Environmental Data Analysis with Mat Lab Lecture 8

Environmental Data Analysis with Mat. Lab Lecture 8: Solving Generalized Least Squares Problems

SYLLABUS Lecture 01 Lecture 02 Lecture 03 Lecture 04 Lecture 05 Lecture 06 Lecture 07 Lecture 08 Lecture 09 Lecture 10 Lecture 11 Lecture 12 Lecture 13 Lecture 14 Lecture 15 Lecture 16 Lecture 17 Lecture 18 Lecture 19 Lecture 20 Lecture 21 Lecture 22 Lecture 23 Lecture 24 Using Mat. Lab Looking At Data Probability and Measurement Error Multivariate Distributions Linear Models The Principle of Least Squares Prior Information Solving Generalized Least Squares Problems Fourier Series Complex Fourier Series Lessons Learned from the Fourier Transform Power Spectra Filter Theory Applications of Filters Factor Analysis Orthogonal functions Covariance and Autocorrelation Cross-correlation Smoothing, Correlation and Spectra Coherence; Tapering and Spectral Analysis Interpolation Hypothesis testing Hypothesis Testing continued; F-Tests Confidence Limits of Spectra, Bootstraps

purpose of the lecture use prior information to solve exemplary problems

review of last lecture
![failure-proof least-squares add information to the problem that guarantees that matrices like [GTG] are failure-proof least-squares add information to the problem that guarantees that matrices like [GTG] are](http://slidetodoc.com/presentation_image_h2/b8c32f0d303045b178e8da07495f975b/image-5.jpg)
failure-proof least-squares add information to the problem that guarantees that matrices like [GTG] are never singular such information is called prior information

examples of prior information soil has density will be around 1500 kg/m 3 give or take 500 or so chemical components sum to 100% pollutant transport is subject to the diffusion equation water in rivers always flows downhill

linear prior information with covariance Ch
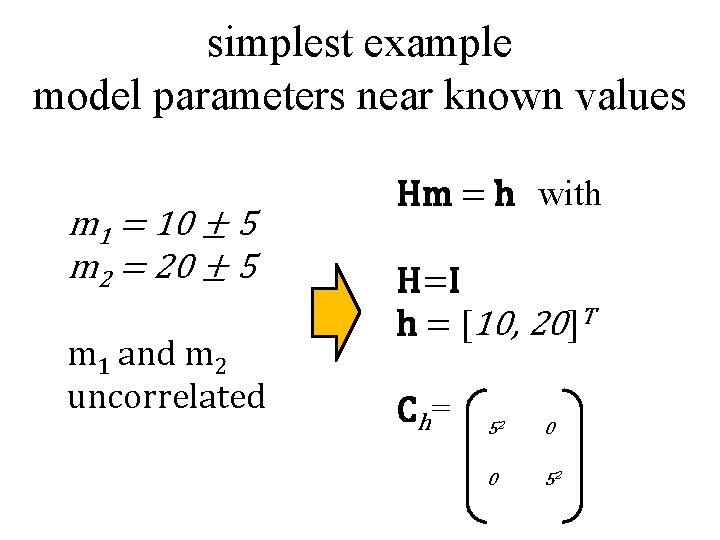
simplest example model parameters near known values m 1 = 10 ± 5 m 2 = 20 ± 5 m 1 and m 2 uncorrelated Hm = h with H=I h = [10, 20]T Ch= 52 0 0 52
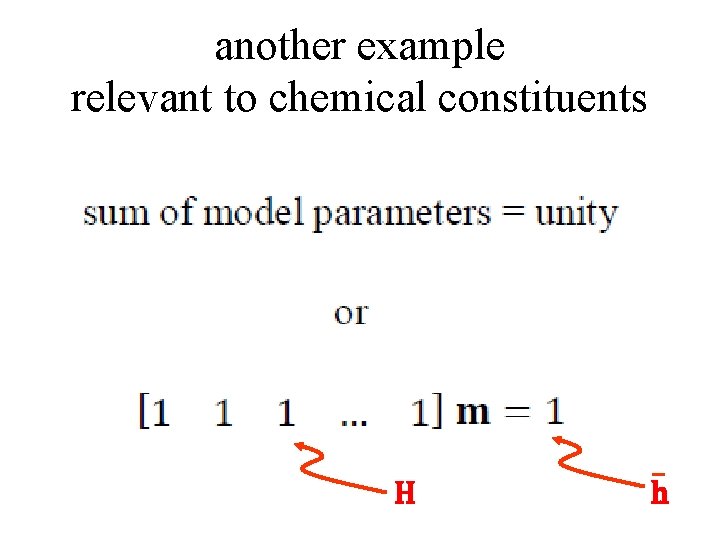
another example relevant to chemical constituents H h

