Environmental DNA Libraries METAGENOME Ju Hyoung Lim Nov

Environmental DNA Libraries METAGENOME Ju Hyoung Lim Nov 24, 2003

Definition of Metagenome Meta-. Gk. meta, of change, beyond. [Handelsman et al. (1998) Chem Biol 5: R 245 -R 249] • The genomes of the total microbiota found in nature • Environmental DNA comprising the genetic blueprints of entire microbial consoritia • The collective genomes directly cloned from all microorganisms present in a habitat at a certain time point • DNA libraries representing the genome of uncultured microbes, as a rich source for isolation of many novel genes

Historical background Pure cultured biota VS Total microbiota • The great plate count anomaly (Staley et al. 1985) • Loss of realistic prokaryotic biodiversity in culture method • Limitation of finding novel genes and protein when using culture method
Historical background 1. The great plate count anomaly (Staley et al. 1985) 1 g of Soil plate cultivation 4, 000 species 40 species • 1 g of soil contains up to 4, 000 different spp. but less than 1% are readily culturable w/ known cultivation method. • The culturability, i. e. the ability to grow as colonies on a rich culture medium is extremely low for natural populations of bacteria. • The reasons remain obscure but it is not difficult to understand that many organisms are not capable of adapting to the artificial and rather restrictive conditions of laboratory pure cultures.

Historical background 1. The great plate count anomaly (Staley et al. 1985) Total vs. cultivatable microbial diversity. Left, microbes on a soil flake were stained with DAPI (4’, 6 -Diamidino-2 -phenylindole) and detected using fluorescence microscopy. Right, colonies grown on enriched LB agar. There seems to be difference between two cases at 2 -3 orders of magnitude. (from Lorenz and Schleper, 2002, J Mol Catal, B Enzym 19 -20: 13)

Historical background 2. Loss of realistic prokaryotic biodiversity in culture method • Traditional cultivation method may result in the loss of major portions of the microbial communities • Even closesly related bacterial species need very different culture conditions. 97. 7% e ra M. l cu lep r be is os u. t M • Phylogenetic study with 16 S r. DNA amplification from environmental DNA (by Pace and colleagues in 1986) : revealed an astonishing number of new microbial groups that had never been picked up by cultivation.
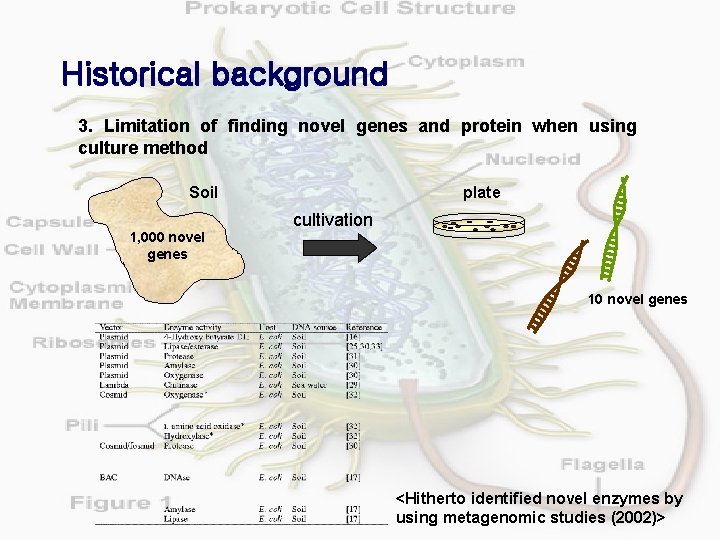
Historical background 3. Limitation of finding novel genes and protein when using culture method Soil 1, 000 novel genes plate cultivation 10 novel genes <Hitherto identified novel enzymes by using metagenomic studies (2002)>
Historical background • Torsvik and Goksoyr (1978) 1990’s to illustrate novel microbial diversity - Environmental DNA • Pace et al. (1986) - 16 S r. DNA from soil DNA • Schmidt et al. (1991) - 16 S r. DNA from aquatic DNA • Handelsman et al. (1998) - Metagenomic library TREND 0’s e 0 0 2 riev s et yme r to enz el v o n
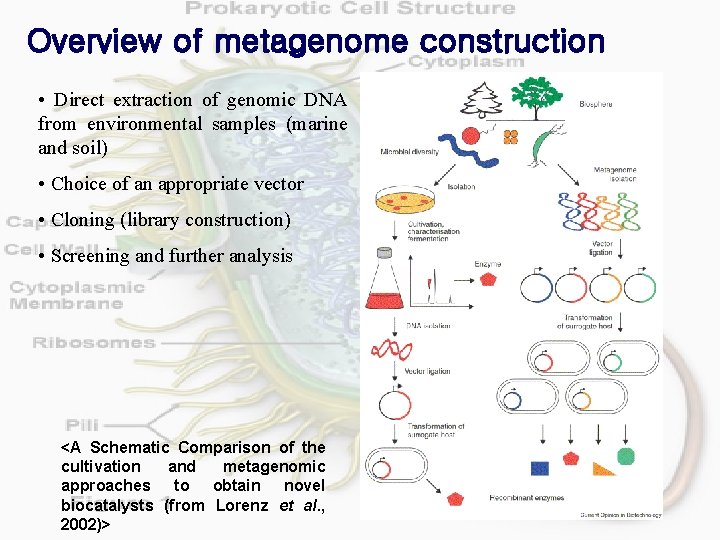
Overview of metagenome construction • Direct extraction of genomic DNA from environmental samples (marine and soil) • Choice of an appropriate vector • Cloning (library construction) • Screening and further analysis <A Schematic Comparison of the cultivation and metagenomic approaches to obtain novel biocatalysts (from Lorenz et al. , 2002)>

