ENZYMES CLASSIFICATION STRUCTURE Enzymes catalysts of biological reactions
ENZYMES: CLASSIFICATION, STRUCTURE
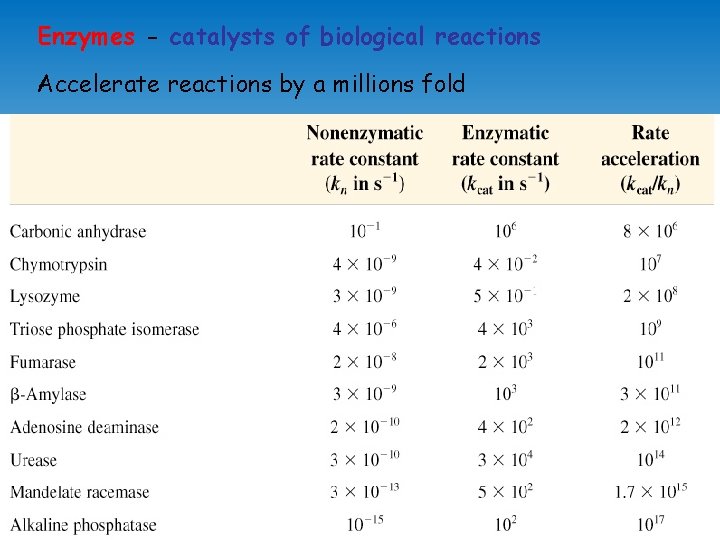
Enzymes - catalysts of biological reactions Accelerate reactions by a millions fold

Common features for enzymes and inorganic catalysts: 1. Catalyze only thermodynamically possible reactions 2. Are not used or changed during the reaction. 3. Don’t change the position of equilibrium and direction of the reaction 4. Usually act by forming a transient complex with the reactant, thus stabilizing the transition state

Specific features of enzymes: 1. Accelerate reactions in much higher degree than inorganic catalysts 2. Specificity of action 3. Sensitivity to temperature 4. Sensitivity to p. H
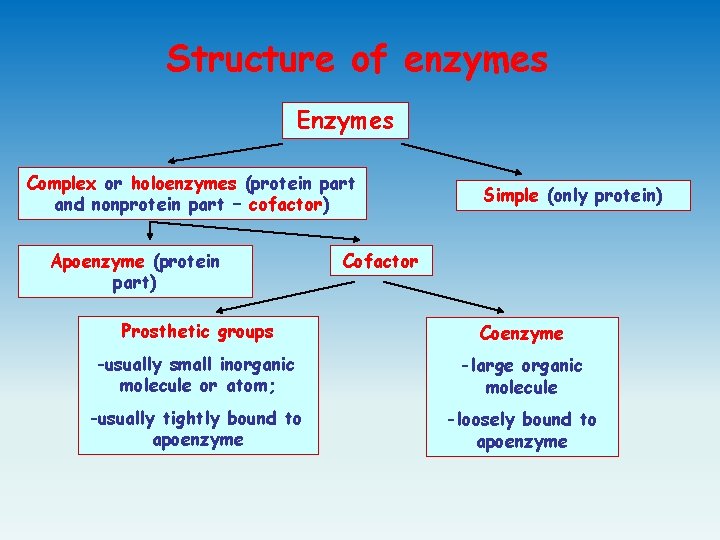