use Normal p. d. f. to represent prior information

Normal p. d. f. defines an “error in prior information” individual errors weighted by their certainty

now suppose that we observe some data: d = dobs with covariance Cd

represent the observations with a Normal p. d. f. p(d) = observations mean of data predicted by the model

this Normal p. d. f. defines an “error in data” prediction error weighted by its certainty

Generalized Principle of Least Squares the best mest is the one that minimizes the total error with respect to m justified by Bayes Theorem in the last lecture

generalized least squares solution pattern same as ordinary least squares … … but with more complicated matrices

(new material) How to use the Generalized Least Squares Equations
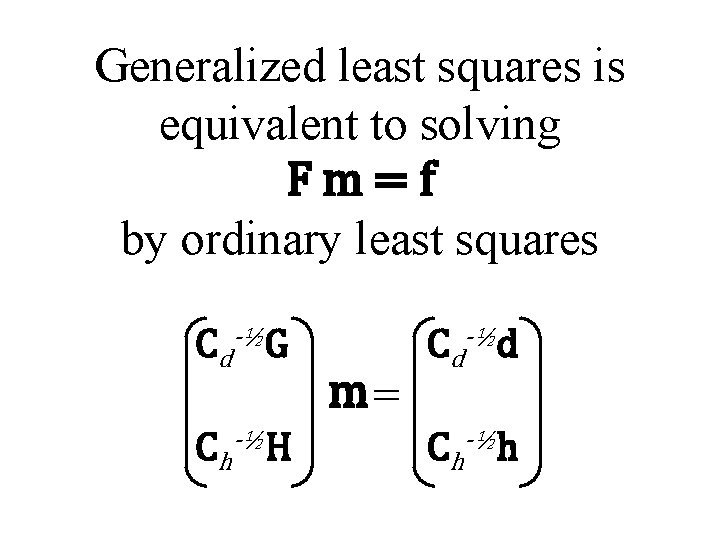
Generalized least squares is equivalent to solving Fm=f by ordinary least squares Cd-½G Ch-½H m= Cd-½d Ch-½h
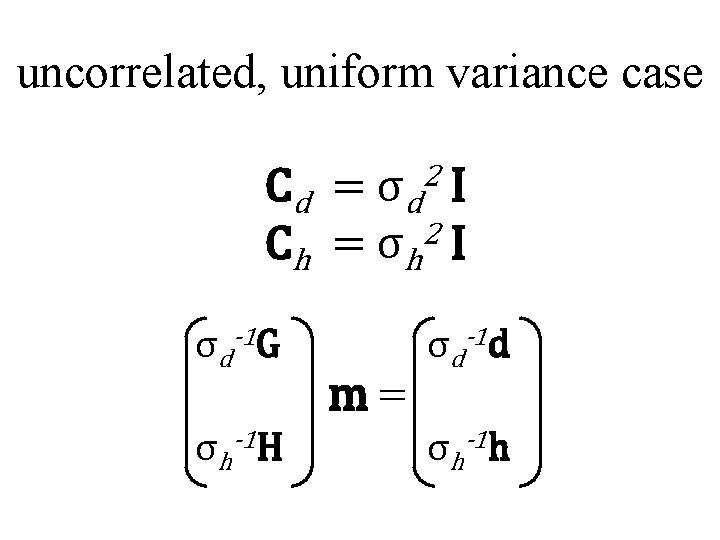
uncorrelated, uniform variance case 2 Cd = σd I C h = σ h 2 I σd-1 G σh-1 H m= σd-1 d σh-1 h
