Enzyme Inhibition and Mechanisms Andy Howard Introductory Biochemistry

Enzyme Inhibition and Mechanisms Andy Howard Introductory Biochemistry, Fall 2010 Monday 20 September 2010 Biochem: Inhibition & Mechanisms 1 09/20/2010
How do enzymes reduce activation energies? We want to understand what is really happening chemically when an enzyme does its job. n We’d also like to know how biochemists probe these systems. n 09/20/2010 Biochem: Inhibition & Mechanisms 2
Inhibition & Mechanism Topics n n Enzyme kinetics, concluded Inhibition n n Reversible & Not Categories Kinetics Drug Design n Mechanisms n n n 09/20/2010 Terminology Transition States Diffusion-controlled Reactions Binding Modes Intermediates Types of reactions Biochem: Inhibition & Mechanisms 3
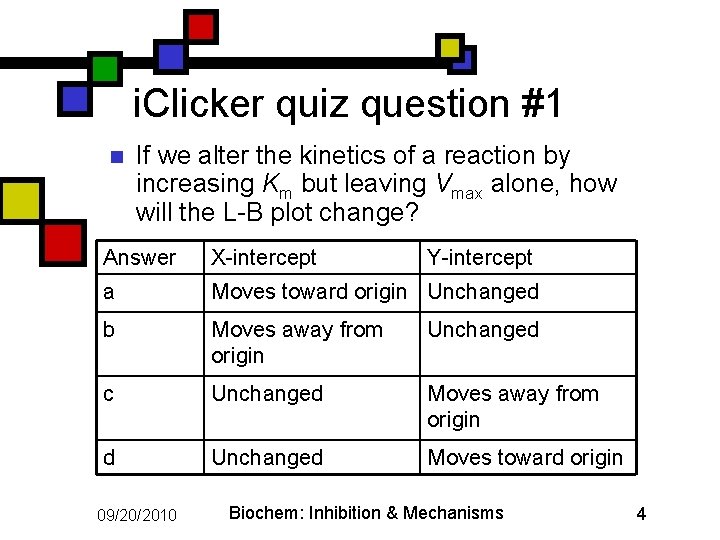
i. Clicker quiz question #1 n If we alter the kinetics of a reaction by increasing Km but leaving Vmax alone, how will the L-B plot change? Answer X-intercept a Moves toward origin Unchanged b Moves away from origin Unchanged c Unchanged Moves away from origin d Unchanged Moves toward origin 09/20/2010 Y-intercept Biochem: Inhibition & Mechanisms 4
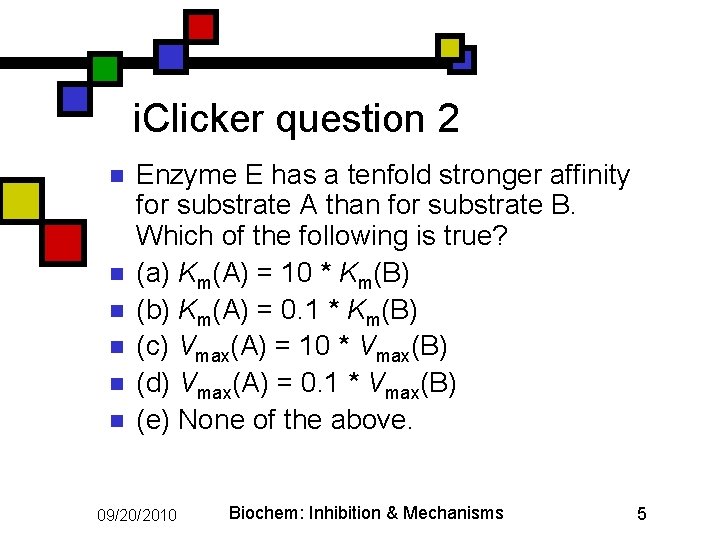
i. Clicker question 2 n n n Enzyme E has a tenfold stronger affinity for substrate A than for substrate B. Which of the following is true? (a) Km(A) = 10 * Km(B) (b) Km(A) = 0. 1 * Km(B) (c) Vmax(A) = 10 * Vmax(B) (d) Vmax(A) = 0. 1 * Vmax(B) (e) None of the above. 09/20/2010 Biochem: Inhibition & Mechanisms 5

Another physical significance of Km n n n Years of experience have led biochemists to a general conclusion: For its preferred substrate, the Km value of an enzyme is usually within a factor of 50 of the steady-state concentration of that substrate. So if we find that Km = 0. 2 m. M for the primary substrate of an enzyme, then we expect that the steady-state concentration of that substrate is between 4 µM and 10 m. M. 09/20/2010 Biochem: Inhibition & Mechanisms 6
Example: hexokinase isozymes n n Mutant human type I hexokinase PDB 1 DGK, 2. 8Å 110 k. Da monomer Hexokinase catalyzes hexose + ATP hexose-6 -P + ADP Most isozymes of hexokinase prefer glucose; some also work okay mannose and fructose Muscle hexokinases have Km ~ 0. 1 m. M so they work efficiently in blood, where [glucose] ~ 4 m. M Liver glucokinase has Km = 10 m. M, which is around the liver [glucose] and can respond to fluctuations in liver [glucose] 09/20/2010 Biochem: Inhibition & Mechanisms 7

