Enzyme Mechanisms and Regulation Andy Howard Introductory Biochemistry

Enzyme Mechanisms and Regulation Andy Howard Introductory Biochemistry, Fall 2010 Wednesday 22 September 2010 Biochem: Mechanisms, Regulation 1 09/22/2010
How do enzymes reduce activation energies? We want to understand what is really happening chemically when an enzyme does its job. n We’d also like to know how biochemists probe these systems. n 09/22/2010 Biochem: Mechanisms, Regulation 2

Mechanism & Regulation Topics n Mechanisms n n n Binding mode catalysis Chemical catalysis Examples Intermediates Applications of Mechanisms n n n Serine proteases Other proteases Lysozyme 09/22/2010 n Regulation n n Thermodynamics Enzyme availability Allostery, revisited Regulation of Globins Biochem: Mechanisms, Regulation 3
How do enzymes reduce activation energies? n We can illustrate mechanistic principles by looking at specific examples; we can also recognize enzyme regulation when we see it. 09/22/2010 Biochem: Mechanisms, Regulation 4

Examining enzyme mechanisms will help us understand catalysis n n Examining general principles of catalytic activity and looking at specific cases will facilitate our appreciation of all enzymes. We can distinguish between bindingmode mechanisms and chemical mechanisms; we’ll look at both 09/22/2010 Biochem: Mechanisms, Regulation 5

Binding modes: proximity n n We describe enzymatic mechanisms in terms of the binding modes of the substrates (or, more properly, the transition-state species) to the enzyme. One of these involves the proximity effect, in which two (or more) substrates are directed down potential-energy gradients to positions where they are close to one another. Thus the enzyme is able to defeat the entropic difficulty of bringing substrates together. 09/22/2010 Biochem: Mechanisms, Regulation William Jencks 6
Binding modes: efficient transition-state binding n n Transition state fits even better (geometrically and electrostatically) in the active site than the substrate would. This improved fit lowers the energy of the transition-state system relative to the substrate. Best competitive inhibitors of an enzyme are those that resemble the transition state rather than the substrate or product. 09/22/2010 Biochem: Mechanisms, Regulation 7

Diffusion-controlled reactions n n Some enzymes are so efficient that the limiting factor in completion of the reaction is diffusion of the substrates into the active site: These are diffusion-controlled reactions. Ultra-high turnover rates: kcat ~ 109 s-1. We can describe kcat / Km as catalytic efficiency of an enzyme. A diffusioncontrolled reaction will have a catalytic efficiency on the order of 108 M-1 s-1. 09/22/2010 Biochem: Mechanisms, Regulation 8
Induced fit n n n Refinement on original Emil Fischer lock-and-key notion: both the substrate (or transitionstate) and the enzyme have flexibility Binding induces conformational changes 09/22/2010 Biochem: Mechanisms, Regulation 9

Ionic reactions n n Define them as reactions that involve charged, or at least polar, intermediates Typically 2 reactants n n Electron rich (nucleophilic) reactant Electron poor (electrophilic) reactant Conventional to describe reaction as attack of nucleophile on electrophile Drawn with nucleophile donating electron(s) to electrophile 09/22/2010 Biochem: Mechanisms, Regulation 10
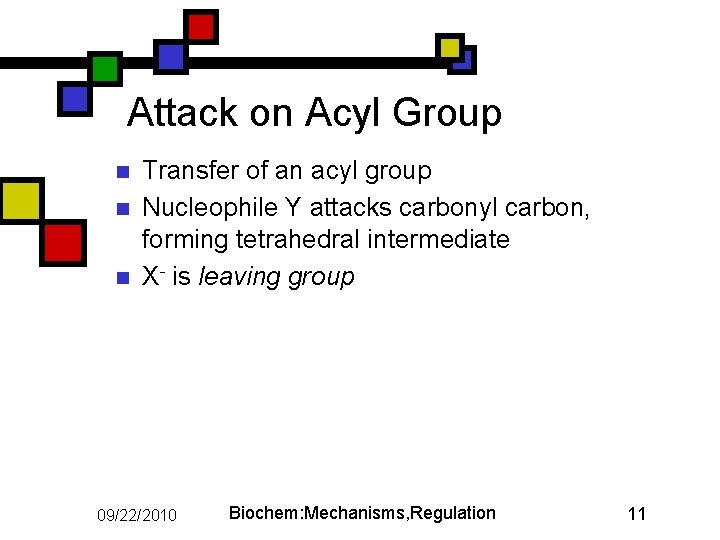
Attack on Acyl Group n n n Transfer of an acyl group Nucleophile Y attacks carbonyl carbon, forming tetrahedral intermediate X- is leaving group 09/22/2010 Biochem: Mechanisms, Regulation 11

Direct Displacement n n Attacking group adds to face of atom opposite to leaving group (scheme 6. 2) Transition state has five ligands; inherently less stable than scheme 6. 1 09/22/2010 Biochem: Mechanisms, Regulation 12

Cleavage Reactions n Both electrons stay with one atom n n n Covalent bond produces carbanion: R 3—C—H R 3—C: - + H+ Covalent bond produces carbocation: R 3—C—H R 3—C+ + : H- One electron stays with each product n Both end up as radicals n R 1 O—OR 2 R 1 O • + • OR 2 Radicals are highly reactive— some more than others n 09/22/2010 Biochem: Mechanisms, Regulation 13

