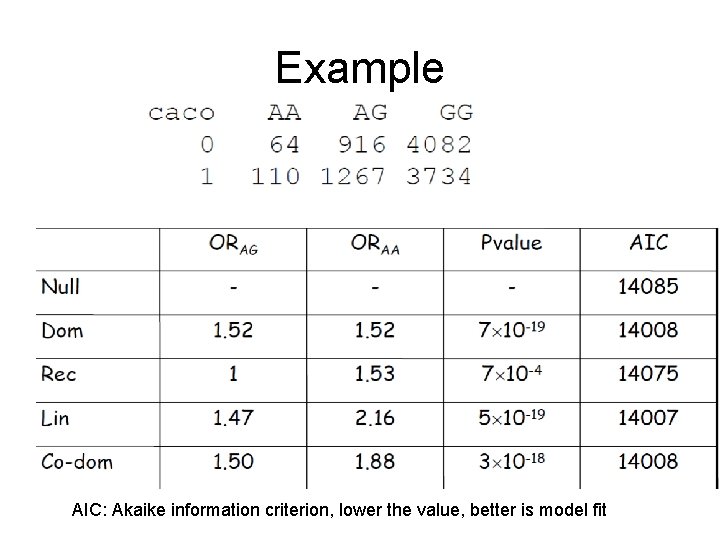
Example AIC: Akaike information criterion, lower the value, better is model fit

Using logistic regression • • Trick: Just code genotype differently Dominant: G=1 if AA or Aa, 0 otherwise Recessive: G=1 if AA, 0 otherwise Trend: G=# A alleles, thus G=2 if AA, =1 if Aa and 0 if aa • Two df test: Create two dummy variables: G 1=1 if Aa and 0 otherwise G 2=1 if AA and 0 otherwise Perform likelihood ratio test of full (G 1 and G 2) vs reduced model (No G 1, G 2). • Adjust for other variables, fit a multivariate model
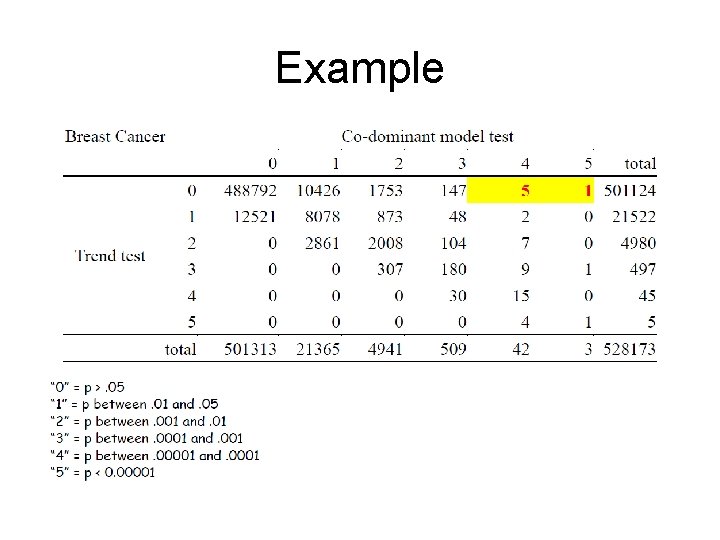
Example

Flip Flops • Under a co-dominant model you see a non -monotone trend, i. e. , • OR(Aa)<1 and OR(AA)>1 • You will likely miss these under the trend test.

Alternative tests • Use alternative maximal test statistic • Calculate dominant, recessive, trend, codominabt: take maximum test statistic • Use permutation to get right P-value Caveats • Resist temptations of going on to a fishing expedition. “MOST SIGNIFICANT CODING” • Mode of inheritance models were developed for simple mendelian disease with near-complete penetrance, much more difficult to believe for complex diseases.

Reliable Test • Co-dominant is model free • Not much loss of power unless AA (homozygous carriers) are very rare. • Log additive is what is reported most of the time to risk some false positives but enhance power.