L-B plots for ordered sequential reactions n n n http: //www-biol. paisley. ac. uk/ kinetics/Chapter_4/chapter 4_3. html Plot 1/v 0 vs. 1/[A] for various [B] values; flatter slopes correspond to larger [B] Lines intersect @ a point in between X intercept and Y intercept 09/20/2010 Biochem: Inhibition & Mechanisms 8
![L-B plots for pingpong reactions n n n Again we plot 1/v vs 1/[A] L-B plots for pingpong reactions n n n Again we plot 1/v vs 1/[A]](http://slidetodoc.com/presentation_image_h/a64387098f7fcb0a273e29df8b95cc76/image-9.jpg)
L-B plots for pingpong reactions n n n Again we plot 1/v vs 1/[A] for various [B] Parallel lines (same kcat/Km); lower lines correspond to larger [B] http: //www-biol. paisley. ac. uk/kinetics/ Chapter_4/chapter 4_3_2. html 09/20/2010 Biochem: Inhibition & Mechanisms 9

Inhibition is important both conceptually and practically n We study inhibition to clarify our understanding of enzyme mechanisms and because knowing how inhibition works helps us design pharmaceuticals. 09/20/2010 Biochem: Inhibition & Mechanisms 10

Why study inhibition? • Let’s look at how enzymes get inhibited. • At least two reasons to do this: • We can use inhibition as a probe for understanding the kinetics and properties of enzymes in their uninhibited state; • Many—perhaps most—drugs are inhibitors of specific enzymes. • We'll see these two reasons for understanding inhibition as we work our way through this topic. 09/20/2010 Biochem: Inhibition & Mechanisms 11
The concept of inhibition n An enzyme is a biological catalyst, i. e. a substance that alters the rate of a reaction without itself becoming permanently altered by its participation in the reaction. n The ability of an enzyme (particularly a proteinaceous enzyme) to catalyze a reaction can be altered by binding small molecules to it: n sometimes at its active site n sometimes at a site distant from the active site. 09/20/2010 Biochem: Inhibition & Mechanisms 12

Inhibitors and accelerators n Usually these alterations involve a reduction in the enzyme's ability to accelerate the reaction; less commonly, they give rise to an increase in the enzyme's ability to accelerate a reaction. 09/20/2010 Biochem: Inhibition & Mechanisms 13

Why more inhibitors than accelerators? n n Natural selection: if there were small molecules that can facilitate the enzyme's propensity to speed up a reaction, nature probably would have found a way to incorporate those facilitators into the enzyme over the billions of years that the enzyme has been available. Most enzymes are already fairly close to optimal in their properties; we can readily mess them up with effectors, but it's more of a challenge to find ways to make enzymes better at their jobs. 09/20/2010 Biochem: Inhibition & Mechanisms 14
Types of inhibitors n Irreversible n n Inhibitor binds without possibility of release Usually covalent Each inhibition event effectively removes a molecule of enzyme from availability Reversible n n n Usually noncovalent (ionic or van der Waals) Several kinds Classifications somewhat superseded by detailed structure-based knowledge of mechanisms, but not entirely 09/20/2010 Biochem: Inhibition & Mechanisms 15
Types of reversible inhibition n n Competitive n Inhibitor binds at active site n Prevents binding of substrate Noncompetitive n Inhibitor binds distant from active site n Interferes with turnover Uncompetitive (rare? ) n Inhibitor binds to ES complex n Removes ES, interferes with turnover Mixed (usually Competitive + Noncompetitive) 09/20/2010 Biochem: Inhibition & Mechanisms 16
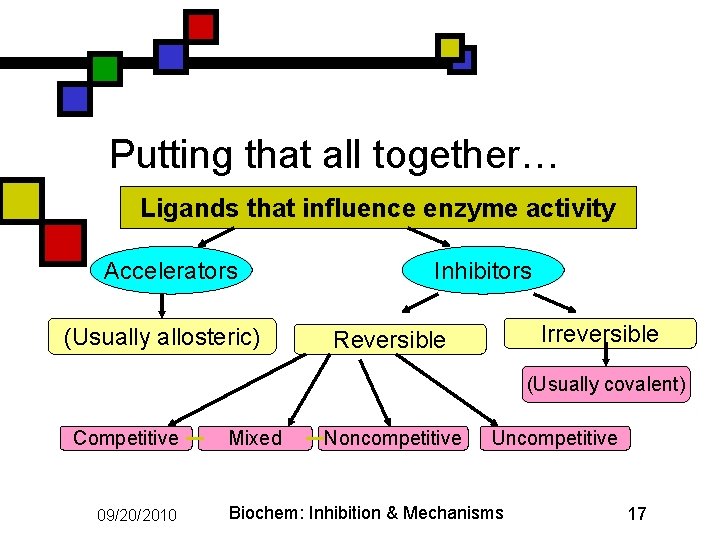
Putting that all together… Ligands that influence enzyme activity Accelerators (Usually allosteric) Inhibitors Irreversible Reversible (Usually covalent) Competitive 09/20/2010 Mixed Noncompetitive Uncompetitive Biochem: Inhibition & Mechanisms 17
How to tell them apart n Reversible vs irreversible n n n dialyze an enzyme-inhibitor complex against a buffer free of inhibitor if turnover or binding still suffers, it’s irreversible Competitive vs. other reversible: n n 09/20/2010 Structural studies if feasible Kinetics Biochem: Inhibition & Mechanisms 18
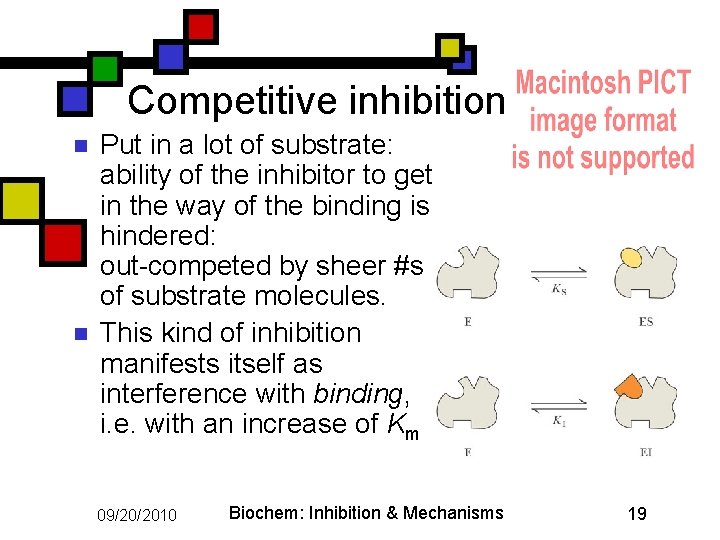
Competitive inhibition n n Put in a lot of substrate: ability of the inhibitor to get in the way of the binding is hindered: out-competed by sheer #s of substrate molecules. This kind of inhibition manifests itself as interference with binding, i. e. with an increase of Km 09/20/2010 Biochem: Inhibition & Mechanisms 19