Oxidation-Reduction Reactions n n Commonplace in biochemistry: EC 1 Oxidation is a loss of electrons Reduction is the gain of electrons In practice, often: n n n oxidation is decrease in # of C-H bonds; reduction is increase in # of C-H bonds Mnemonic: OIL RIG n n Oxidation is loss of electrons Reduction is gain of electrons 09/22/2010 Biochem: Mechanisms, Regulation 14
Redox, continued n n n Intermediate electron acceptors and donors are organic moieties or metals Ultimate electron acceptor in aerobic organisms is usually dioxygen (O 2) Anaerobic organisms usually employ other electron acceptors 09/22/2010 Biochem: Mechanisms, Regulation 15
Biological redox reactions n n n n Generally 2 -electron transformations Often involve alcohols, aldehydes, ketones, carboxylic acids, C=C bonds: R 1 R 2 CH-OH + X R 1 R 2 C=O + XH 2 R 1 HC=O + X + OH- R 1 COO- + XH 2 X is usually NAD, NADP, FAD, FMN A few biological redox systems involve metal ions or Fe-S complexes Usually reduced compounds are higher-energy than the corresponding oxidized compounds 09/22/2010 Biochem: Mechanisms, Regulation 16
Examples illustrating transition state stabilization n n Numerous enzymes act by providing stabilization of a transition state or an intermediate Giveaway is extremely effective competitive inhibitors that resemble the transition state that is being stabilized 09/22/2010 Biochem: Mechanisms, Regulation 17