Structure of enzymes Enzymes Complex or holoenzymes (protein part and nonprotein part – cofactor) Apoenzyme (protein part) Simple (only protein) Cofactor Prosthetic groups Coenzyme -usually small inorganic molecule or atom; -large organic molecule -usually tightly bound to apoenzyme -loosely bound to apoenzyme
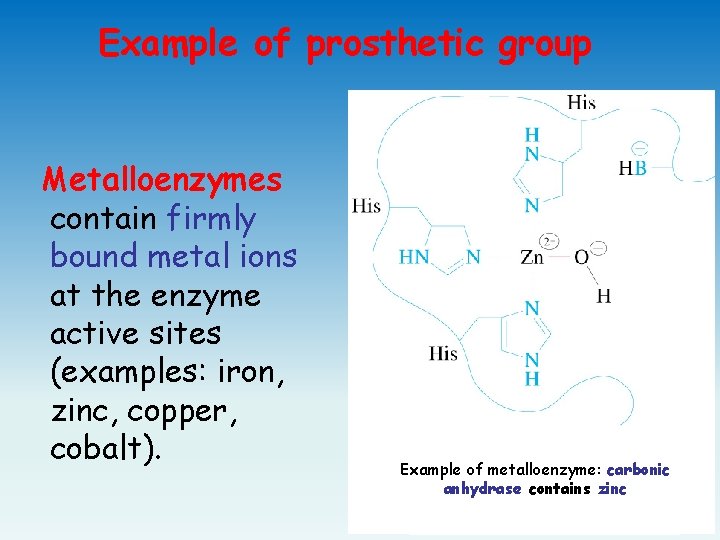
Example of prosthetic group Metalloenzymes contain firmly bound metal ions at the enzyme active sites (examples: iron, zinc, copper, cobalt). Example of metalloenzyme: carbonic anhydrase contains zinc

Coenzymes • Coenzymes act as group-transfer reagents • Hydrogen, electrons, or groups of atoms can be transferred Coenzyme classification (1) Metabolite coenzymes - synthesized from common metabolites (2) Vitamin-derived coenzymes - derivatives of vitamins Vitamins cannot be synthesized by mammals, but must be obtained as nutrients
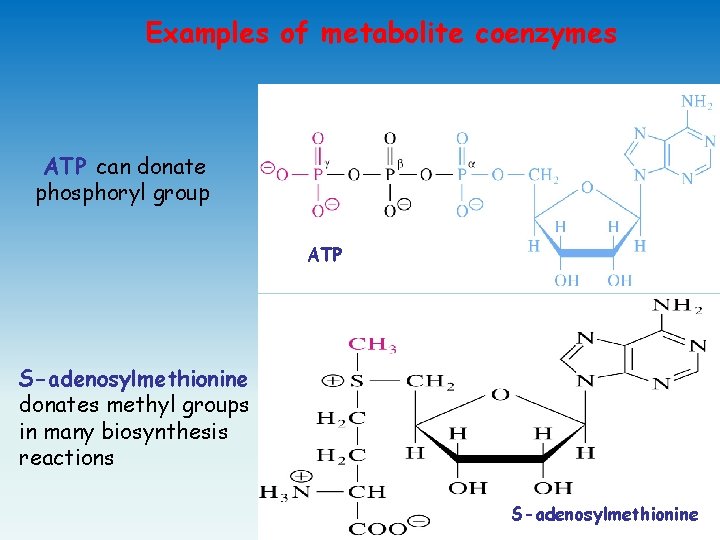