QQ Plots • Nice visual tools for checking association and systematic biases • Plot observed (-log 10)P-values versus expected under the global null (i. e. quantiles draw from U[0, 1]) • Since vast majority (>99%) of tested markers are not associated with the trait, plot should fall along y=x line (if we are lucky we will see a few departures in the tail). • Departures could be due to stratification, cryptic relatedness, differential genotyping error, incorrect test.
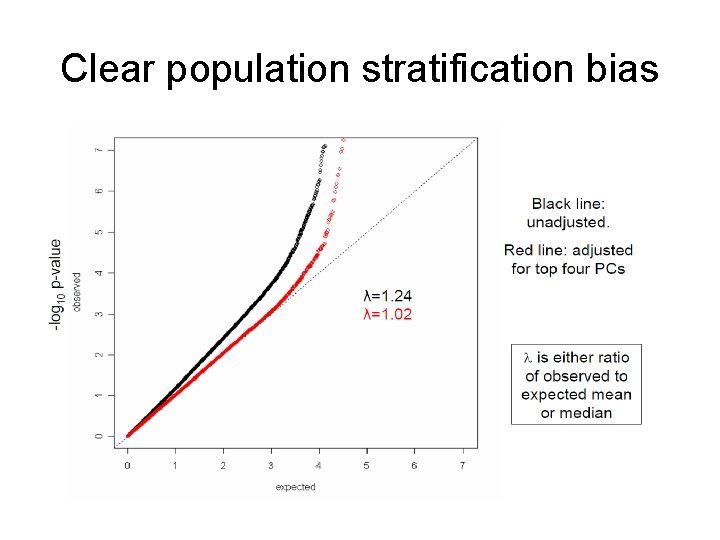
Clear population stratification bias

Family Based Studies • You heard about population stratification. • Solutions: -Match on self-reported ethnicity -Adjust for Principal components extracted from markers. -Use family based controls Case-sib (conditional logistic regression) Case-parent (TDT, FBAT) Nuclear Families Extended Pedigrees

Why use families • Robust tests. • Detect genotyping error with inheritance impossibilities. (mother and father AA, offspring Aa). • Do not have to think about selection of “good” controls.

Why not use families • Case-control typically more powerful unless the disease is very rare. • Not just logistic regression or trend chisquared as analysis tools. • Much harder to recruit (depending on the disease : late onset or childhood disorder).

Hypotheses • Case-control design • Family-based designs have no power to detect association unless linkage is present. • When testing for association in family-based design, HA is always: both linkage and association is present between the marker and disease susceptibility locus (DSL) underlying the trait.

The null hypotheses

Mendelian Transmission

Transmission Disequilibrium Test • Spielman et al, AJHG, 1998 • H 0: Association but no linkage

A simple statistical test • No variation in diagonal elements (homozygote parents have no uncertainty in determining the conditional genotype distribution of the offspring). • Under the null (i. e. , Mendelian transmission) x| x+y~Binomial (n=x+y, p=1/2) • Similarly, like Mc. Nemar’s test:

TDT Example Test statistic:

FBAT : Family based association tests • Extending TDT beyond case-parent trios • Test statistic for FBAT mimics a natural covariance function between trait and genotype. • i: family j: individual Sum over all i and j • T: Trait (centered) X: Coded Genotype • S: parental genotype or a sufficient statistic for parental genotype

Details • The E(X|S) is calculated under Mendelian transmission. • X-E(X|S): residual of the transmission of parental genotype to offspring. • You basically assess whethere is any association between the trait and this genotype residual.

Test statistic • Under all three nulls, E(U)=0. Test Statistic: • Note all expectation and variance are on X, conditional on parental genotype and trait T.


Another Tool: Conditional Likelihood • Used in case-sib studies, in general in matched studies. Strata: Family/pair. • Breslow et al (1978) first proposed this tool for matched case-control data. • R function clogit does this. Underlying codes use survival model as there is connection with partial likelihood from Cox’s proportional hazard model.



• For case-sib studies, sib is the control and the genotype is the exposure. The contribution of a given pair to the conditional likelihood is: • exp[β Genotype(case)]+exp[β Genotype(control)] -Obtain variance-covariance of conditional MLE using inverse Fisher information.

Summary • Different tests for association in casecontrol studies: 2 by 3 table and logistic regression. • Family based studies: tests and hypotheses. • Study design choices, power, recruitment.
- Slides: 74