Adenosine deaminase with transition-state analog n n Transition-state analog: Ki~10 -8 * substrate Km Wilson et al (1991) Science 252: 1278 09/22/2010 Biochem: Mechanisms, Regulation 18
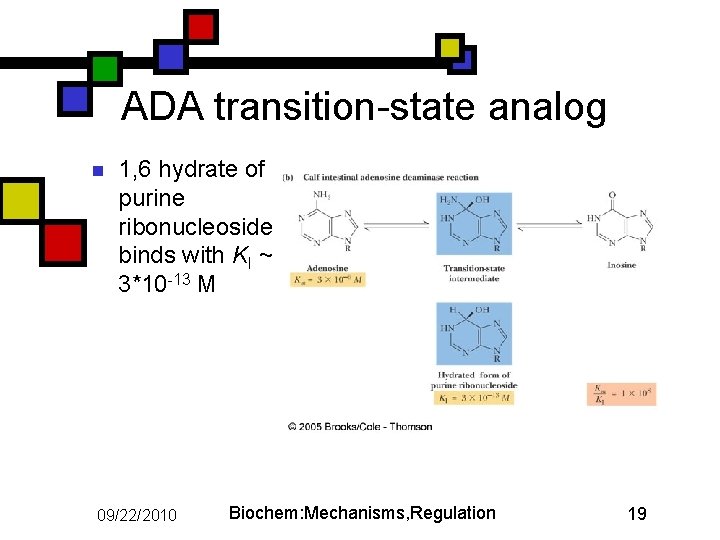
ADA transition-state analog n 1, 6 hydrate of purine ribonucleoside binds with KI ~ 3*10 -13 M 09/22/2010 Biochem: Mechanisms, Regulation 19
Example of induced fit: hexokinase n n n Glucose + ATP Glucose-6 -P + ADP Risk: unproductive reaction with water Enzyme exists in open & closed forms Glucose induces conversion to closed form; water can’t do that Energy expended moving to closed form 09/22/2010 Biochem: Mechanisms, Regulation 20
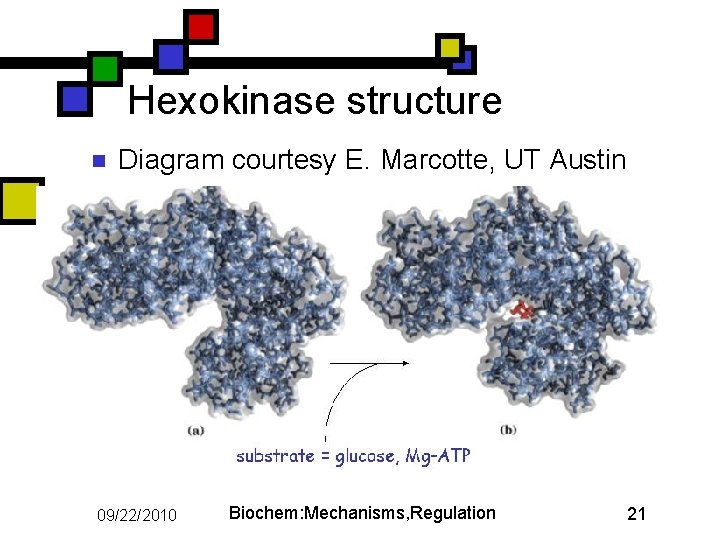
Hexokinase structure n Diagram courtesy E. Marcotte, UT Austin 09/22/2010 Biochem: Mechanisms, Regulation 21
Tight binding of ionic intermediates n n Quasi-stable ionic species strongly bound by ion-pair and H-bond interactions Similar to notion that transition states are the most tightly bound species, but these are more stable 09/22/2010 Biochem: Mechanisms, Regulation 22
Serine protease mechanism n n n Only detailed mechanism that we’ll ask you to memorize One of the first to be elucidated Well studied structurally Illustrates many other mechanisms Instance of convergent and divergent evolution 09/22/2010 Biochem: Mechanisms, Regulation 23
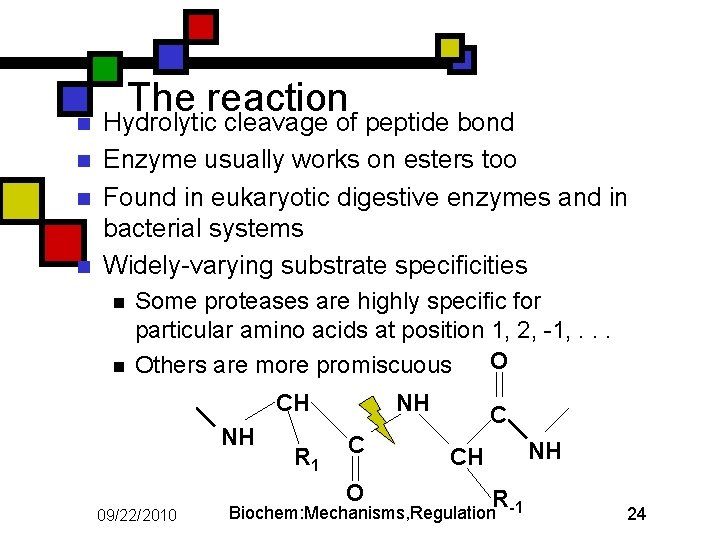
n n The reaction Hydrolytic cleavage of peptide bond Enzyme usually works on esters too Found in eukaryotic digestive enzymes and in bacterial systems Widely-varying substrate specificities n n Some proteases are highly specific for particular amino acids at position 1, 2, -1, . . . O Others are more promiscuous CH NH R 1 NH C O 09/22/2010 C CH R Biochem: Mechanisms, Regulation -1 NH 24
Mechanism n n Active-site serine —OH … Without neighboring amino acids, it’s fairly unreactive becomes powerful nucleophile because OH proton lies near unprotonated N of His This N can abstract the hydrogen at nearneutral p. H Resulting + charge on His is stabilized by its proximity to a nearby carboxylate group on an aspartate side-chain. 09/22/2010 Biochem: Mechanisms, Regulation 25