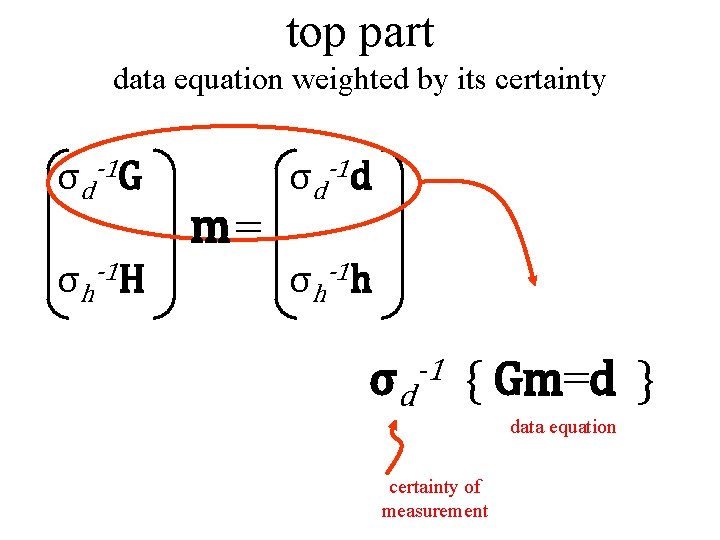
top part data equation weighted by its certainty σd-1 G σh-1 H m= σd-1 d σh-1 h σd -1 { Gm=d } data equation certainty of measurement
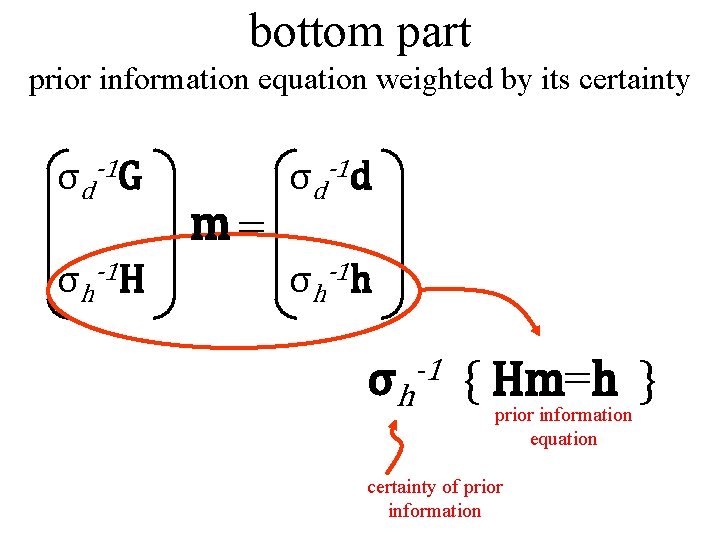
bottom part prior information equation weighted by its certainty σd-1 G σh-1 H m= σd-1 d σh-1 h σh -1 { Hm=h } prior information equation certainty of prior information
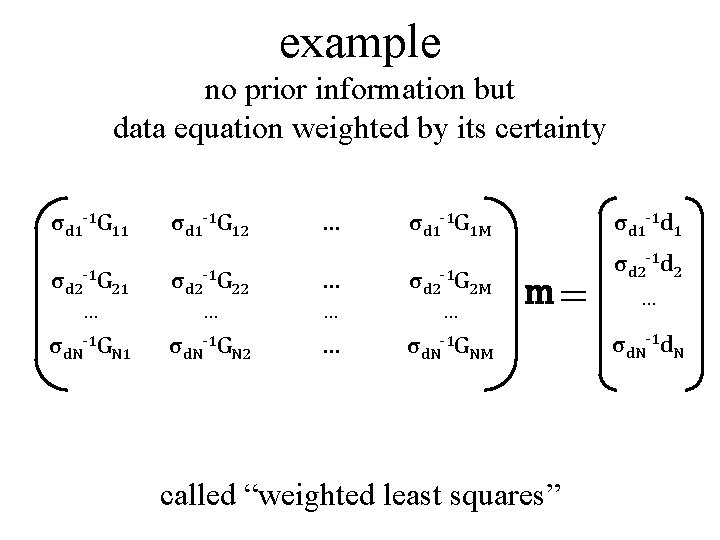
example no prior information but data equation weighted by its certainty σd 1 -1 G 11 σd 1 -1 G 12 … σd 1 -1 G 1 M σd 2 -1 G 21 σd 2 -1 G 22 … σd 2 -1 G 2 M … … σd. N-1 GN 1 σd. N-1 GN 2 … σd. N-1 GNM σd 1 -1 d 1 m= called “weighted least squares” σd 2 -1 d 2 … σd. N-1 d. N
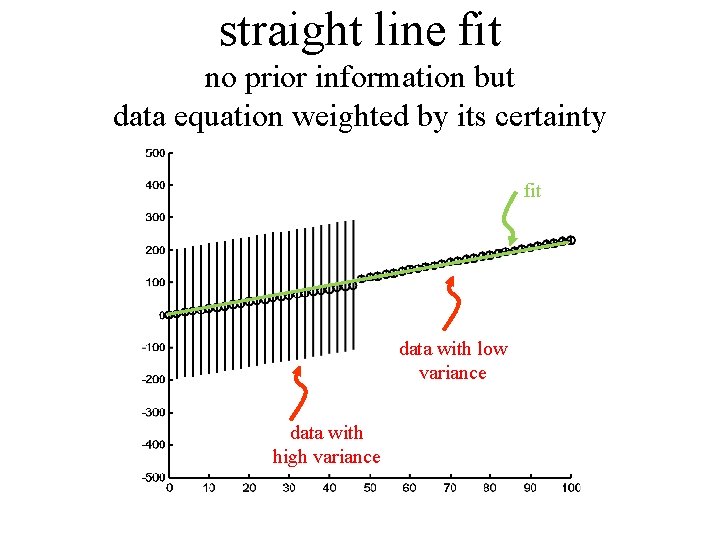
straight line fit no prior information but data equation weighted by its certainty fit data with low variance data with high variance
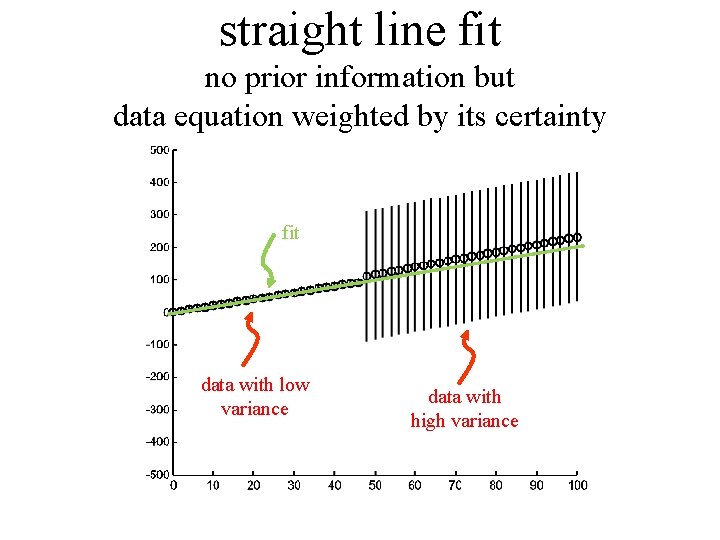
straight line fit no prior information but data equation weighted by its certainty fit data with low variance data with high variance