Metagenomic library construction • Isolation of environmental DNA • Choice of an appropriate vector • Cloning (library construction) • Screening and further analysis
Isolation of environmental DNA (soil) • Marine environment is easier to access than soil environment. • Up to 40 Kb or even more than 100 Kb DNA fragments have been cloned and they have served to characterize novel genomic fragments of abundant marine archaea and bacteria (the late 1990’s). • Soil environments is more difficult to prepare metagenomic libraries because of the presence of high MW inhibitor such as humic acid, fulvic acid, polyphenolic compounds in soil samples -> poor quality and quantity of DNA. Therefore numerous strategies were developed. Soil DNA extraction: basically falling into two ways… lysis Mechanical in situ extraction agitation lysis w/o mechanical agitation Physical separation ex situ extraction
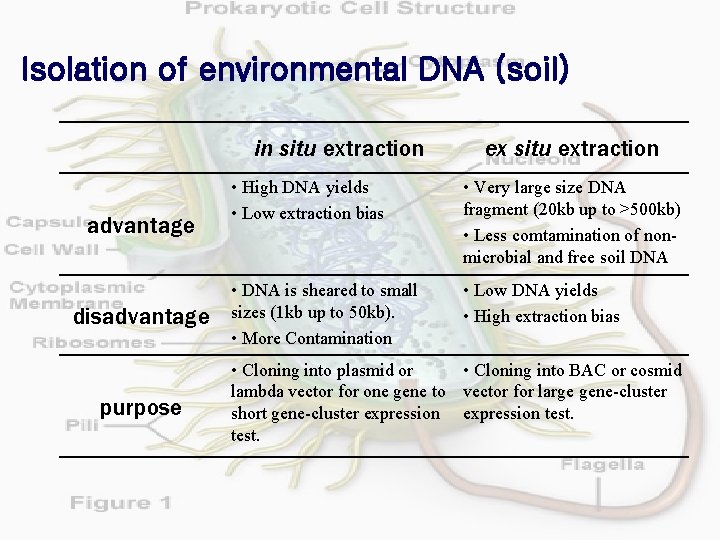
Isolation of environmental DNA (soil) in situ extraction advantage disadvantage purpose ex situ extraction • High DNA yields • Low extraction bias • Very large size DNA fragment (20 kb up to >500 kb) • Less comtamination of nonmicrobial and free soil DNA • DNA is sheared to small sizes (1 kb up to 50 kb). • More Contamination • Low DNA yields • High extraction bias • Cloning into plasmid or • Cloning into BAC or cosmid lambda vector for one gene to vector for large gene-cluster short gene-cluster expression test.
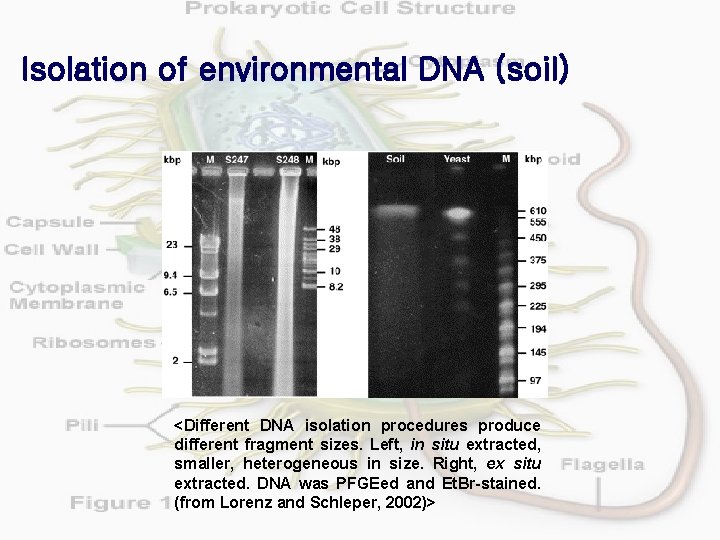
Isolation of environmental DNA (soil) <Different DNA isolation procedures produce different fragment sizes. Left, in situ extracted, smaller, heterogeneous in size. Right, ex situ extracted. DNA was PFGEed and Et. Br-stained. (from Lorenz and Schleper, 2002)>