Catalytic triad n The catalytic triad of asp, his, and ser is found in an approximately linear arrangement in all the serine proteases, all the way from non-specific, secreted bacterial proteases to highly regulated and highly specific mammalian proteases. 09/22/2010 Biochem: Mechanisms, Regulation 26
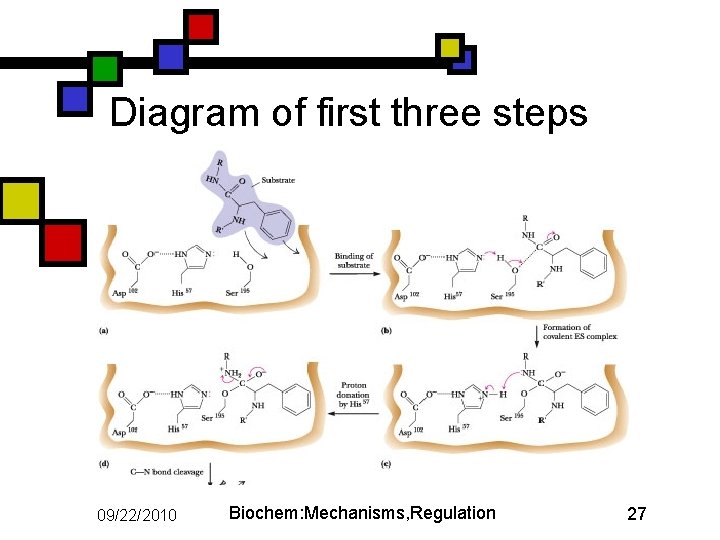
Diagram of first three steps 09/22/2010 Biochem: Mechanisms, Regulation 27
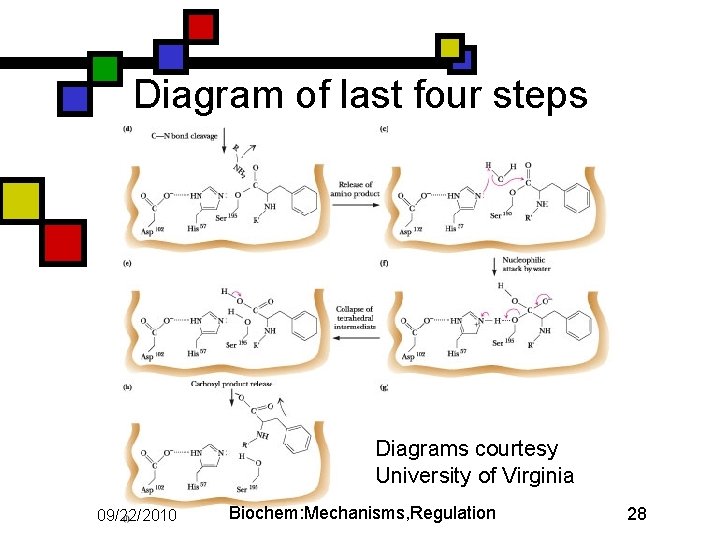
Diagram of last four steps Diagrams courtesy University of Virginia 09/22/2010 Biochem: Mechanisms, Regulation 28
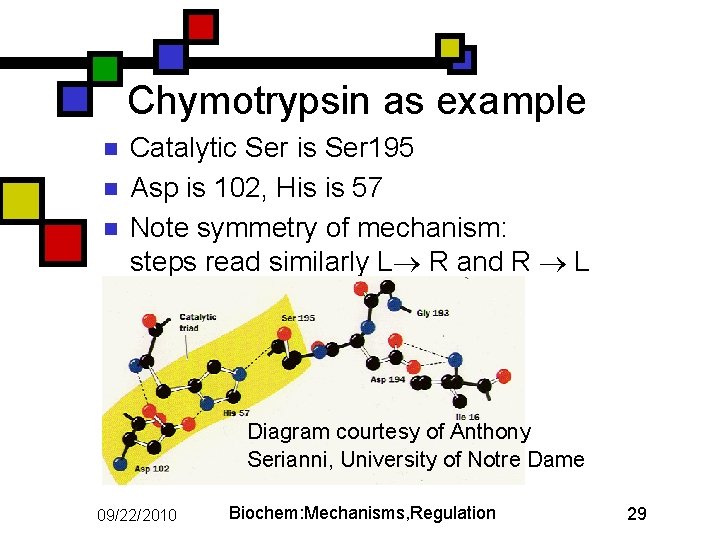
Chymotrypsin as example n n n Catalytic Ser is Ser 195 Asp is 102, His is 57 Note symmetry of mechanism: steps read similarly L R and R L Diagram courtesy of Anthony Serianni, University of Notre Dame 09/22/2010 Biochem: Mechanisms, Regulation 29
Oxyanion hole n n When his-57 accepts proton from Ser-195: it creates an R—O- ion on Ser sidechain In reality the Ser O immediately becomes covalently bonded to substrate carbonyl carbon, moving negative charge to the carbonyl O. Oxyanion is on the substrate's oxygen Oxyanion stabilized by additional interaction in addition to the protonated his 57: main-chain NH group from gly 193 H-bonds to oxygen atom (or ion) from the substrate, further stabilizing the ion. 09/22/2010 Biochem: Mechanisms, Regulation 30
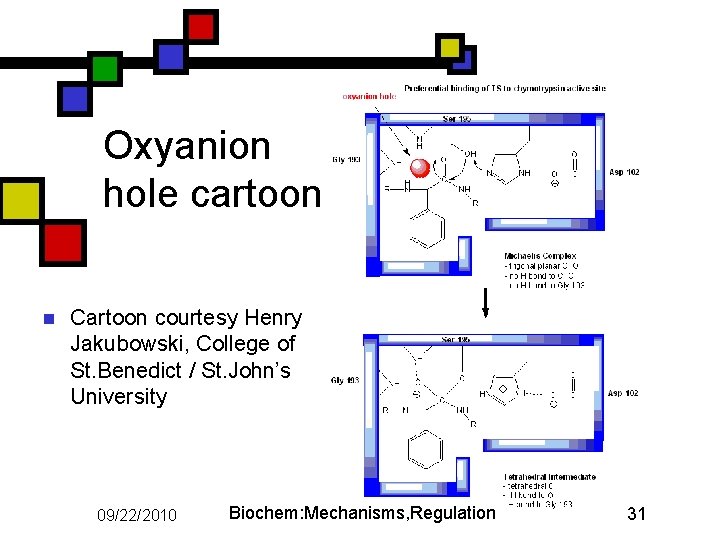
Oxyanion hole cartoon n Cartoon courtesy Henry Jakubowski, College of St. Benedict / St. John’s University 09/22/2010 Biochem: Mechanisms, Regulation 31
Modes of catalysis in serine proteases n n Proximity effect: gathering of reactants in steps 1 and 4 Acid-base catalysis at histidine in steps 2 and 4 Covalent catalysis on serine hydroxymethyl group in steps 2 -5 So both chemical (acid-base & covalent) and binding modes (proximity & transition-state) are used in this mechanism 09/22/2010 Biochem: Mechanisms, Regulation 32