Examples of metabolite coenzymes ATP can donate phosphoryl group ATP S-adenosylmethionine donates methyl groups in many biosynthesis reactions S-adenosylmethionine
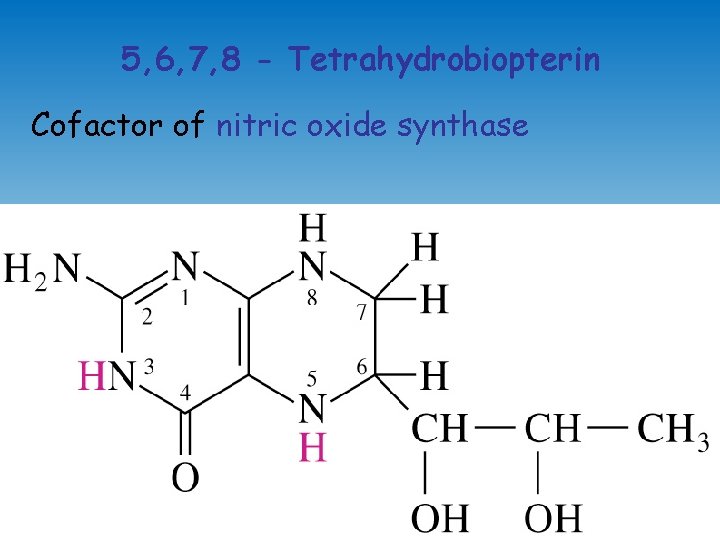
5, 6, 7, 8 - Tetrahydrobiopterin Cofactor of nitric oxide synthase

Vitamin-Derived Coenzymes • Vitamins are required for coenzyme synthesis and must be obtained from nutrients • Most vitamins must be enzymatically transformed to the coenzyme • Deficit of vitamin and as result correspondent coenzyme results in the disease
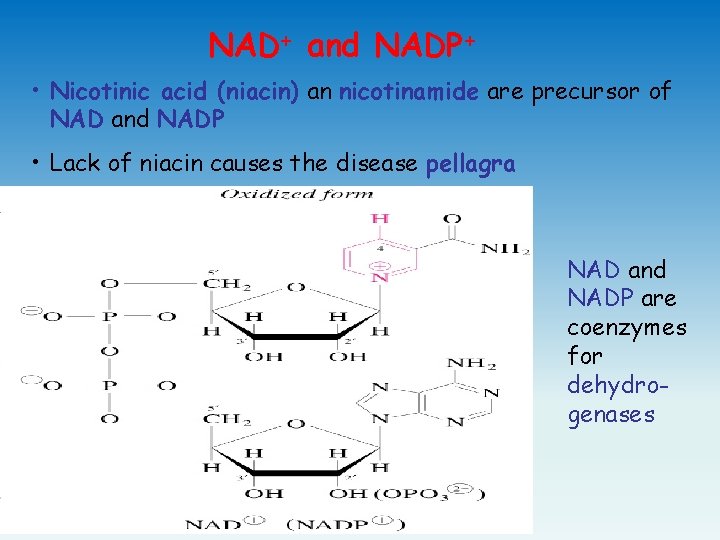
NAD+ and NADP+ • Nicotinic acid (niacin) an nicotinamide are precursor of NAD and NADP • Lack of niacin causes the disease pellagra NAD and NADP are coenzymes for dehydrogenases
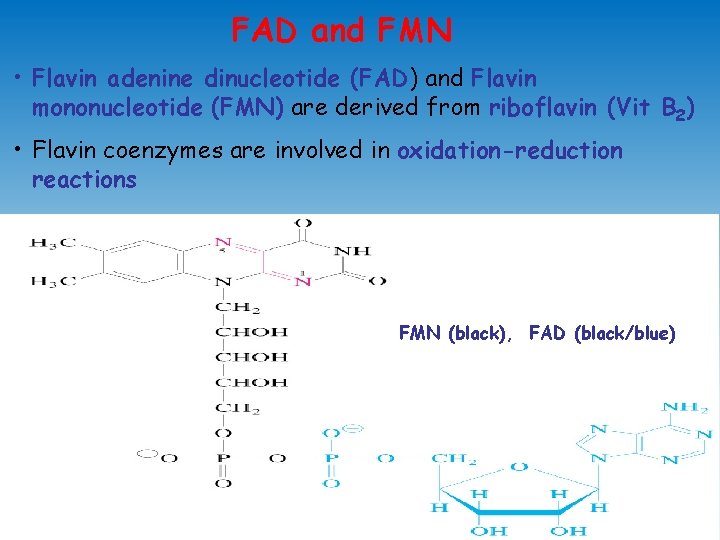