Metagenomic library construction • Isolation of environmental DNA • Choice of an appropriate vector • Cloning (library construction) • Screening and further analysis
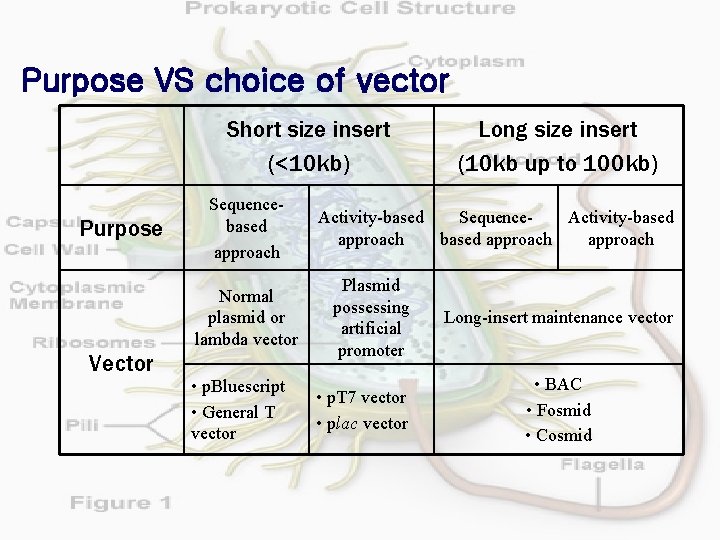
Purpose VS choice of vector Short size insert (<10 kb) Purpose Sequencebased approach Normal plasmid or lambda vector Vector • p. Bluescript • General T vector Long size insert (10 kb up to 100 kb) Activity-based Sequence. Activity-based approach Plasmid possessing artificial promoter • p. T 7 vector • plac vector Long-insert maintenance vector • BAC • Fosmid • Cosmid

Sequence-based approach • PCR-amplification for searching specific genes using conserved primer • Southern hybridization using conserved-sequence probe clone 2148 clone 25 - clones clone 124 + clones - It is rather conservative method, so the range of discovery could be narrow. Yet molecular diversity is so great that numerous novel enzymes can be retrieved in this way. - Long-insert cloning is better for this method - Vector: BAC, cosmid, fosmid, …>>…, plasmid, lambda vector