Specificity n n n Active site catalytic triad is nearly invariant for eukaryotic serine proteases Remainder of cavity where reaction occurs varies significantly from protease to protease. In chymotrypsin hydrophobic pocket just upstream of the position where scissile bond sits This accommodates large hydrophobic side chain like that of phe, and doesn’t comfortably accommodate hydrophilic or small side chain. Thus specificity is conferred by the shape and electrostatic character of the site. 09/22/2010 Biochem: Mechanisms, Regulation 33

Chymotrypsin active site n n Comfortably accommodates aromatics at S 1 site Differs from other mammalian serine proteases in specificity Diagram courtesy School of Crystallography, Birkbeck College 09/22/2010 Biochem: Mechanisms, Regulation 34
Divergent evolution n Ancestral eukaryotic serine proteases presumably have differentiated into forms with different side-chain specificities Chymotrypsin is substantially conserved within eukaryotes, but is distinctly different from elastase Primary differences are in P 1 side chain pocket, but that isn’t inevitable 09/22/2010 Biochem: Mechanisms, Regulation 35
i. Clicker quiz, question 1 Why would the nonproductive hexokinase reaction H 2 O + ATP ADP + Pi be considered nonproductive? n (a) Because it needlessly soaks up water n (b) Because the enzyme undergoes a wasteful conformational change n (c) Because the energy in the high-energy phosphate bond is unavailable for other purposes n (d) Because ADP is poisonous n (e) None of the above 09/22/2010 Biochem: Mechanisms, Regulation 36

i. Clicker Quiz question 2 What would bind tightest in the TIM active site? n (a) DHAP (substrate) n (b) D-glyceraldehyde (product) n (c) 2 -phosphoglycolate (Transition-state analog) n (d) They would all bind equally well n (e) None of them would bind at all. 09/22/2010 Biochem: Mechanisms, Regulation 37
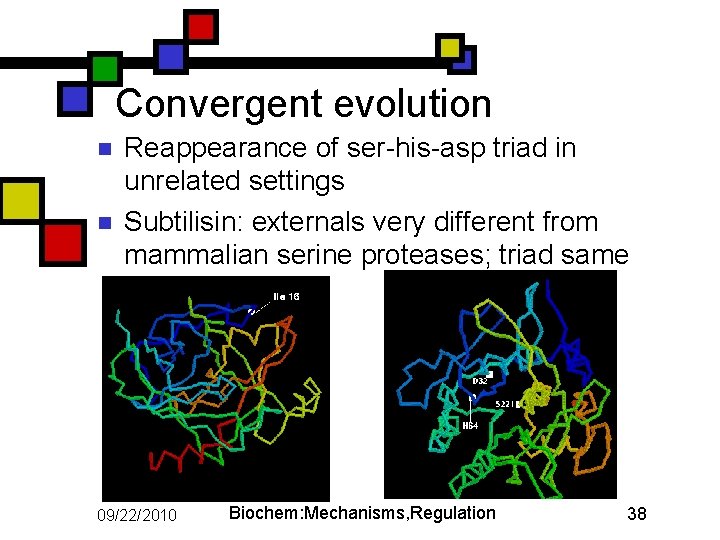
Convergent evolution n n Reappearance of ser-his-asp triad in unrelated settings Subtilisin: externals very different from mammalian serine proteases; triad same 09/22/2010 Biochem: Mechanisms, Regulation 38

Subtilisin mutagenesis n n n Substitutions for any of the amino acids in the catalytic triad has disastrous effects on the catalytic activity, as measured by kcat. Km affected only slightly, since the structure of the binding pocket is not altered very much by conservative mutations. An interesting (and somewhat non-intuitive) result is that even these "broken" enzymes still catalyze the hydrolysis of some test substrates at much higher rates than buffer alone would provide. I would encourage you to think about why that might be true. 09/22/2010 Biochem: Mechanisms, Regulation 39

Cysteinyl proteases n n n Ancestrally related to ser proteases? Cathepsins, caspases, papain Contrasts: n n n Cys —SH is more basic than ser —OH Residue is less hydrophilic S- is a weaker nucleophile than O- 09/22/2010 Diagram courtesy of Mariusz Jaskolski, U. Poznan Biochem: Mechanisms, Regulation 40
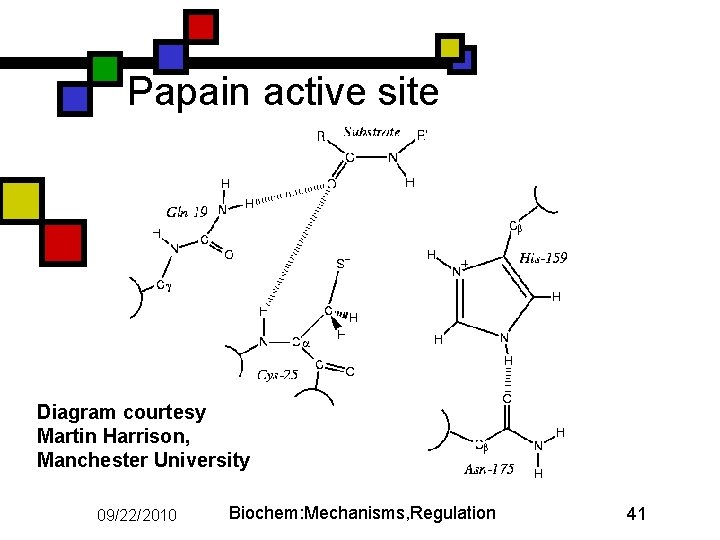
Papain active site Diagram courtesy Martin Harrison, Manchester University 09/22/2010 Biochem: Mechanisms, Regulation 41
Hen egg-white lysozyme n n n Antibacterial protectant of growing chick embryo Hydrolyzes bacterial cell-wall peptidoglycans “hydrogen atom of structural biology” n n HEWL PDB 2 vb 1 0. 65Å 15 k. Da Commercially available in pure form Easy to crystallize and do structure work Available in multiple crystal forms Mechanism is surprisingly complex 09/22/2010 Biochem: Mechanisms, Regulation 42
Mechanism of lysozyme n n n Strain-induced destabilization of substrate makes the substrate look more like the transition state Long arguments about the nature of the intermediates Accepted answer: covalent intermediate between D 52 and glycosyl C 1 (Garrett & Grisham, fig. 14. 39 B) 09/22/2010 Biochem: Mechanisms, Regulation 43
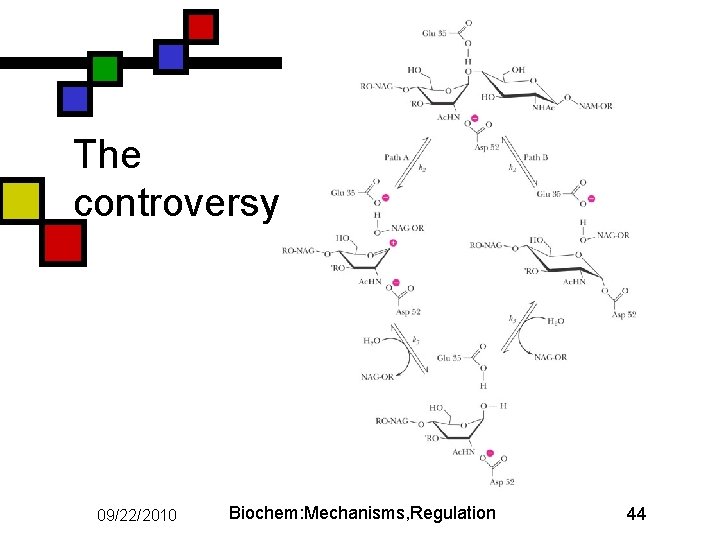
The controversy 09/22/2010 Biochem: Mechanisms, Regulation 44

