Ensemble Results of PIM 1 15 6 PIM
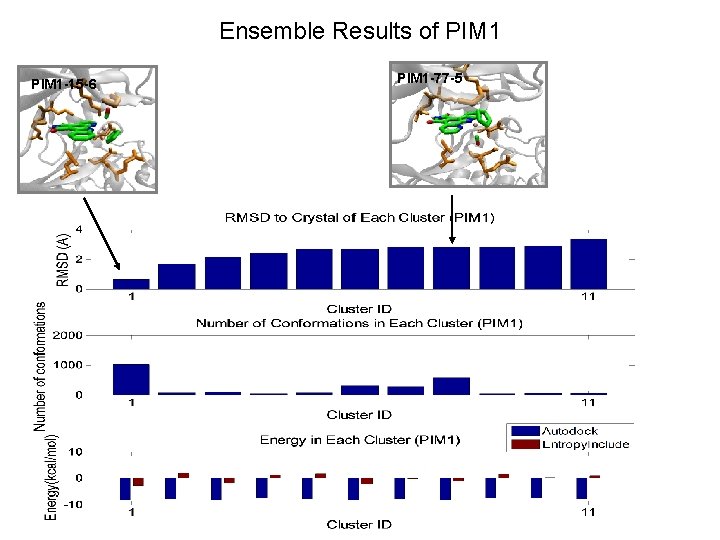
Ensemble Results of PIM 1 -15 -6 PIM 1 -77 -5
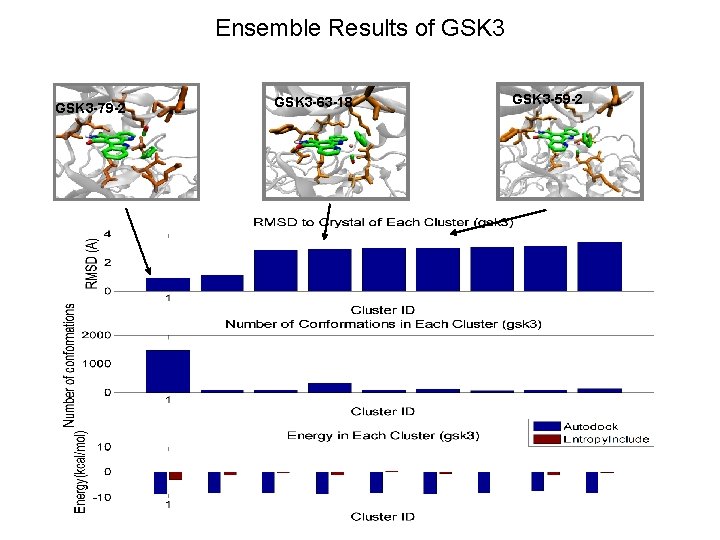
Ensemble Results of GSK 3 -79 -2 GSK 3 -63 -18 GSK 3 -59 -2
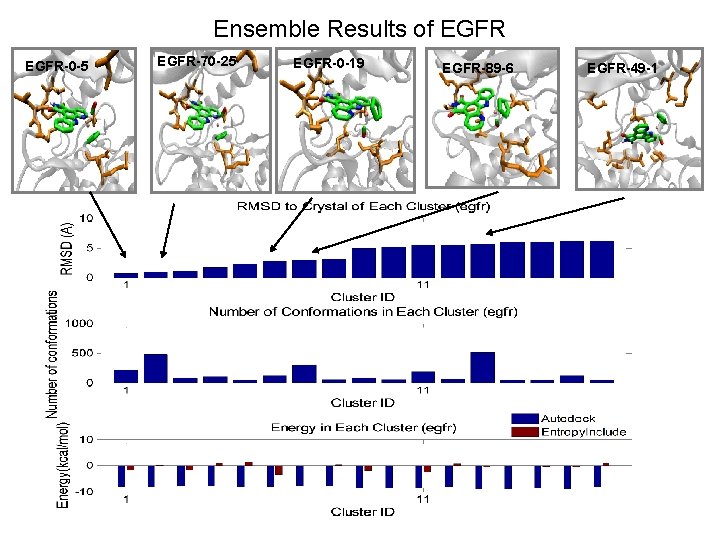
Ensemble Results of EGFR-0 -5 EGFR-70 -25 EGFR-0 -19 EGFR-89 -6 EGFR-49 -1
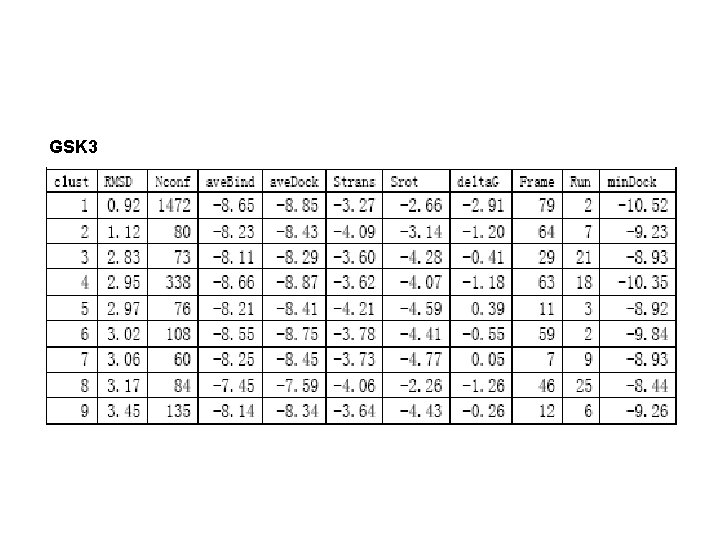
GSK 3
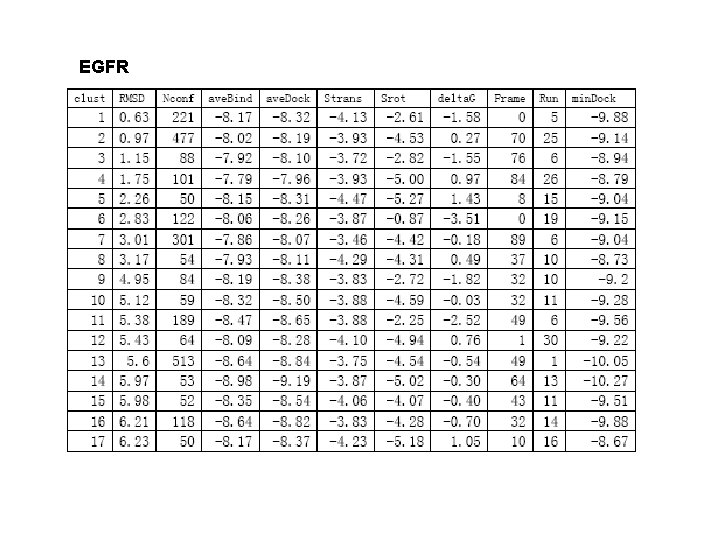
EGFR
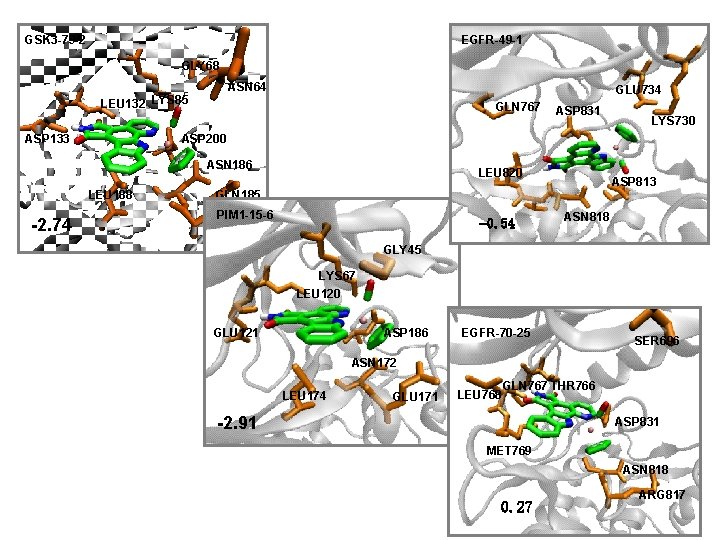