FAD and FMN • Flavin adenine dinucleotide (FAD) and Flavin mononucleotide (FMN) are derived from riboflavin (Vit B 2) • Flavin coenzymes are involved in oxidation-reduction reactions FMN (black), FAD (black/blue)
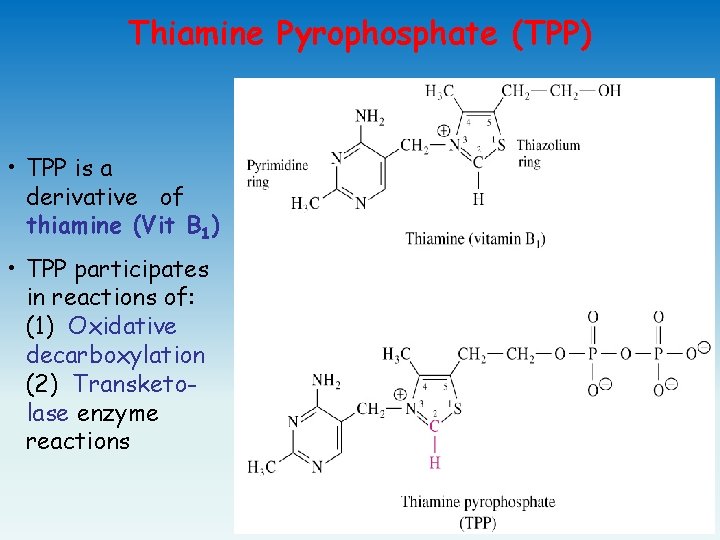
Thiamine Pyrophosphate (TPP) • TPP is a derivative of thiamine (Vit B 1) • TPP participates in reactions of: (1) Oxidative decarboxylation (2) Transketolase enzyme reactions
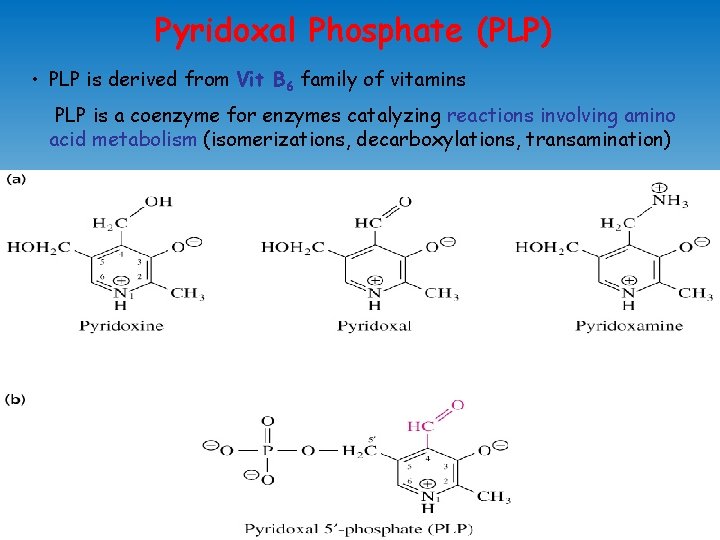
Pyridoxal Phosphate (PLP) • PLP is derived from Vit B 6 family of vitamins PLP is a coenzyme for enzymes catalyzing reactions involving amino acid metabolism (isomerizations, decarboxylations, transamination)

Enzymes active sites Substrate usually is relatively small molecule Enzyme is large protein molecule Therefore substrate binds to specific area on the enzyme Active site – specific region in the enzyme to which substrate molecule is bound