Regulation of enzymes n n The very catalytic proficiency for which enzymes have evolved means that their activity must not be allowed to run amok Activity is regulated in many ways: n n n Thermodynamics Enzyme availability Allostery Post-translational modification Protein-protein interactions 09/22/2010 Biochem: Mechanisms, Regulation 45

Thermodynamics as a regulatory force n n n Remember that Go (or Go’) is not the determiner of spontaneity: G is. Therefore: local product and substrate concentrations determine whether the enzyme is catalyzing reversible reactions to the left or to the right Rule of thumb: Go’ < -20 k. J mol-1 is irreversible 09/22/2010 Biochem: Mechanisms, Regulation 46

Enzyme availability n n n The enzyme has to be where the reactants are in order for it to act Even a highly proficient enzyme has to have a nonzero concentration How can the cell control [E]tot? n n n Transcription (and translation) Protein processing (degradation) Compartmentalization 09/22/2010 Biochem: Mechanisms, Regulation 47
Transcriptional control n n n m. RNAs have short lifetimes Therefore once a protein is degraded, it will be replaced and available only if new transcriptional activity for that protein occurs Many types of transcriptional effectors n n n Proteins can bind to their own gene Small molecules can bind to gene Promoters can be turned on or off 09/22/2010 Biochem: Mechanisms, Regulation 48
Protein degradation n n All proteins have finite half-lives; Enzymes’ lifetimes often shorter than structural or transport proteins Degraded by slings & arrows of outrageous fortune; or Activity of the proteasome, a molecular machine that tags proteins for degradation and then accomplishes it 09/22/2010 Biochem: Mechanisms, Regulation 49
How the proteasome works n n Proteins in need of degradation are tagged by covalent linkage to the small protein ubiquitin, or to a chain of several ubiquitin molecules Proteasome is molecular machine that recognized ubiquitinated proteins n n Cleaves off the ubiquitin(s) for re-use Proteolytically degrades our initial protein 09/22/2010 Biochem: Mechanisms, Regulation 50
Compartmentalization n If the enzyme is in one compartment and the substrate in another, it won’t catalyze anything Several mitochondrial catabolic enzyme act on substrates produced in the cytoplasm; these require elaborate transport mechanisms to move them in Therefore, control of the transporters confers control over the enzymatic system 09/22/2010 Biochem: Mechanisms, Regulation 51
Allostery n n Remember we defined this as an effect on protein activity in which binding of a ligand to a protein induces a conformational change that modifies the protein’s activity Ligand may be the same molecule as the substrate or it may be a different one (homotropic vs. heterotropic allostery) Ligand may bind to the same subunit or a different one These effects happen to non-enzymatic proteins as well as enzymes 09/22/2010 Biochem: Mechanisms, Regulation 52

Substrates as allosteric effectors (homotropic) n n n Standard example: binding of O 2 to one subunit of tetrameric hemoglobin induces conformational change that facilitates binding of 2 nd (& 3 rd & 4 th) O 2’s So the first oxygen is an allosteric effector of the activity in the other subunits Effect can be inhibitory or accelerative 09/22/2010 Biochem: Mechanisms, Regulation 53