GSK 3 -79 -2 EGFR-49 -1 GLY 68 ASN 64 LEU 132 LYS 85 ASP 133 GLU 734 GLN 767 LYS 730 ASP 200 ASN 186 LEU 188 -2. 74 ASP 831 LEU 820 ASP 813 GLN 185 PIM 1 -15 -6 -0. 54 ASN 818 GLY 45 LYS 67 LEU 120 GLU 121 ASP 186 EGFR-70 -25 SER 696 ASN 172 LEU 174 GLU 171 LEU 768 GLN 767 THR 766 -2. 91 ASP 831 MET 769 ASN 818 0. 27 ARG 817
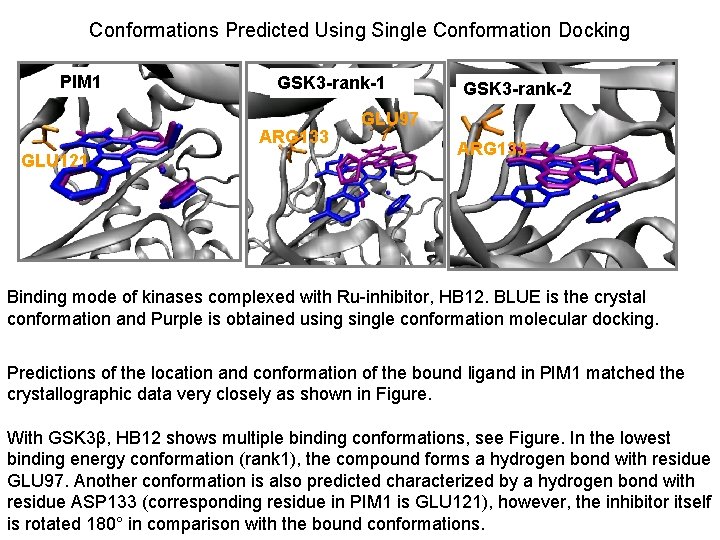
Conformations Predicted Using Single Conformation Docking PIM 1 GSK 3 -rank-1 ARG 133 GLU 121 GSK 3 -rank-2 GLU 97 ARG 133 Binding mode of