Sequence-based approach (continued) • Partial or full sequencing -> ORF assigning and blast <Physical map of a cosmid clone identified in oral metagenome (from Voget et al. 2003)> - useful to identify an entire gene clusters encoding mulifunctional modular enzyme e. g. polyketide synthases occur in clusters exceeding 100 kbp of contiguous DNA. - Vector: BAC, cosmid, fosmid, …>>…, plasmid, lambda vector • Advantage of sequence-based approach: - It overcomes possibilities of excluding some enzymes which are expressed at subthreshold level or not expressed in the heterogenous host. - Incomplete genes resulted from partial cloning can be detected.
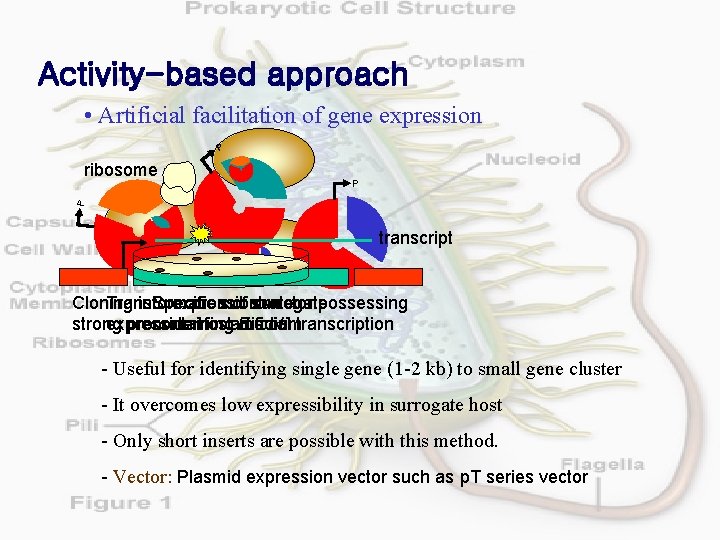
Activity-based approach • Artificial facilitation of gene expression P P P ribosome transcript Cloning Transformation into Specific expression substrate of surrogate vector possessing strong expression promoter containing host for artificial E. medium coli transcription - Useful for identifying single gene (1 -2 kb) to small gene cluster - It overcomes low expressibility in surrogate host - Only short inserts are possible with this method. - Vector: Plasmid expression vector such as p. T series vector

Activity-based approach (continued) • Gene expression from insert-borne promoters clone 25 Specific substrate containing medium - Useful for identifying single gene to large clusters - Eugenes, not pseudogenes, can be identified. - Problem of heterologous expression - Vector: BAC, cosmid, fosmid