Competitive inhibitors don’t affect turnover n Within the active site of any given molecule of the enzyme, one of three states are possible: n n The inhibitor is present The substrate is present Nothing is present Therefore the rate of turnover isn’t affected by the inhibitor: just the availability of binding sites. 09/20/2010 Biochem: Inhibition & Mechanisms 20

Kinetics of competition n Competitive inhibitor hinders binding of substrate but not reaction velocity: Affects the Km of the enzyme, not Vmax. Which way does it affect it? n n n Km = amount of substrate that needs to be present to run the reaction velocity up to half its saturation velocity. Competitive inhibitor requires us to shove more substrate into the reaction in order to achieve that half-maximal velocity. So: competitive inhibitor increases Km 09/20/2010 Biochem: Inhibition & Mechanisms 21
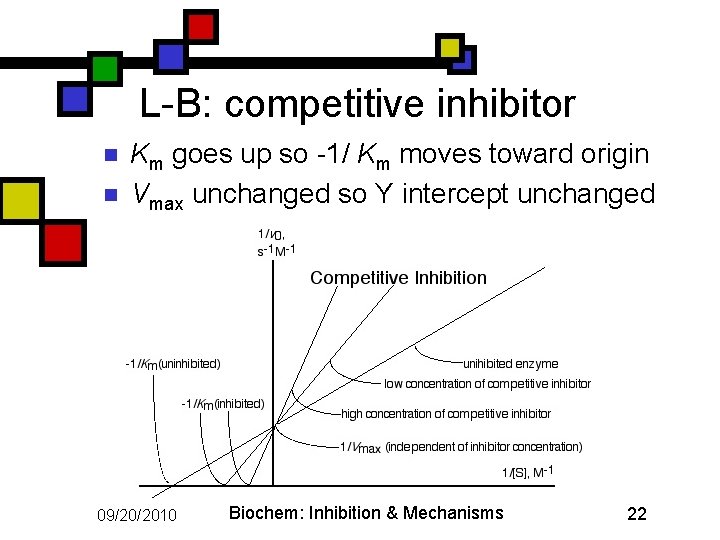
L-B: competitive inhibitor n n Km goes up so -1/ Km moves toward origin Vmax unchanged so Y intercept unchanged 09/20/2010 Biochem: Inhibition & Mechanisms 22
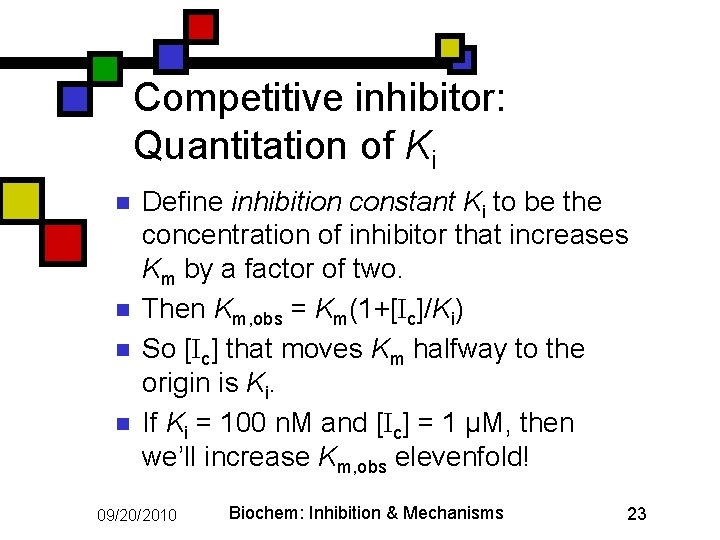
Competitive inhibitor: Quantitation of Ki n n Define inhibition constant Ki to be the concentration of inhibitor that increases Km by a factor of two. Then Km, obs = Km(1+[Ic]/Ki) So [Ic] that moves Km halfway to the origin is Ki. If Ki = 100 n. M and [Ic] = 1 µM, then we’ll increase Km, obs elevenfold! 09/20/2010 Biochem: Inhibition & Mechanisms 23