kinases complexed with Ru-inhibitor, HB 12. BLUE is the crystal conformation and Purple is obtained usingle conformation molecular docking. Predictions of the location and conformation of the bound ligand in PIM 1 matched the crystallographic data very closely as shown in Figure. With GSK 3β, HB 12 shows multiple binding conformations, see Figure. In the lowest binding energy conformation (rank 1), the compound forms a hydrogen bond with residue GLU 97. Another conformation is also predicted characterized by a hydrogen bond with residue ASP 133 (corresponding residue in PIM 1 is GLU 121), however, the inhibitor itself is rotated 180° in comparison with the bound conformations.
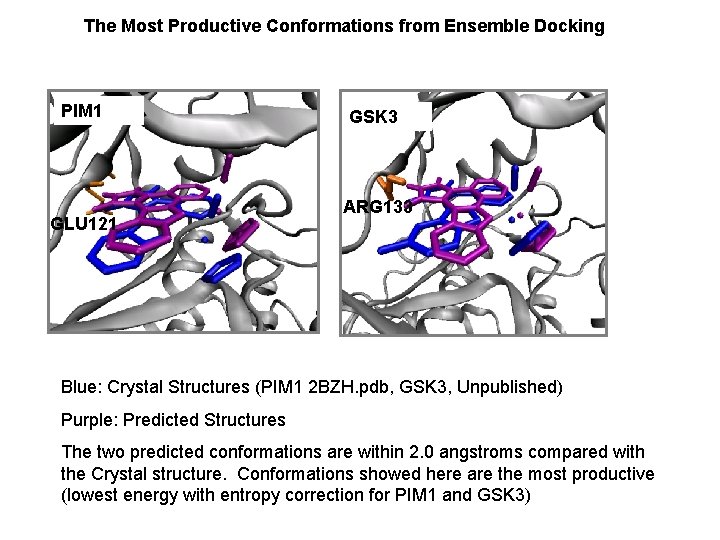
The Most Productive Conformations from Ensemble Docking PIM 1 GLU 121 GSK 3 ARG 133 Blue: Crystal Structures (PIM 1 2 BZH. pdb, GSK 3, Unpublished) Purple: Predicted Structures The two predicted conformations are within 2. 