another example prior information that the model parameters are small m≈0 H=I h=0 assume uncorrelated with uniform variances Cd = σ d 2 I C h = σ h 2 I
![Fm=h σd-1 G σh-1 I m= σd-1 d σh-10 [FTF]-1 FTm=f m=[GTG + ε Fm=h σd-1 G σh-1 I m= σd-1 d σh-10 [FTF]-1 FTm=f m=[GTG + ε](http://slidetodoc.com/presentation_image_h2/b8c32f0d303045b178e8da07495f975b/image-26.jpg)
Fm=h σd-1 G σh-1 I m= σd-1 d σh-10 [FTF]-1 FTm=f m=[GTG + ε 2 I]-1 GTd with ε= σd/σm
![called “damped least squares” m=[GTG + ε 2 I]-1 GTd with ε= σd/σm ε=0: called “damped least squares” m=[GTG + ε 2 I]-1 GTd with ε= σd/σm ε=0:](http://slidetodoc.com/presentation_image_h2/b8c32f0d303045b178e8da07495f975b/image-27.jpg)
called “damped least squares” m=[GTG + ε 2 I]-1 GTd with ε= σd/σm ε=0: minimize the prediction error ε→∞: minimize the size of the model parameters 0<ε<∞: minimize a combination of the two
![m=[GTG + ε 2 I]-1 GTd with ε= σd/σm advantages: really easy to code m=[GTG + ε 2 I]-1 GTd with ε= σd/σm advantages: really easy to code](http://slidetodoc.com/presentation_image_h2/b8c32f0d303045b178e8da07495f975b/image-28.jpg)
m=[GTG + ε 2 I]-1 GTd with ε= σd/σm advantages: really easy to code mest = (G’*G+(e^2)*eye(M))(G’*d); always works disadvantages: often need to determine ε empirically prior information that the model parameters are small not always sensible