Noncompetitive I inhibition n n S Noncompetitive inhibitor has no influence on how available the binding site for substrate is, so it doesn’t affect Km at all However, it has a profound inhibitory influence on the speed of the reaction, i. e. turnover. So it reduces Vmax and has no influence on Km. 09/20/2010 Biochem: Inhibition & Mechanisms 24
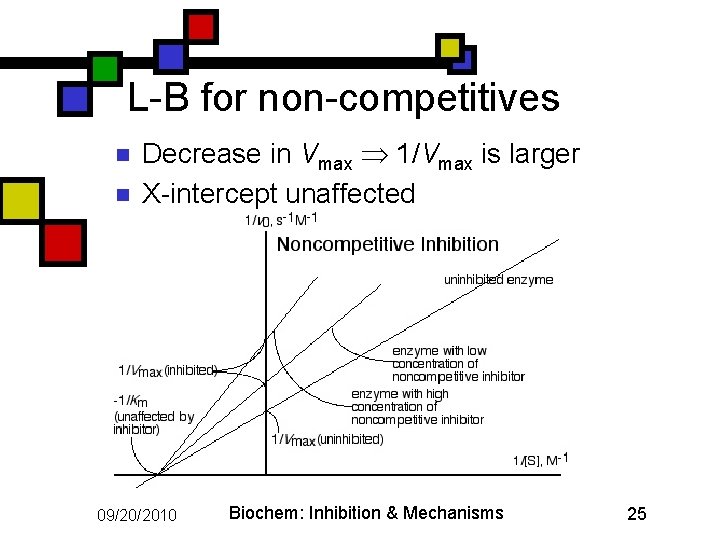
L-B for non-competitives n n Decrease in Vmax 1/Vmax is larger X-intercept unaffected 09/20/2010 Biochem: Inhibition & Mechanisms 25
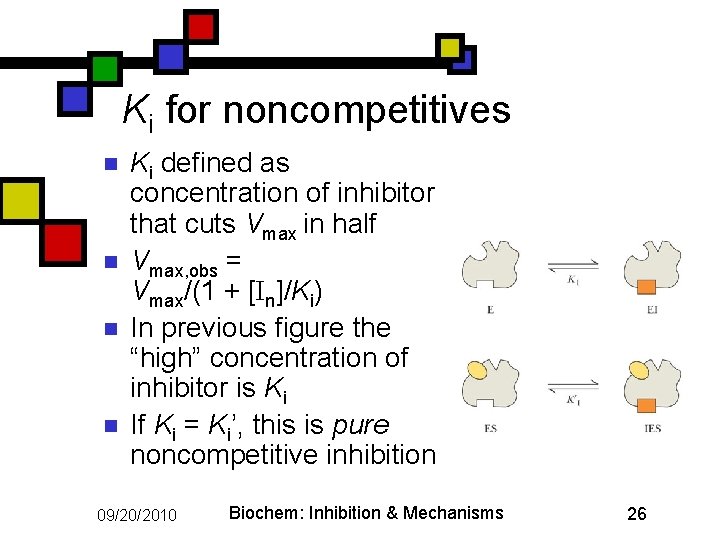
Ki for noncompetitives n n Ki defined as concentration of inhibitor that cuts Vmax in half Vmax, obs = Vmax/(1 + [In]/Ki) In previous figure the “high” concentration of inhibitor is Ki If Ki = Ki’, this is pure noncompetitive inhibition 09/20/2010 Biochem: Inhibition & Mechanisms 26
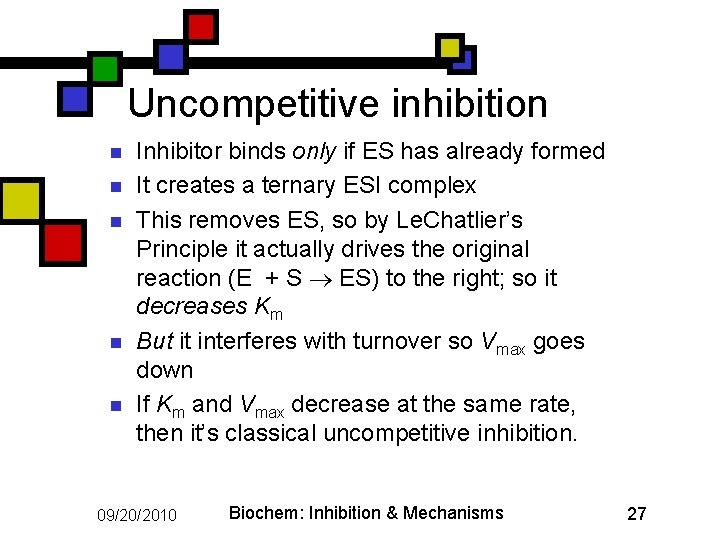
Uncompetitive inhibition n n Inhibitor binds only if ES has already formed It creates a ternary ESI complex This removes ES, so by Le. Chatlier’s Principle it actually drives the original reaction (E + S ES) to the right; so it decreases Km But it interferes with turnover so Vmax goes down If Km and Vmax decrease at the same rate, then it’s classical uncompetitive inhibition. 09/20/2010 Biochem: Inhibition & Mechanisms 27
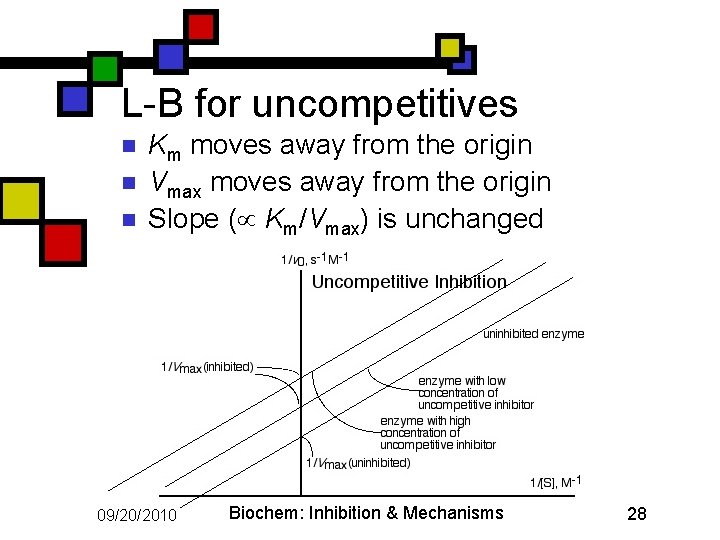
L-B for uncompetitives n n n Km moves away from the origin Vmax moves away from the origin Slope ( Km/Vmax) is unchanged 09/20/2010 Biochem: Inhibition & Mechanisms 28
Ki for uncompetitives Defined as inhibitor concentration that cuts Vmax or Km in half n Easiest to read from Vmax value n Iu labeled “high” is Ki in this plot n 09/20/2010 Biochem: Inhibition & Mechanisms 29