0 angstroms compared with the Crystal structure. Conformations showed here are the most productive (lowest energy with entropy correction for PIM 1 and GSK 3)
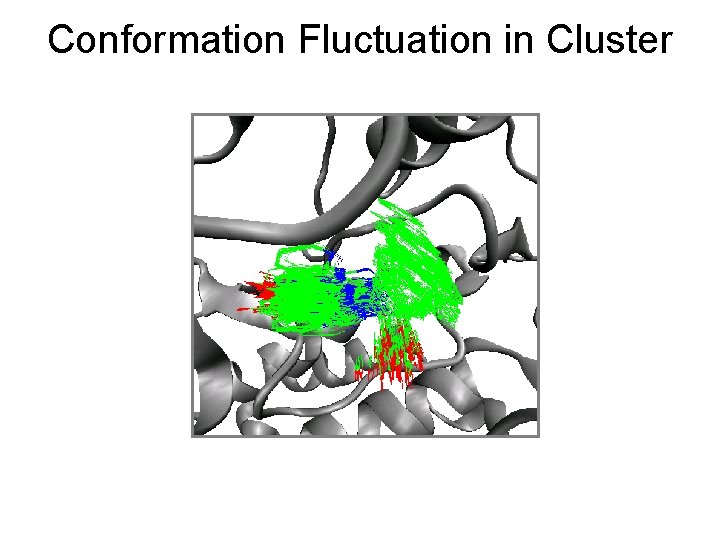
Conformation Fluctuation in Cluster

• Slices after this are backup slices which show the conformations in detail.
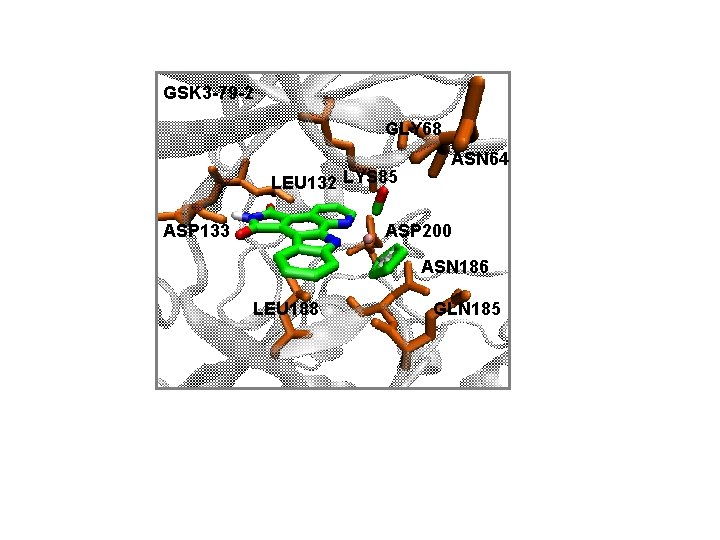
GSK 3 -79 -2 GLY 68 LEU 132 LYS 85 ASP 133 ASN 64 ASP 200 ASN 186 LEU 188 GLN 185
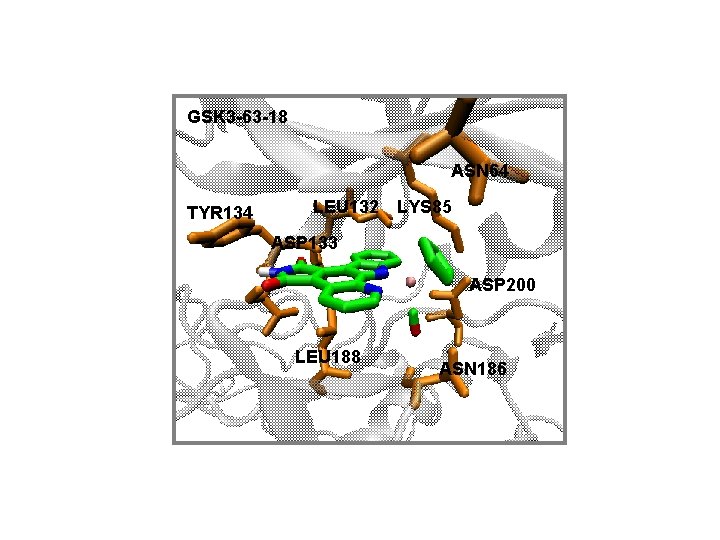