smoothness as prior information

model parameters represent the values of a function m(x) at equally spaced increments along the x-axis

function approximated by its values at a sequence of x’s mi m(x) mi+1 xi xi+1 Δx m(x) → m=[m 1, m 2, m 3, …, m. M]T x

rough function has large second derivative a smooth function is one that is not rough a smooth function has a small second derivative

approximate expressions for second derivative
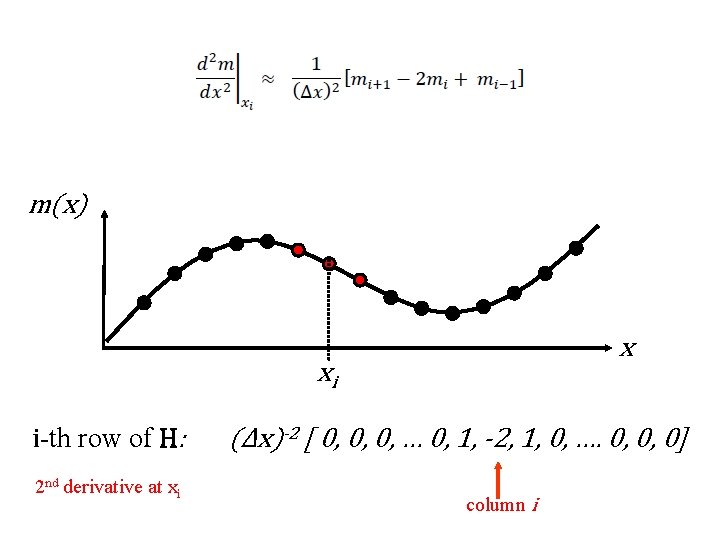
m(x) x xi i-th row of H: 2 nd derivative at xi (Δx)-2 [ 0, 0, 0, … 0, 1, -2, 1, 0, …. 0, 0, 0] column i

what to do about m 1 and m. M? not enough points for 2 nd derivative two possibilities no prior information for m 1 and m. M or prior information about flatness (first derivative)
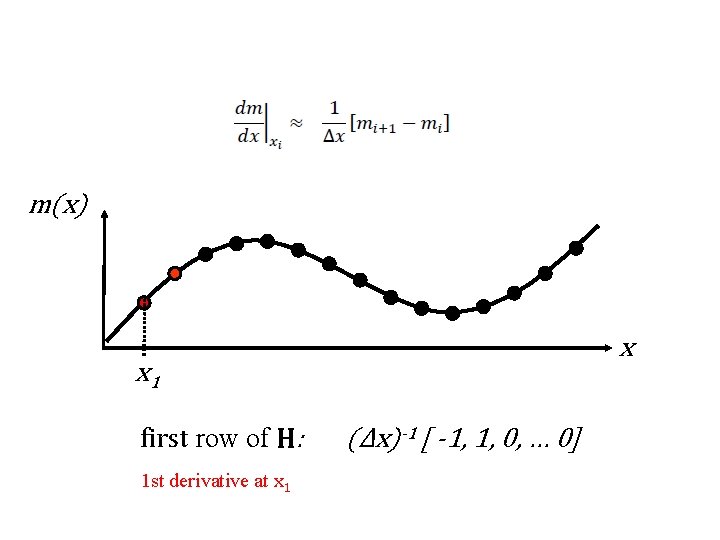
m(x) x x 1 first row of H: 1 st derivative at x 1 (Δx)-1 [ -1, 1, 0, … 0]