i. Clicker question 3 What assumptions does the standard derivation of the Michaelis-Menten equation depend on? n n n (a) d[ES]/dt = 0 for some reasonable period (b) Formation of product is rate-limiting (c) [P] = 0 at time t=0, or else k-2 = 0 (d) All of the above (e) None of the above 09/20/2010 Biochem: Inhibition & Mechanisms 30
i. Clicker quiz, question 4 Treatment of enzyme E with compound Y doubles Km and leaves Vmax unchanged. Compound Y is: n (a) an accelerator of the reaction n (b) a competitive inhibitor n (c) a non-competitive inhibitor n (d) an uncompetitive inhibitor 09/20/2010 Biochem: Inhibition & Mechanisms 31
i. Clicker quiz, question 5 Treatment of enzyme E with compound X doubles Vmax and leaves Km unchanged. Compound X is: (a) an accelerator of the reaction n (b) a competitive inhibitor n (c) a non-competitive inhibitor n (d) an uncompetitive inhibitor n 09/20/2010 Biochem: Inhibition & Mechanisms 32
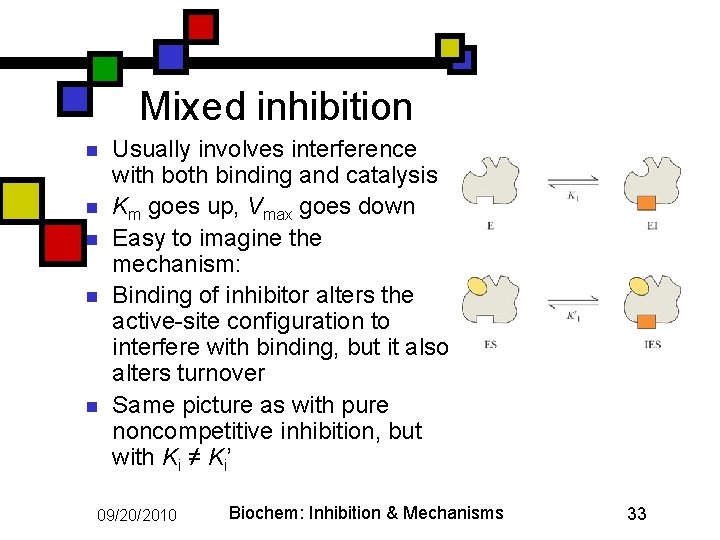
Mixed inhibition n n Usually involves interference with both binding and catalysis Km goes up, Vmax goes down Easy to imagine the mechanism: Binding of inhibitor alters the active-site configuration to interfere with binding, but it also alters turnover Same picture as with pure noncompetitive inhibition, but with Ki ≠ Ki’ 09/20/2010 Biochem: Inhibition & Mechanisms 33