GSK 3 -63 -18 ASN 64 TYR 134 LEU 132 LYS 85 ASP 133 ASP 200 LEU 188 ASN 186
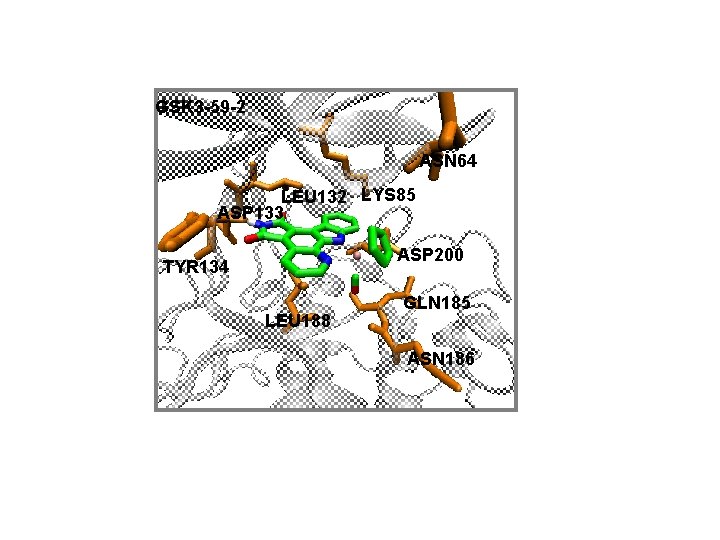
GSK 3 -59 -2 ASN 64 LEU 132 LYS 85 ASP 133 ASP 200 TYR 134 LEU 188 GLN 185 ASN 186
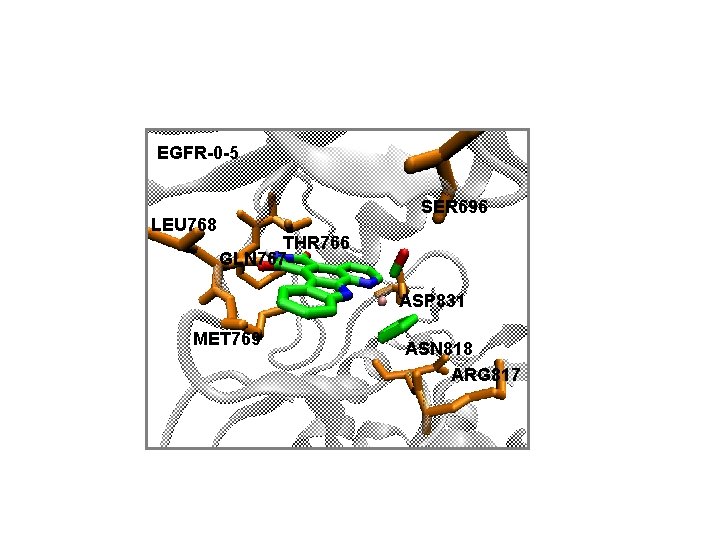
EGFR-0 -5 LEU 768 SER 696 THR 766 GLN 767 ASP 831 MET 769 ASN 818 ARG 817
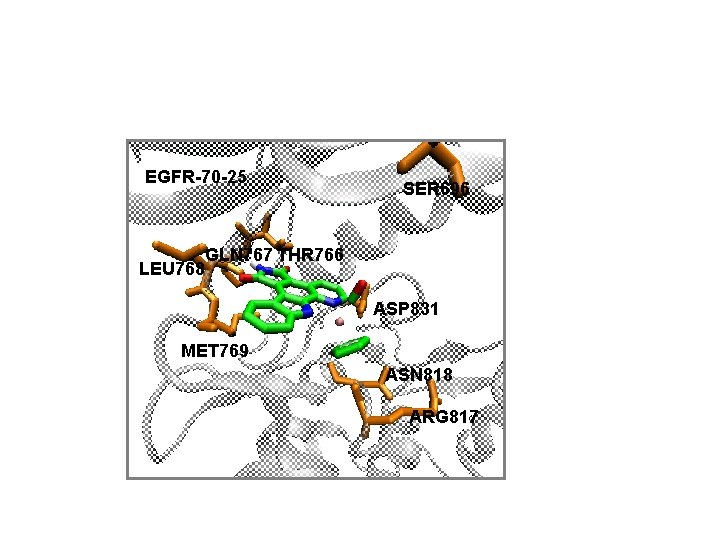
EGFR-70 -25 LEU 768 SER 696 GLN 767 THR 766 ASP 831 MET 769 ASN 818 ARG 817
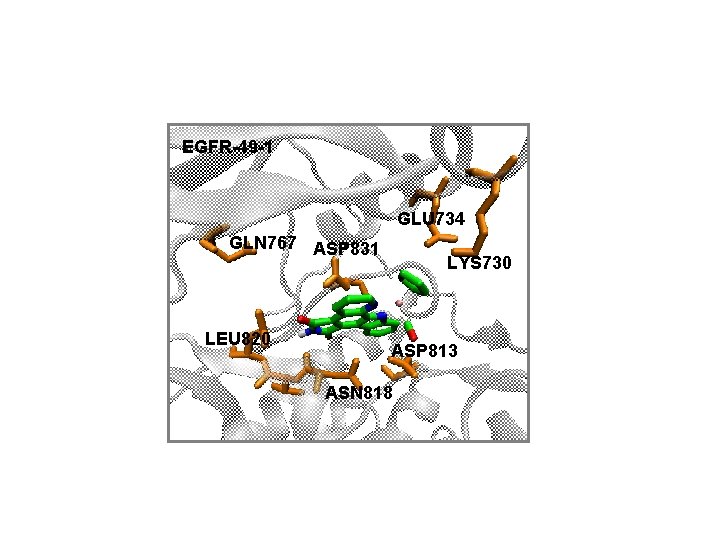
EGFR-49 -1 GLU 734 GLN 767 ASP 831 LEU 820 LYS 730 ASP 813 ASN 818