History of BAC vector • Larger the insert DNA, more difficult maintenance of the clone - Extra energy for replication of so long non-chromosomal DNA - Deleterious recombination events • More copy number of vector, easier the loss of the extrachromosome • Normal plasmid ori can’t afford to very long stretch of replicon. YAC (yeast artificial chromosome) vector (Burke et al. Science 1987) • developed to maintain clones with large sizes (>500 kb) • However, fatal instability (YAC clone rescue failure) and chimera problems have been observed. BAC (bacteria artificial chromosome) vector (Shizuya et al. PNAS 1992) • capable of maintaining 1, 000 kb fragment. 1 -2 copies. • Insert DNA is stable, easy to manipulate, low recombination
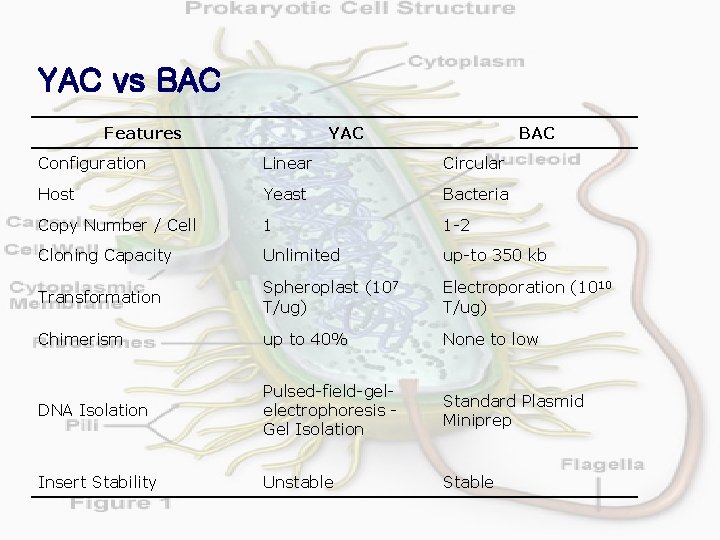
YAC vs BAC Features YAC BAC Configuration Linear Circular Host Yeast Bacteria Copy Number / Cell 1 1 -2 Cloning Capacity Unlimited up-to 350 kb Transformation Spheroplast (107 T/ug) Electroporation (1010 T/ug) Chimerism up to 40% None to low DNA Isolation Pulsed-field-gelelectrophoresis Gel Isolation Standard Plasmid Miniprep Insert Stability Unstable Stable
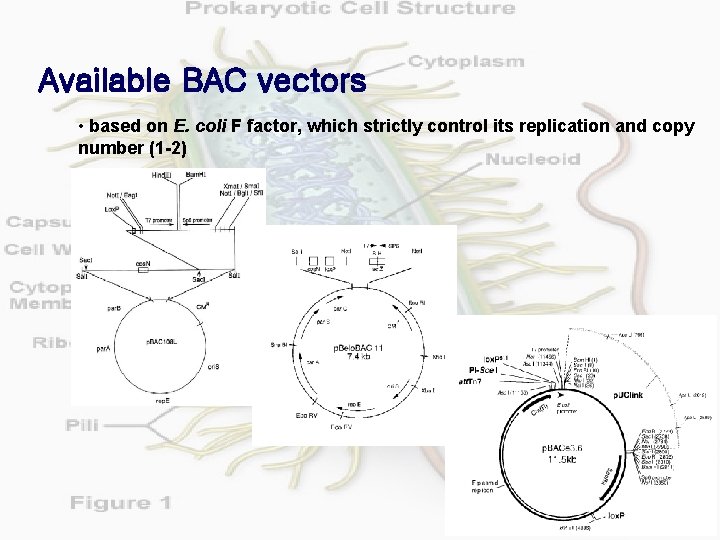
Available BAC vectors • based on E. coli F factor, which strictly control its replication and copy number (1 -2)

Available BAC vectors (continued) Name Cloning sites Recombinant selection p. BAC 108 L (6. 7 kb) Hind. III, Bam. HI no Shizuya et al. , 1992 p. Belo. BAC 11 (7. 4 kb) Hind. III, Bam. HI, Sph. I lac. Z Kim et al. , 1996 p. ECSBAC 4 (9. 3 kb) Eco. RI, Hind. III, Bam. HI lac. Z Frijters et al. , 1996 BIBAC 2 (23. 5 kb) Bam. HI sac. BII Plant Transformation via Agrobacterium Hamilton et al. , 1996 p. BACwich (11 kb) Hind. III, Bam. HI, Sph. I lac. Z Plant Transformation via Site-Specific Recombination Choi et al. , unpublished p. BACe 3. 6 (11. 5 kb) Bam. HI, Sac. II, Mlu. I, Eco. RI, Ava. III sac. BII High copy number is available de Jong et al. , unpublished p. Clasper (9. 7 kb) homologous recombination in yeast LEU 2 Yeast and bacteria shuttle vector Bradshaw et al. , 1995 Features Reference

Metagenomic library construction • Isolation of environmental DNA • Choice of an appropriate vector • Cloning (library construction) • Screening and further analysis

Cloning (library construction) • Insert preparation 1. Size fractionation - Pulse field gel electrophoresis (PFGE) & gel extraction of appropriatesized fragements - Restriction enzyme cutting or mechanical shearing 2. Polishing ends - Blunt ends formation (polymerase filling or nuclease trimming) - Sticky ends formation (restriction digest) - 3’ ends A-overhang (blud ends formation followed by Taq pol treatment) • Vector preparation - Restriction digest and 5’ end dephosphorylation