Characteristics of active sites § Specificity (absolute, relative (group), stereospecificity) § Small three dimensional region of the protein. Substrate interacts with only three to five amino acid residues. Residues can be far apart in sequence § Binds substrates through multiple weak interactions (noncovalent bonds) § There are contact and catalytic regions in the active site
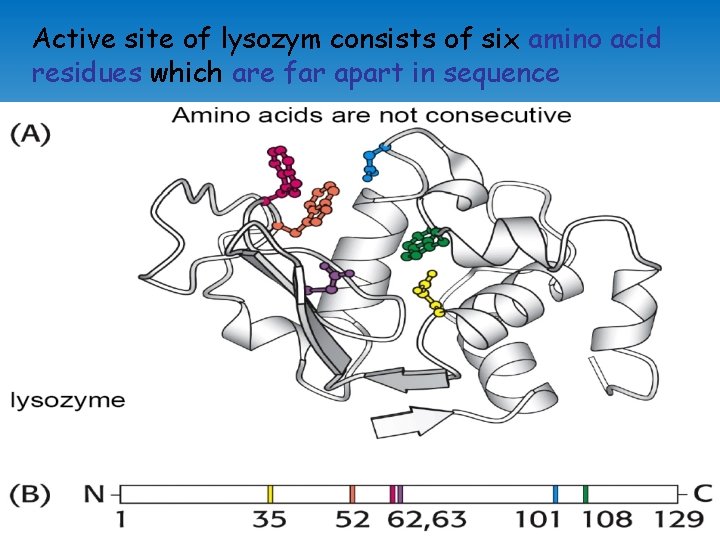
Active site of lysozym consists of six amino acid residues which are far apart in sequence
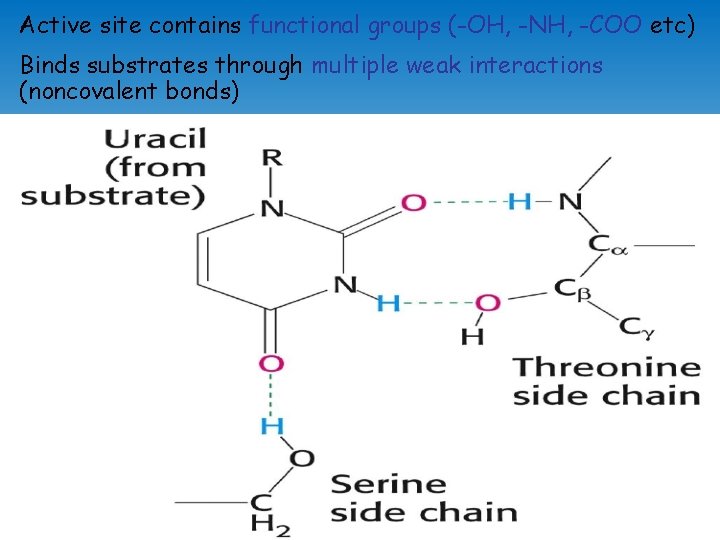
Active site contains functional groups (-OH, -NH, -COO etc) Binds substrates through multiple weak interactions (noncovalent bonds)
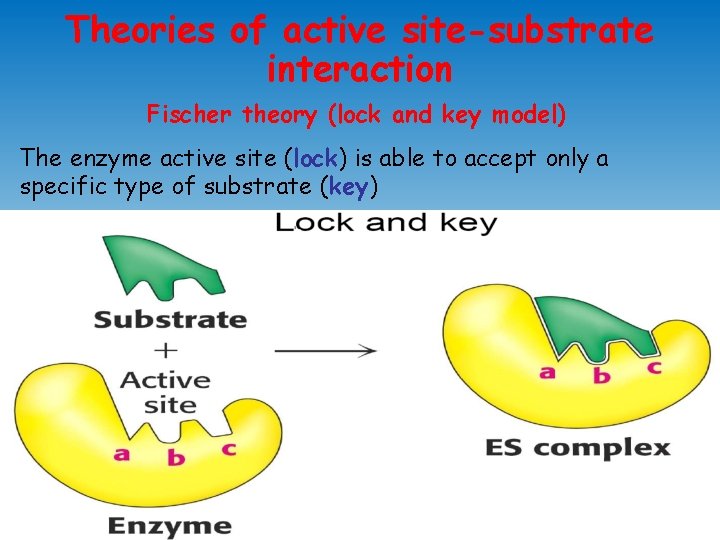
Theories of active site-substrate interaction Fischer theory (lock and key model) The enzyme active site (lock) is able to accept only a specific type of substrate (key)
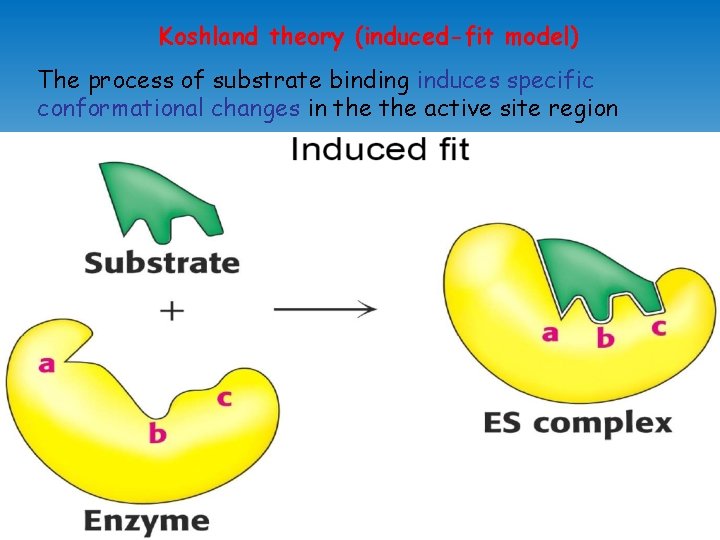