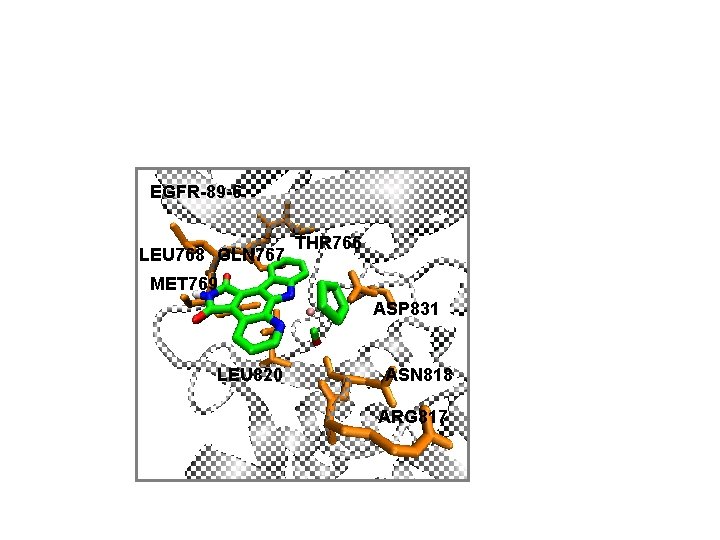
EGFR-89 -6 LEU 768 GLN 767 THR 766 MET 769 ASP 831 LEU 820 ASN 818 ARG 817
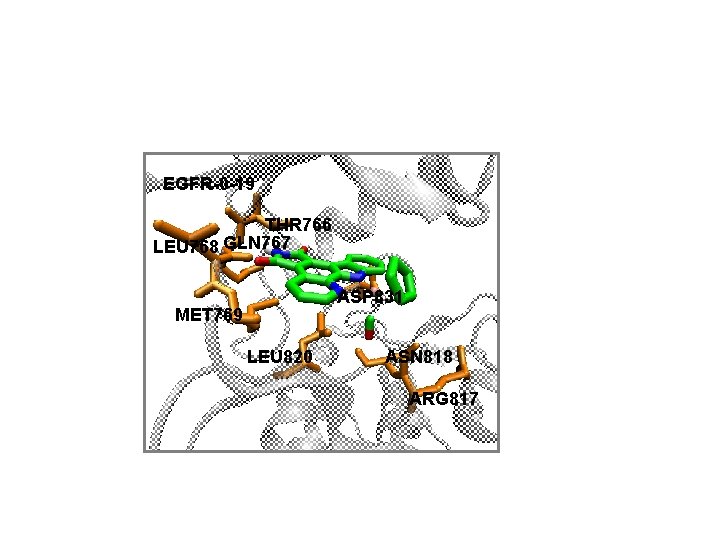
EGFR-0 -19 THR 766 LEU 768 GLN 767 ASP 831 MET 769 LEU 820 ASN 818 ARG 817
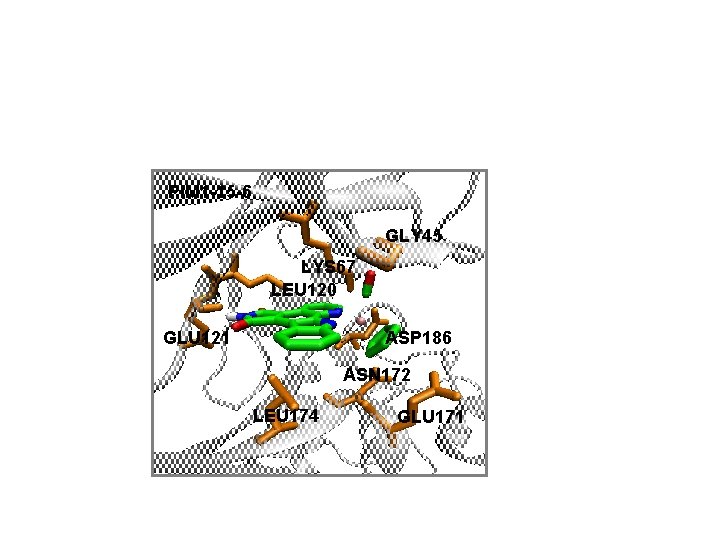
PIM 1 -15 -6 GLY 45 LYS 67 LEU 120 GLU 121 ASP 186 ASN 172 LEU 174 GLU 171
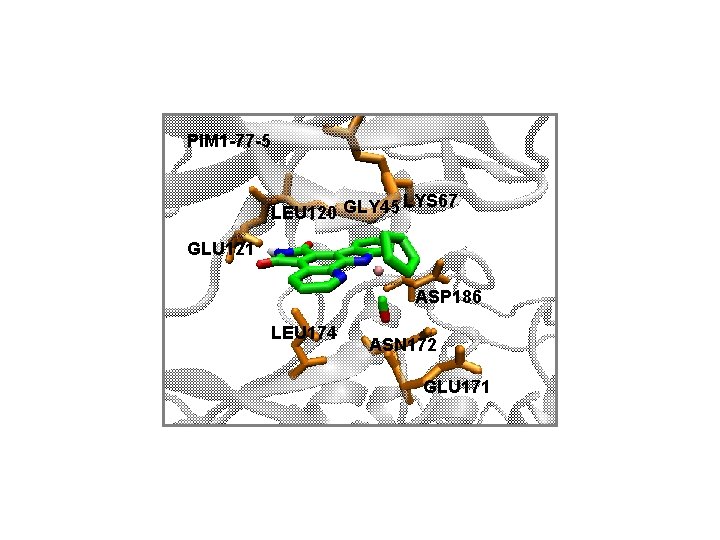
PIM 1 -77 -5 LYS 67 LEU 120 GLY 45 GLU 121 ASP 186 LEU 174 ASN 172 GLU 171
- Slides: 21