Engineering SelfModelling Systems Application to Biology Carole Bernon

Engineering Self-Modelling Systems: Application to Biology Carole Bernon, Davy Capera*, Jean-Pierre Mano SMAC Team (Cooperative Multi-Agent Systems) Institut de Recherche en Informatique de Toulouse *UPEtec www. irit. fr/SMAC - www. upetec. fr 20 janvier 2005

Outline v. Making complex systems self-build Self-organisation by cooperation Four-layer model v. A domain of application: Biology micro. Mega specific case Agents and Biology v. Model applied to micro. Mega Architecture o Agents o Behaviours Preliminary results v. Conclusion Engineering Self-Modelling Systems: Application to Biology 2

Statement v. Systems: more and more complex v. Environments: more and more open and dynamic v. Biological domain is no exception Huge volumes of data o To be gathered, processed, exploited, visualised… Interaction networks o Large-scale o Interactions are incompletely known o Experimental data incomplete and heterogeneous Model integration o Building a whole o By assembling coupled parts o In order to explain a higher level of functioning Engineering Self-Modelling Systems: Application to Biology 3
![Towards Self-building Systems v. Complexity “autonomic computing” [IBM 03] v. Alleviate the designer’s task Towards Self-building Systems v. Complexity “autonomic computing” [IBM 03] v. Alleviate the designer’s task](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-4.jpg)
Towards Self-building Systems v. Complexity “autonomic computing” [IBM 03] v. Alleviate the designer’s task Initial expertise Some minimal feedback from time to time v. Let the system self-build v. Autonomous change of the organisation of the system v. Autonomous change of the behaviour of its components Ability to learn what is unknown (or incompletely known) Ability to interact in a different way Ability to appear/disappear Engineering Self-Modelling Systems: Application to Biology 4
![Self-organisation by Cooperation v. Adaptive Multi-Agent Systems theory [Camps 98, Capera 03] v. Social Self-organisation by Cooperation v. Adaptive Multi-Agent Systems theory [Camps 98, Capera 03] v. Social](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-5.jpg)
Self-organisation by Cooperation v. Adaptive Multi-Agent Systems theory [Camps 98, Capera 03] v. Social attitude of agents Perceive: Perceptions are understood without ambiguity Decide: Perceptions enable conclusion(s) Act: Actions are useful for the environment (and itself) v. A cooperative agent acts to Avoid Prevent Remove vsituations that it judges as being cooperative failures Engineering Self-Modelling Systems: Application to Biology 5
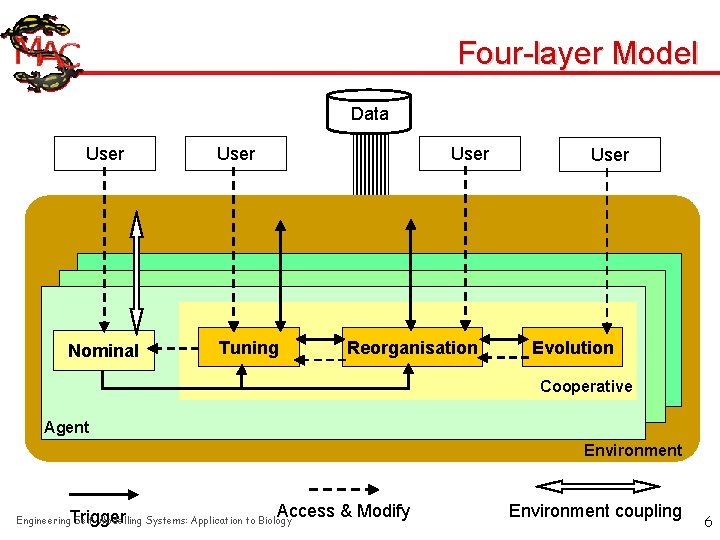
Four-layer Model Data User Nominal User Tuning Reorganisation User Evolution Cooperative Agent Environment Trigger Access & Modify Engineering Self-Modelling Systems: Application to Biology Environment coupling 6

Outline v. Making complex systems self-build Self-organisation by cooperation Four-layer model v. A domain of application: Biology micro. Mega specific case Agents and Biology v. Model applied to micro. Mega Architecture o Agents o Behaviours Preliminary results v. Conclusion Engineering Self-Modelling Systems: Application to Biology 7

Complexity and Biological Systems v. Theories are often missing v. Modelling and simulation (Gepasi [Mendes 93], Copasi…) v. Different approaches Mathematical models Petri nets Cellular automata Neural networks … v. Drawbacks Black boxes Models often static Far from a biological reality Engineering Self-Modelling Systems: Application to Biology 8

micro. Mega v. National project LISBP, INSA biologists o « Génie microbiologique » team o « Physiologie microbienne des eucaryotes » team LAAS, Disco team mathematicians LSP, UPS statisticians v. Multi-agent modelling of the genetic-metabolic interaction of a yeast (Saccharomyces Cerevisiae) v. From: Transcriptomic data: genes Macroscopic data: components v. In order to get free from experimental conditions v. Feasibility study Engineering Self-Modelling Systems: Application to Biology 9