Most pharmaceuticals are enzyme inhibitors n n n Some are inhibitors of enzymes that are necessary for functioning of pathogens Others are inhibitors of some protein whose inappropriate expression in a human causes a disease. Others are targeted at enzymes that are produced more energetically by tumors than they are by normal tissues. 09/20/2010 Biochem: Inhibition & Mechanisms 34
Characteristics of Pharmaceutical Inhibitors n n Usually competitive, i. e. they raise Km without affecting Vmax Some are mixed, i. e. Km up, Vmax down Iterative design work will decrease Ki from millimolar down to nanomolar Sometimes design work is purely blind HTS; other times, it’s structure-based 09/20/2010 Biochem: Inhibition & Mechanisms 35

Amprenavir n n Competitive inhibitor of HIV protease, Ki = 0. 6 n. M for HIV-1 No longer sold: mutual interference with rifabutin, which is an antibiotic used against a common HIV secondary bacterial infection, Mycobacterium avium 09/20/2010 Biochem: Inhibition & Mechanisms 36
When is a good inhibitor a good drug? n n n It needs to be bioavailable and nontoxic Beautiful 20 n. M inhibitor is often neither Modest sacrifices of Ki in improving bioavailability and non-toxicity are okay if Ki is low enough when you start sacrificing 09/20/2010 Biochem: Inhibition & Mechanisms 37

How do we lessen toxicity and improve bioavailability? n n n Increase solubility… that often increases Ki because the van der Waals interactions diminish Solubility makes it easier to get the compound to travel through the bloodstream Toxicity is often associated with fat storage, which is more likely with insoluble compounds 09/20/2010 Biochem: Inhibition & Mechanisms 38
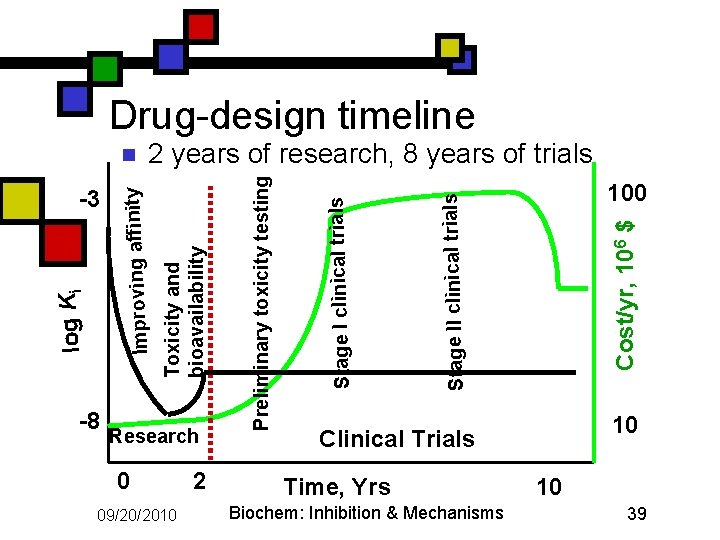
Drug-design timeline Research 0 09/20/2010 2 100 Cost/yr, 106 $ Stage II clinical trials Stage I clinical trials Preliminary toxicity testing -8 Toxicity and bioavailability log Ki -3 2 years of research, 8 years of trials Improving affinity n 10 Clinical Trials Time, Yrs Biochem: Inhibition & Mechanisms 10 39
Atomic-Level Mechanisms n n n We want to understand atomic-level events during an enzymatically catalyzed reaction. Sometimes we want to find a way to inhibit an enzyme in other cases we're looking for more fundamental knowledge, viz. the ways that biological organisms employ chemistry and how enzymes make that chemistry possible. 09/20/2010 Biochem: Inhibition & Mechanisms 40
How we study mechanisms n There a variety of experimental tools available for understanding mechanisms, including isotopic labeling of substrates, structural methods, and spectroscopic kinetic techniques. 09/20/2010 Biochem: Inhibition & Mechanisms 41
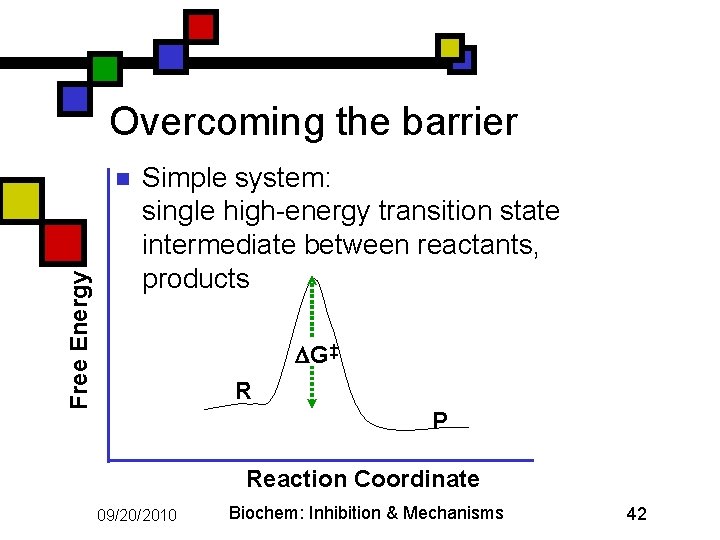
Overcoming the barrier Free Energy n Simple system: single high-energy transition state intermediate between reactants, products G‡ R P Reaction Coordinate 09/20/2010 Biochem: Inhibition & Mechanisms 42
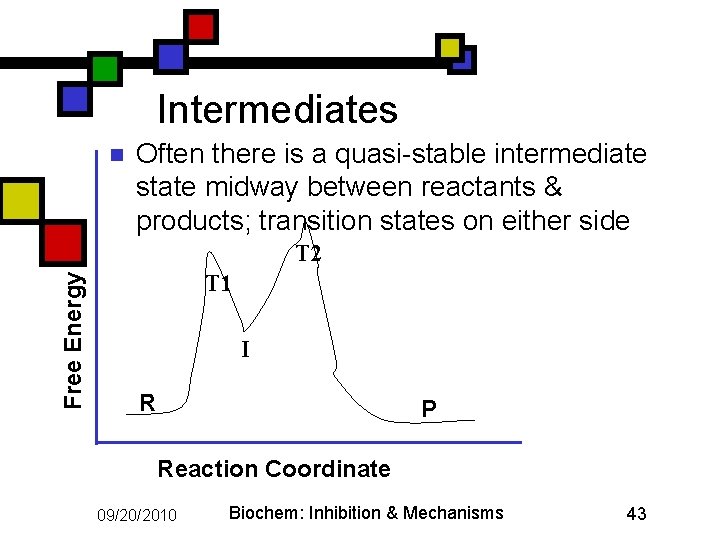
Intermediates n Often there is a quasi-stable intermediate state midway between reactants & products; transition states on either side Free Energy T 2 T 1 I R P Reaction Coordinate 09/20/2010 Biochem: Inhibition & Mechanisms 43