Koshland theory (induced-fit model) The process of substrate binding induces specific conformational changes in the active site region

Properties of Enzymes Specificity of enzymes 1. Absolute – one enzyme acts only on one substrate (example: urease decomposes only urea; arginase splits only arginine) 2. Relative – one enzyme acts on different substrates which have the same bond type (example: pepsin splits different proteins) 3. Stereospecificity – some enzymes can catalyze the transformation only substrates which are in certain geometrical configuration, cis- or trans-
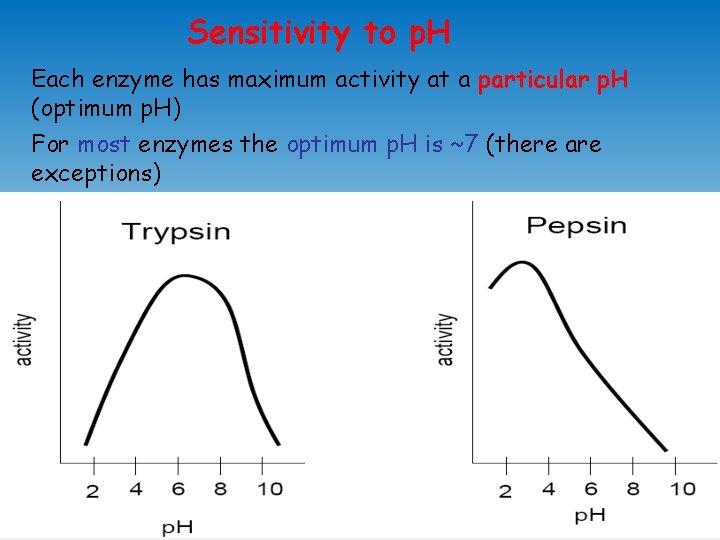
Sensitivity to p. H Each enzyme has maximum activity at a particular p. H (optimum p. H) For most enzymes the optimum p. H is ~7 (there are exceptions)
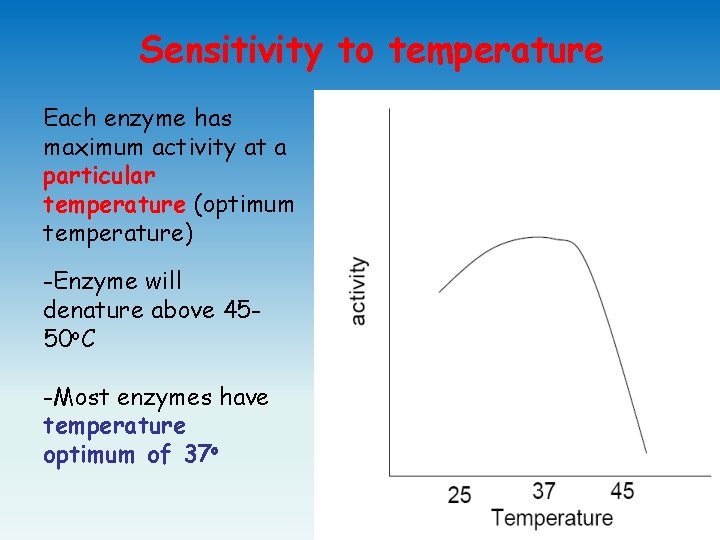