Cloning (library construction), continued • Ligation • Transformation - electroporation • Confirming insert clone, not self-ligate • Library grouping - Not 1 digestion (5’-GCGGCCGC-3’): high G+C group/ low G+C group - 16 S r. DNA amplifying and sequencing, restriction pattern, PCR fingerprint : metagenomic diversity • Transfer of the clones to 96 -well plates and conservation

Metagenomic library construction • Isolation of environmental DNA • Choice of an appropriate vector • Cloning (library construction) • Screening and further analysis
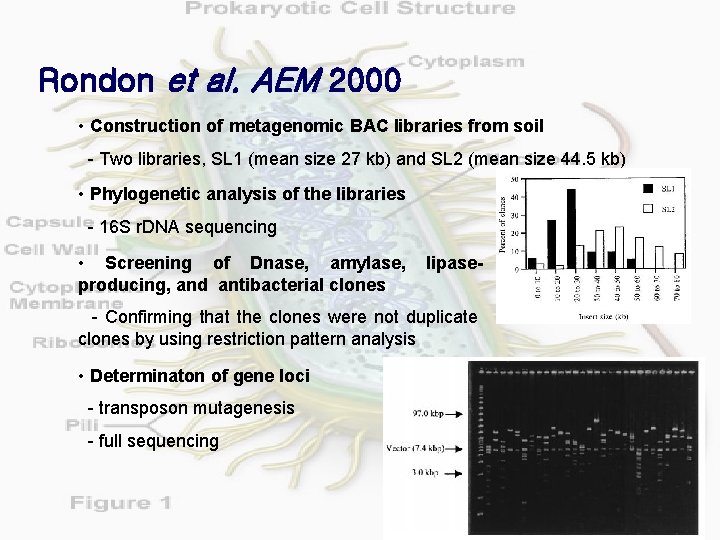
Rondon et al. AEM 2000 • Construction of metagenomic BAC libraries from soil - Two libraries, SL 1 (mean size 27 kb) and SL 2 (mean size 44. 5 kb) • Phylogenetic analysis of the libraries - 16 S r. DNA sequencing • Screening of Dnase, amylase, producing, and antibacterial clones lipase- - Confirming that the clones were not duplicate clones by using restriction pattern analysis • Determinaton of gene loci - transposon mutagenesis - full sequencing

Voget et al. AEM 2003 • Laboratory enrichment of soil microbiota showing agarolytic activities - for increasing cloning efficiency and fast isolation of specific metagenome • Construction of cosmid DNA libraries - Insert DNA was partially digested by Sau 3 A and cloned into cosmid p. WE 15 • Full sequencing and ORF assignment of amidase, two cellulases, αamylase, and pectate lyase genes • PCR cloning of entire ORFs into expression vector • Enzyme activities of protein extracts

Diaz-Torres et al. Antimicrob Agents Chemother 2003 • Oral metagenome was constructed from dental plaques and saliva - Insert DNA was sheared by ultrasonication. - Mung bean nuclease treated inserts were electrophoresed. - 800 to 3, 000 bp sized fragments were recovered and 3’ adenylated by Taq pol incubation. - Prepared inserts were cloned into TA expression vector. • Of 450 transformants, 18 (4%) colonies were screened on LB medium as tetracyclin resistant clones • Confirming that if the resistance is derived from previous known mechanism. PCR using tet gene primers.

Perspectives in metagenomic study • The microbial world seems to offer the greatest natural resource of molecular diversity. Classical cultivation method is valid and powerful but severely restricted in scope. So, it needs to be complemented by metagenomic study. • Yet methodological problems will set the limits of this approach: Heterologous gene expression in surrogate host is not always successful. • Using a variety of different hosts e. g. Streptomyces lividans and Bacillus sp. in addition to E. coli should significantly boost the success rate in heterologous expression screens
INDEX • • Definition of metagenome Historical background Overview of metagenome construction Isolation of environmental DNA Choice of an appropriate vector Cloning (library construction) Screening and further analysis Perspectived in metagenomic study
- Slides: 32