Activation energy & temperature n n It’s intuitively sensible that higher temperatures would make it easier to overcome an activation barrier Rate k(T) = Q 0 exp(- G‡/RT) G‡ = activation energy or Arrhenius energy This provides tool for measuring G‡ 09/20/2010 Biochem: Inhibition & Mechanisms Svante Arrhenius 44
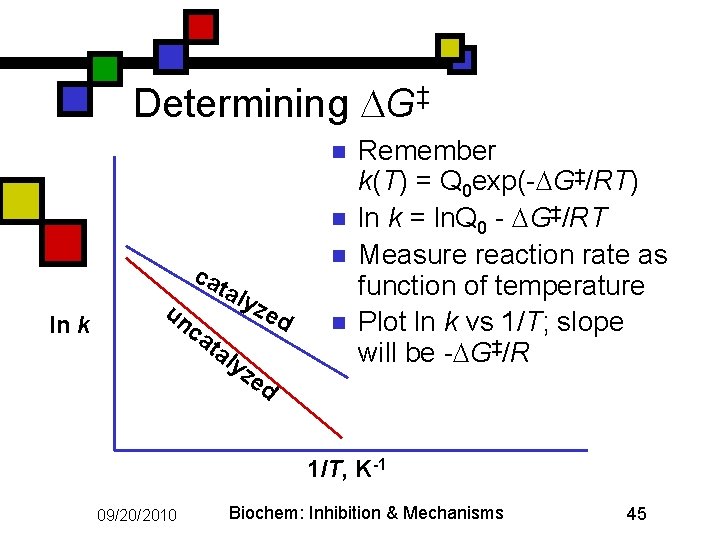
Determining G‡ n n ca ln k un ca n tal y ta ly ze d ze n Remember k(T) = Q 0 exp(- G‡/RT) ln k = ln. Q 0 - G‡/RT Measure reaction rate as function of temperature Plot ln k vs 1/T; slope will be - G‡/R d 1/T, K-1 09/20/2010 Biochem: Inhibition & Mechanisms 45

How enzymes alter G‡ n n Enzymes reduce G‡ by allowing the binding of the transition state into the active site Binding of the transition state needs to be tighter than the binding of either the reactants or the products. 09/20/2010 Biochem: Inhibition & Mechanisms 46
G‡ and Entropy n n n Effect is partly entropic: When a substrate binds, it loses a lot of entropy. Thus the entropic disadvantage of (say) a bimolecular reaction is soaked up in the process of binding the first of the two substrates into the enzyme's active site. 09/20/2010 Biochem: Inhibition & Mechanisms 47
Enthalpy and transition states n n Often an enthalpic component to the reduction in G‡ as well Ionic or hydrophobic interactions between the enzyme's active site residues and the components of the transition state make that transition state more stable. 09/20/2010 Biochem: Inhibition & Mechanisms 48
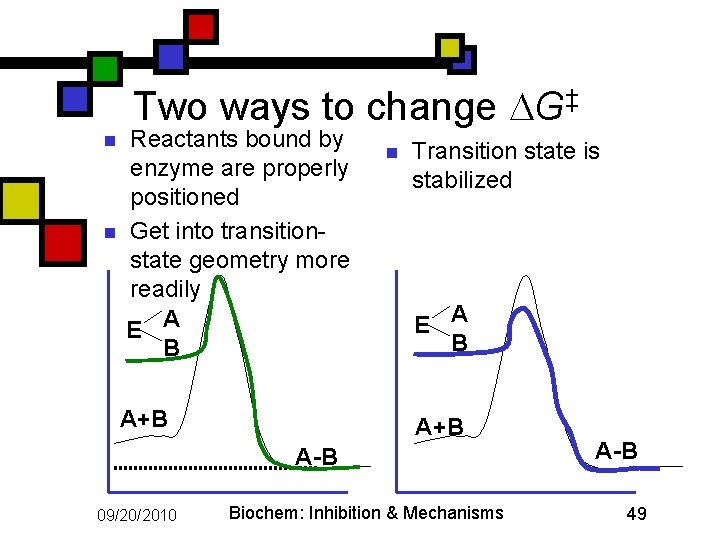
Two ways to change G‡ n n Reactants bound by enzyme are properly positioned Get into transitionstate geometry more readily E A B A+B n Transition state is stabilized E A B A+B A-B 09/20/2010 Biochem: Inhibition & Mechanisms A-B 49
How do enzymes reduce activation energies? n We can illustrate mechanistic principles by looking at specific examples; we can also recognize enzyme regulation when we see it. 09/20/2010 Biochem: Inhibition & Mechanisms 50
Examining enzyme mechanisms will help us understand catalysis n Examining general principles of catalytic activity and looking at specific cases will facilitate our appreciation of all enzymes. 09/20/2010 Biochem: Inhibition & Mechanisms 51