“smooth interior” / “flat ends” version of Hm=h h=0
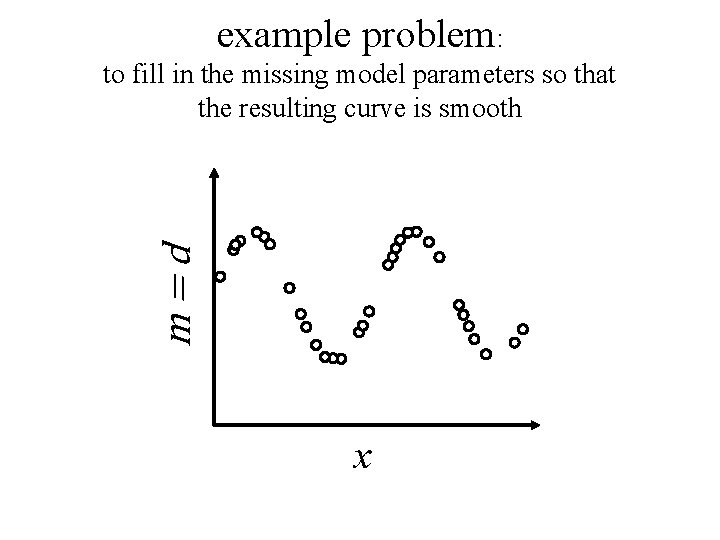
example problem: m=d to fill in the missing model parameters so that the resulting curve is smooth x
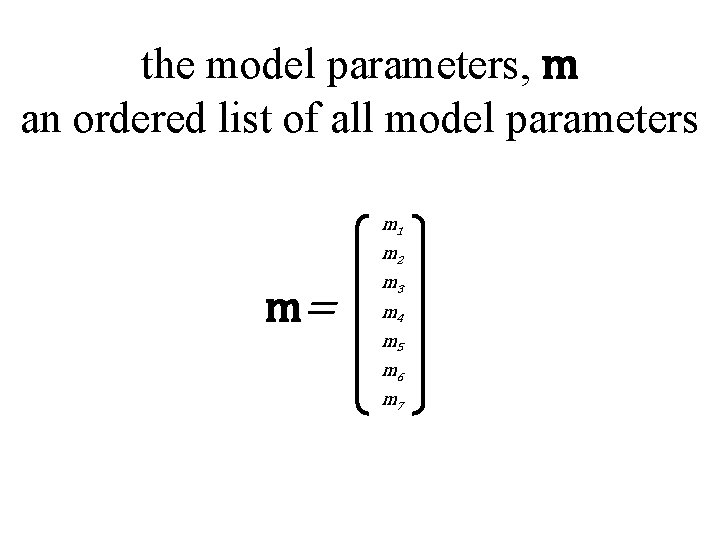
the model parameters, m an ordered list of all model parameters m 1 m 2 m= m 3 m 4 m 5 m 6 m 7
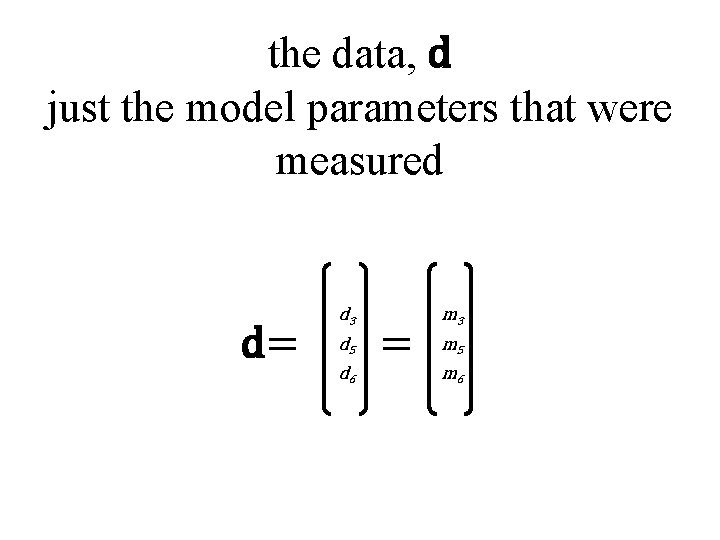
the data, d just the model parameters that were measured d= d 3 d 5 d 6 = m 3 m 5 m 6
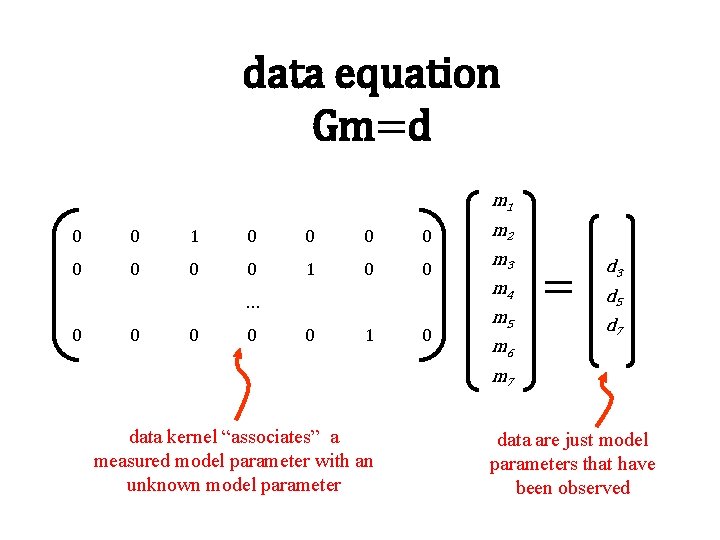
data equation Gm=d m 1 0 0 0 0 m 2 0 0 1 0 0 m 3 … 0 0 0 1 0 m 4 m 5 m 6 = d 3 d 5 d 7 m 7 data kernel “associates” a measured model parameter with an unknown model parameter data are just model parameters that have been observed