Sensitivity to temperature Each enzyme has maximum activity at a particular temperature (optimum temperature) -Enzyme will denature above 4550 o. C -Most enzymes have temperature optimum of 37 o

Naming of Enzymes Common names are formed by adding the suffix –ase to the name of substrate Example: - tyrosinase catalyzes oxidation of tyrosine; - cellulase catalyzes the hydrolysis of cellulose Common names don’t describe the chemistry of the reaction Trivial names Example: pepsin, catalase, trypsin. Don’t give information about the substrate, product or chemistry of the reaction

Principle of the international classification All enzymes are classified into six categories according to the type of reaction they catalyze Each enzyme has an official international name ending in –ase Each enzyme has classification number consisting of four digits: EC: 2. 3. 4. 2 First digit refers to a class of enzyme, second to a subclass, third – to a subsubclass, and fourth means the ordinal number of enzyme in subsubclass

The Six Classes of Enzymes 1. Oxidoreductases • Catalyze oxidation-reduction reactions - oxidases - peroxidases - dehydrogenases
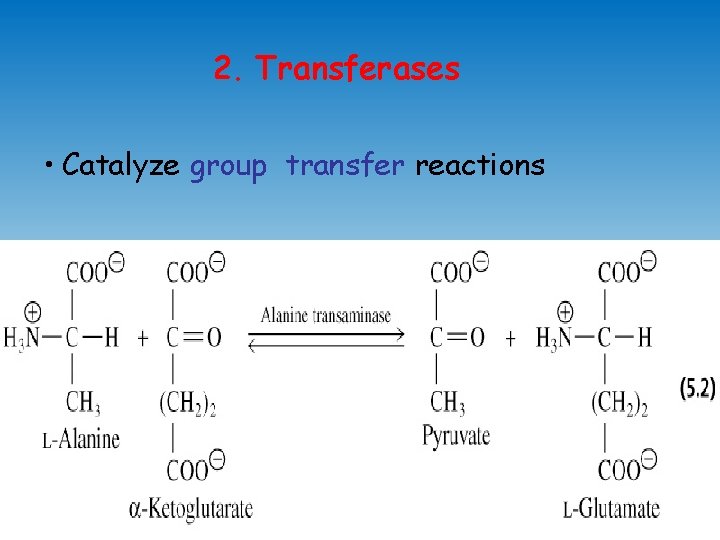
2. Transferases • Catalyze group transfer reactions

3. Hydrolases • Catalyze hydrolysis reactions where water is the acceptor of the transferred group - esterases - peptidases - glycosidases
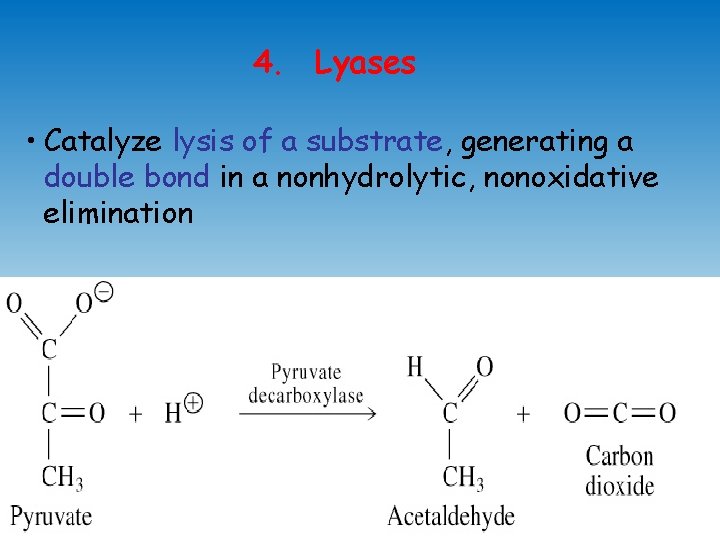
4. Lyases • Catalyze lysis of a substrate, generating a double bond in a nonhydrolytic, nonoxidative elimination
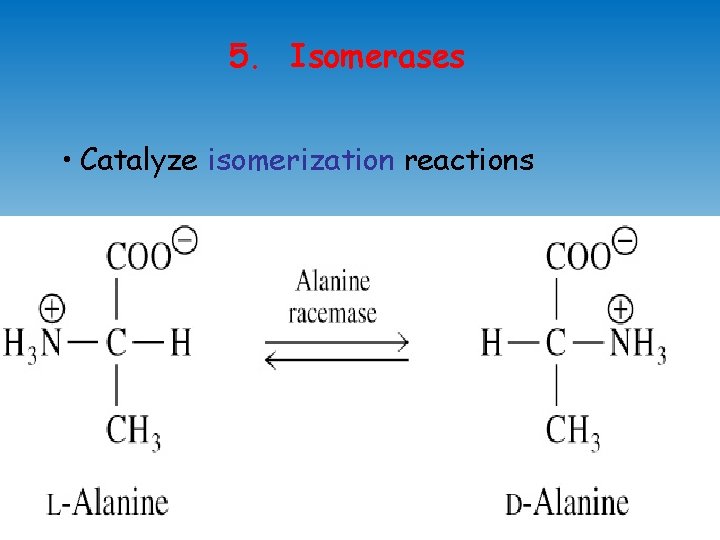
5. Isomerases • Catalyze isomerization reactions
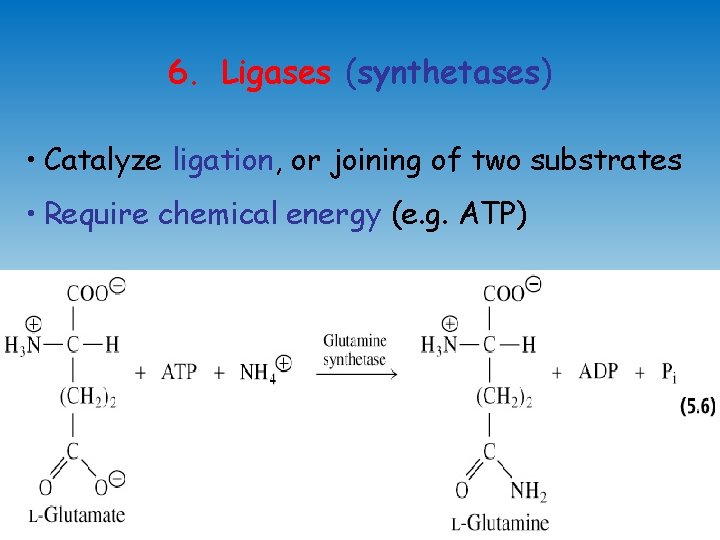
6. Ligases (synthetases) • Catalyze ligation, or joining of two substrates • Require chemical energy (e. g. ATP)
- Slides: 31