Binding modes: proximity n n We describe enzymatic mechanisms in terms of the binding modes of the substrates (or, more properly, the transition-state species) to the enzyme. One of these involves the proximity effect, in which two (or more) substrates are directed down potential-energy gradients to positions where they are close to one another. Thus the enzyme is able to defeat the entropic difficulty of bringing substrates together. 09/20/2010 Biochem: Inhibition & Mechanisms William Jencks 52
Binding modes: efficient transition-state binding n n Transition state fits even better (geometrically and electrostatically) in the active site than the substrate would. This improved fit lowers the energy of the transition-state system relative to the substrate. Best competitive inhibitors of an enzyme are those that resemble the transition state rather than the substrate or product. 09/20/2010 Biochem: Inhibition & Mechanisms 53

Diffusion-controlled reactions n n Some enzymes are so efficient that the limiting factor in completion of the reaction is diffusion of the substrates into the active site: These are diffusion-controlled reactions. Ultra-high turnover rates: kcat ~ 109 s-1. We can describe kcat / Km as catalytic efficiency of an enzyme. A diffusioncontrolled reaction will have a catalytic efficiency on the order of 108 M-1 s-1. 09/20/2010 Biochem: Inhibition & Mechanisms 54
Induced fit n n n Refinement on original Emil Fischer lock-and-key notion: both the substrate (or transitionstate) and the enzyme have flexibility Binding induces conformational changes 09/20/2010 Biochem: Inhibition & Mechanisms 55

Ionic reactions n n Define them as reactions that involve charged, or at least polar, intermediates Typically 2 reactants n n Electron rich (nucleophilic) reactant Electron poor (electrophilic) reactant Conventional to describe reaction as attack of nucleophile on electrophile Drawn with nucleophile donating electron(s) to electrophile 09/20/2010 Biochem: Inhibition & Mechanisms 56

Attack on Acyl Group n n n Transfer of an acyl group: section 14. 6 Nucleophile Y attacks carbonyl carbon, forming tetrahedral intermediate X- is leaving group 09/20/2010 Biochem: Inhibition & Mechanisms 57

Direct Displacement n n Attacking group adds to face of atom opposite to leaving group (scheme 6. 2) Transition state has five ligands; inherently less stable than scheme 6. 1 09/20/2010 Biochem: Inhibition & Mechanisms 58

Cleavage Reactions n Both electrons stay with one atom n n n Covalent bond produces carbanion: R 3—C—H R 3—C: - + H+ Covalent bond produces carbocation: R 3—C—H R 3—C+ + : H- One electron stays with each product n Both end up as radicals n R 1 O—OR 2 R 1 O • + • OR 2 Radicals are highly reactive— some more than others n 09/20/2010 Biochem: Inhibition & Mechanisms 59

Oxidation-Reduction Reactions n n Commonplace in biochemistry: EC 1 Oxidation is a loss of electrons Reduction is the gain of electrons In practice, often: n n n oxidation is decrease in # of C-H bonds; reduction is increase in # of C-H bonds Mnemonic: OIL RIG n n Oxidation is loss of electrons Reduction is gain of electrons 09/20/2010 Biochem: Inhibition & Mechanisms 60
Redox, continued n n n Intermediate electron acceptors and donors are organic moieties or metals Ultimate electron acceptor in aerobic organisms is usually dioxygen (O 2) Anaerobic organisms usually employ other electron acceptors 09/20/2010 Biochem: Inhibition & Mechanisms 61
Biological redox reactions n n n n Generally 2 -electron transformations Often involve alcohols, aldehydes, ketones, carboxylic acids, C=C bonds: R 1 R 2 CH-OH + X R 1 R 2 C=O + XH 2 R 1 HC=O + X + OH- R 1 COO- + XH 2 X is usually NAD, NADP, FAD, FMN A few biological redox systems involve metal ions or Fe-S complexes Usually reduced compounds are higher-energy than the corresponding oxidized compounds 09/20/2010 Biochem: Inhibition & Mechanisms 62
- Slides: 62