The prior information equation, Hm=h “smooth interior” / “flat ends” h=0
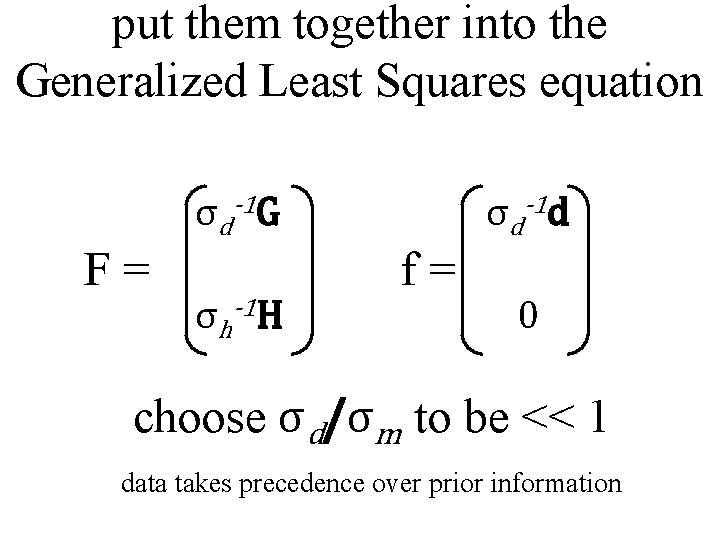
put them together into the Generalized Least Squares equation σd-1 G F= σh-1 H σd-1 d f= 0 choose σd/σm to be << 1 data takes precedence over prior information

the solution using Mat. Lab
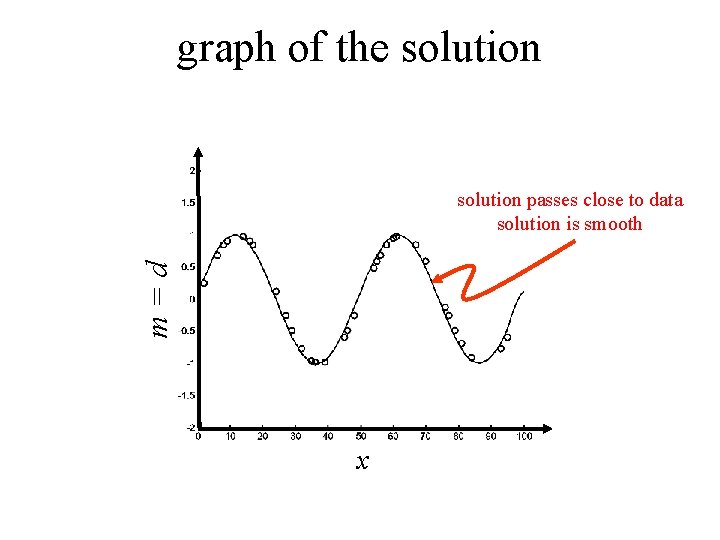
graph of the solution m=d solution passes close to data solution is smooth x