Other allosteric effectors (heterotropic) n n n Covalent modification of an enzyme by phosphate or other PTM molecules can turn it on or off Usually catabolic enzymes are stimulated by phosphorylation and anabolic enzymes are turned off, but not always Phosphatases catalyze dephosphorylation; these have the opposite effects 09/22/2010 Biochem: Mechanisms, Regulation 54
Cyclic AMP-dependent protein kinases n n n Enzymes phosphorylate proteins with S or T within sequence R(R/K)X(S*/T*) Intrasteric control: regulatory subunit or domain has a sequence that looks like the target sequence; this binds and inactivates the kinase’s catalytic subunit When regulatory subunits binds c. AMP, it releases from the catalytic subunit so it can do its thing 09/22/2010 Biochem: Mechanisms, Regulation 55
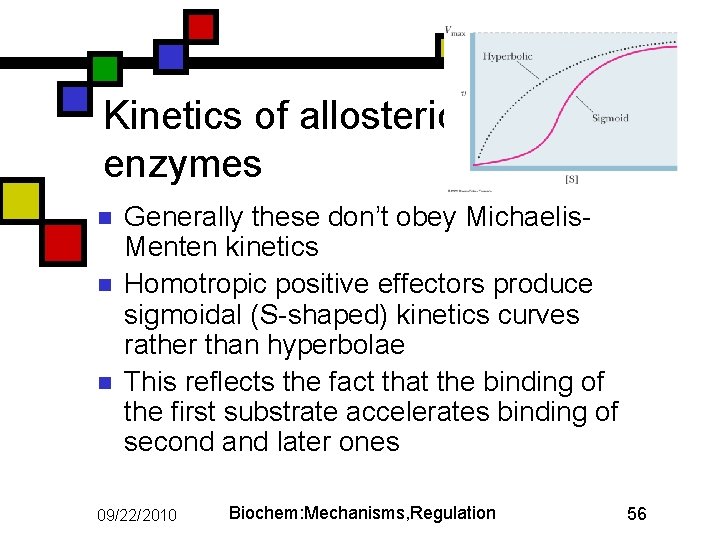
Kinetics of allosteric enzymes n n n Generally these don’t obey Michaelis. Menten kinetics Homotropic positive effectors produce sigmoidal (S-shaped) kinetics curves rather than hyperbolae This reflects the fact that the binding of the first substrate accelerates binding of second and later ones 09/22/2010 Biochem: Mechanisms, Regulation 56
T R State transitions n n Many allosteric effectors influence the equilibrium between two conformations One is typically more rigid and inactive, the other is more flexible and active The rigid one is typically called the “tight” or “T” state; the flexible one is called the “relaxed” or “R” state Allosteric effectors shift the equilibrium toward R or toward T 09/22/2010 Biochem: Mechanisms, Regulation 57

Examples you’ve already seen n Glycogen phosphorylase: R state is more effective at phosphorylating glycogen Tetrameric hemoglobin: R state binds oxygen better Hexokinase: R state binds ATP 09/22/2010 Biochem: Mechanisms, Regulation 58
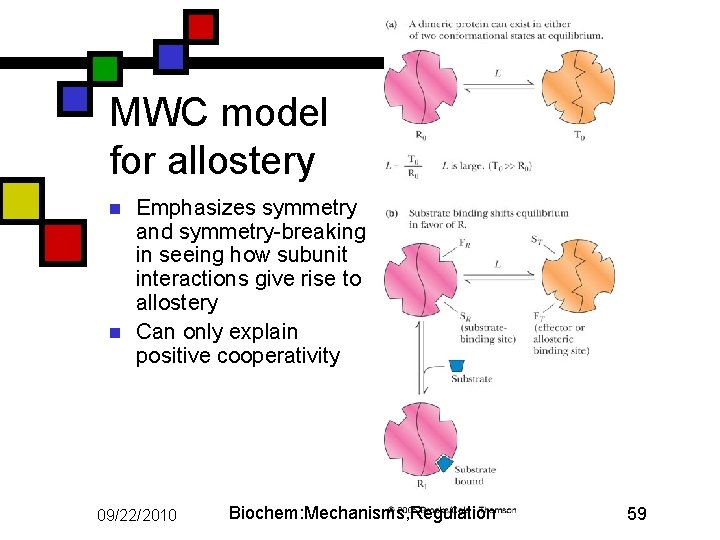
MWC model for allostery n n Emphasizes symmetry and symmetry-breaking in seeing how subunit interactions give rise to allostery Can only explain positive cooperativity 09/22/2010 Biochem: Mechanisms, Regulation 59
Koshland (KNF) model n n Emphasizes conformational changes from one state to another, induced by binding of effector Ligand binding and conformational transitions are distinct steps … so this is a sequential model for allosteric transitions Allows for negative cooperativity as well as positive cooperativity 09/22/2010 Biochem: Mechanisms, Regulation 60
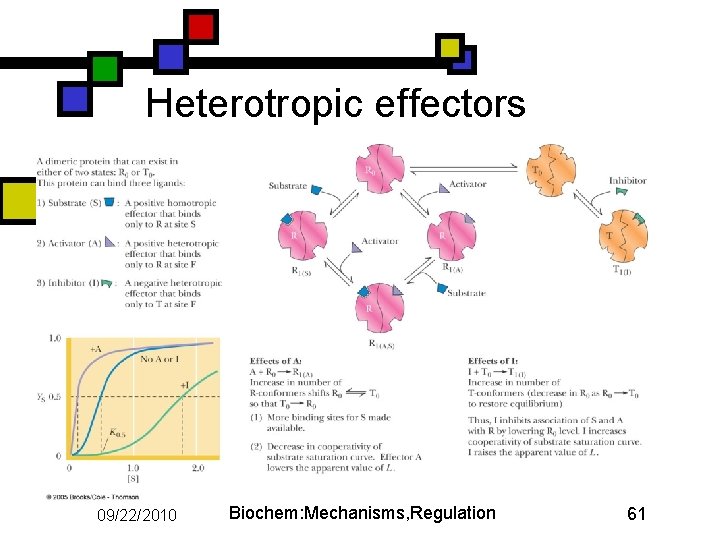
Heterotropic effectors 09/22/2010 Biochem: Mechanisms, Regulation 61