Two Mat. Lab issues Issue 1: matrices like G and F can be quite big, but contain mostly zeros. Solution 1: Use “sparse matrices” which don’t store the zeros Issue 2: matrices like GTG and FTF are not as sparse as G and F Solution 2: Solve equation by a method, such as “biconjugate gradients” that doesn’t require the calculation of GTG and FTF

Using “sparse matrices” which don’t store the zeros: N=200000; M=100000; F=spalloc(N, M, 3*M); “sparse allocate” creates a 200000× 100000 matrix that can hold up to 300000 non-zero elements. note that an ordinary matrix would have 20, 000, 000 elements

Once allocated, sparse matrices are used just like ordinary matrices … … they just consume less memory.

Issue 2: Use biconjugate gradient solver to avoid calculating GTG and FTF Suppose that we want to solve FTF m = FTf The standard way would be: mest = (F’F)(F’f); but that requires that we compute F’F

a “biconjugate gradient” solver requires only that we be able to multiply a vector, v, by GTG, where the solver supplies the vector, v. so we have to calculate y=GTG v the trick is to calculate t=Gv first, and then calculate y=G’t this is done in a Matlab function, afun()

function y = afun(v, transp_flag) global F; t = F*v; ignore this variable; its y = F'*t; never used return

the bicg()solver is passed a “handle” to this function so, the new way of solving the generalized inverse problem is: clear F; put at the top of the Mat. Lab script global F; … mest=bicg(@afun, F'*f, 1 e-10, 3*L); for “biconjugate” “handle” to the function

tolerance maximum number of iterations mest=bicg(@afun, F'*f, 1 e-10, Niter); for “biconjugate” r. h. s of equation FTFm=FTf “handle” to the multiply function The solution is by iterative improvement of an initial guess. The iterations stop when the tolerance falls beneath the specified level (good) or, regardless, when the maximum number of iterations is reached (bad).

example of a large problem fill in the missing model parameters that represents a 2 D function m(x, y) so that the function passes through measured data points m(xi, yi) = di and the function satisfies the diffusion equation d 2 m/dx 2 + d 2 m/dy 2 = 0
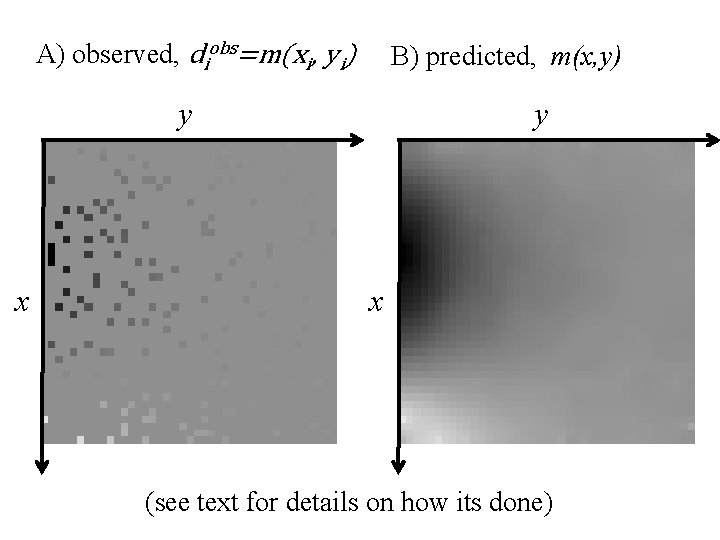
A) observed, diobs=m(xi, yi) B) predicted, m(x, y) y x (see text for details on how its done)
- Slides: 55