Post-translational modification n n We’ve already looked at phosphorylation Proteolytic cleavage of the enzyme to activate it is another common PTM mode Some proteases cleave themselves (auto -catalysis); in other cases there’s an external protease involved Blood-clotting cascade involves a series of catalytic activations 09/22/2010 Biochem: Mechanisms, Regulation 62
Zymogens n n n As mentioned earlier, this is a term for an inactive form of a protein produced at the ribosome Proteolytic post-translational processing required for the zymogen to be converted to its active form Cleavage may happen intracellularly, during secretion, or extracellularly 09/22/2010 Biochem: Mechanisms, Regulation 63
Blood clotting n n n Seven serine proteases in cascade Final one (thrombin) converts fibrinogen to fibrin, which can aggregate to form an insoluble mat to prevent leakage Two different pathways: n n n Intrinsic: blood sees injury directly Extrinsic: injured tissues release factors that stimulate process Come together at factor X 09/22/2010 Biochem: Mechanisms, Regulation 64
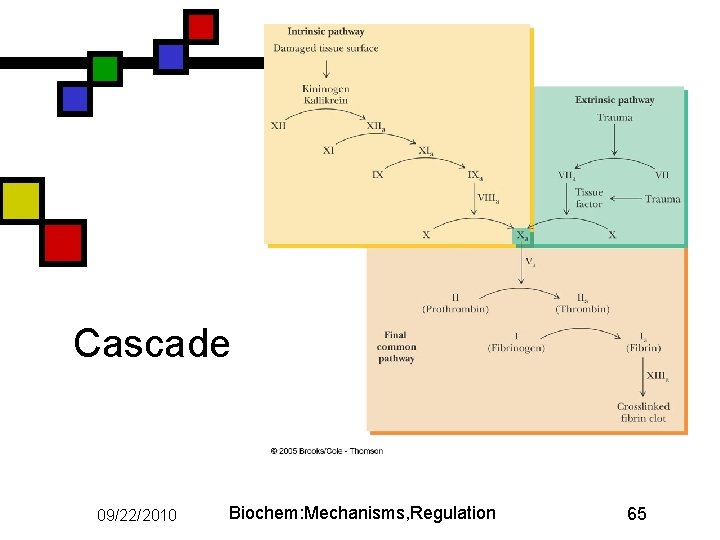
Cascade 09/22/2010 Biochem: Mechanisms, Regulation 65
Protein-protein interactions n n n One major change in biochemistry in the last 20 years is the increasing emphasis on protein interactions in understanding biological activities Many proteins depend on exogenous partners for modulating their activity up or down Example: cholera toxin’s enzymatic component depends on interaction with human protein ARF 6 09/22/2010 Biochem: Mechanisms, Regulation 66

Globins as aids to understanding n n Myoglobin and hemoglobin are wellunderstood non-enzymatic proteins whose properties help us understand enzyme regulation Hemoglobin is described as an “honorary enzyme” in that it “catalyzes” the reaction O 2(lung) O 2 (peripheral tissues) 09/22/2010 Biochem: Mechanisms, Regulation 67
Setting the stage for this story n n n Myoglobin is a 16 k. Da monomeric O 2 -storage protein found in peripheral tissues Has Fe-containing prosthetic group called heme; iron must be in Fe 2+ state to bind O 2 It yields up dioxygen to various oxygenrequiring processes, particularly oxidative phosphorylation in mitochondria in rapidly metabolizing tissues 09/22/2010 Biochem: Mechanisms, Regulation 68
Why is myoglobin needed? n Free heme will bind O 2 nicely; why not just rely on that? n Protein has 3 functions: Immobilizes the heme group n Discourages oxidation of Fe 2+ to Fe 3+ n Provides a pocket that oxygen can fit into n 09/22/2010 Biochem: Mechanisms, Regulation 69
Setting the stage II n n n Hemoglobin (in vertebrates, at least) is a tetrameric, 64 k. Da transport protein that carries oxygen from the lungs to peripheral tissues It also transports acidic CO 2 the opposite direction Its allosteric properties are what we’ll discuss 09/22/2010 Biochem: Mechanisms, Regulation 70

n Structure determinations Myoglobin & hemoglobin were the first 2 proteins to have their 3 -D structures determined experimentally n n Photo courtesy EMBL Myoglobin: Kendrew, 1958 Hemoglobin: Perutz, 1958 Most of the experimental tools that crystallographers rely on were developed for these structure determinations Nobel prizes for both, 1965 (small T!) 09/22/2010 Biochem: Mechanisms, Regulation Photo courtesy Oregon State Library 71
- Slides: 71