Agents and Biology v. Agent and multi-agent technologies are rising [Lints 05, Merelli 06, Amigoni 07] v. Bioinformatics [Luck 05] or systems biology Protein folding/docking [Armano 05, Bortolussi 05] Pathways [Khan 03, Gonzalez 03, Querrec 03] Cell simulation [Webb 06, Lints 05, Boss 06, Jonker 08] Cell population simulation [Emonet 05, Troisi 05, D’Inverno 05, Guo 07] v. Discover new phenomena? Organisation is often fixed in MAS Laws considered as known Disruptions are not taken into account o Some exceptions [Querrec 03, Shafaei 08] Engineering Self-Modelling Systems: Application to Biology 10
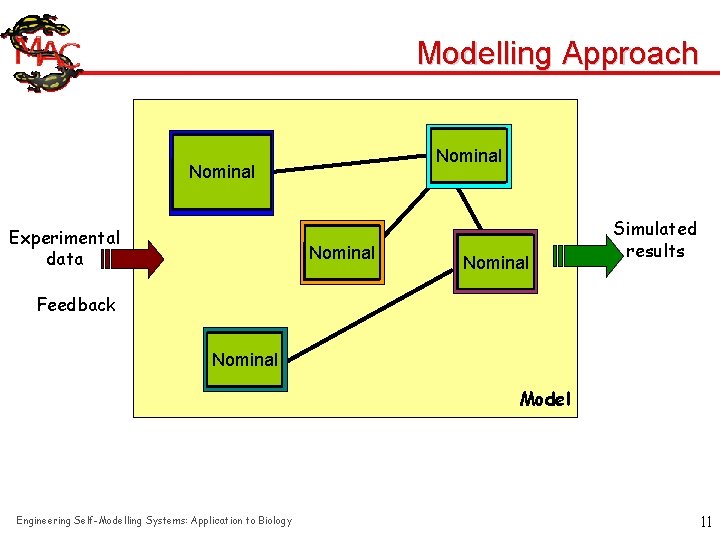
Modelling Approach Nominal Cooperative Nominal R E T Experimental data Cooperative Nominal T R E Nominal Cooperative Nominal T Simulated results R E Feedback Nominal Cooperative Nominal T R E Engineering Self-Modelling Systems: Application to Biology Model 11

Outline v. Making complex systems self-build Self-organisation by cooperation Four-layer model v. A domain of application: Biology micro. Mega specific case Agents and Biology v. Model applied to micro. Mega Architecture o Agents o Behaviours Preliminary results v. Conclusion Engineering Self-Modelling Systems: Application to Biology 12

Architecture of micro. Mega v. AMAS simulating chemical reactions v. Two kinds of cooperative agents Functional agents o Physical elements o Reactions o Interactions § Element consumption/production § Reactions regulation Viewer agents o Interactions with users o Data injection o Specific constraints Engineering Self-Modelling Systems: Application to Biology 13

Functional Agents v. Elements Represent common attributes for each element within the cell Quantity associated v. Reactions Genes o Confirm data about transcripts Transporters o Move an element quantity from one compartment to another o Passive / Active (ATP consumption) Catalysis o Transform a metabolite quantity into two o Catalysis may be regulated Synthesis o Assemble two metabolites o Synthesis may be regulated Engineering Self-Modelling Systems: Application to Biology 14
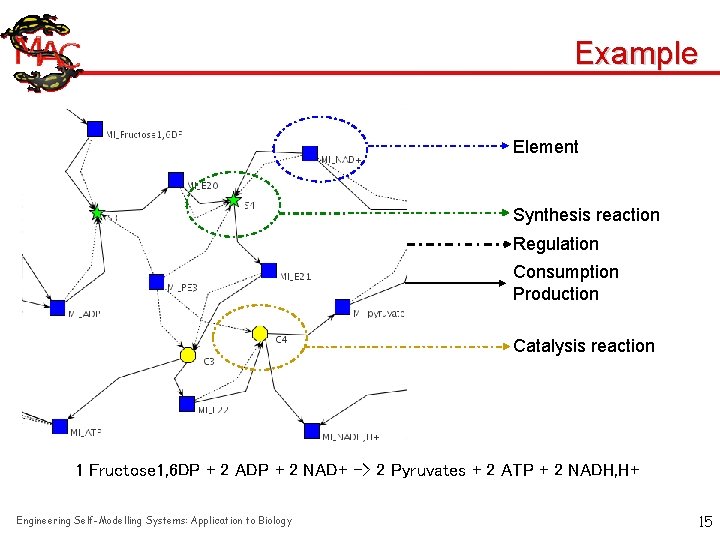
Example Element Synthesis reaction Regulation Consumption Production Catalysis reaction 1 Fructose 1, 6 DP + 2 ADP + 2 NAD+ -> 2 Pyruvates + 2 ATP + 2 NADH, H+ Engineering Self-Modelling Systems: Application to Biology 15

Viewer Agents v. Element. Viewer. Agent Gather quantities of a list of element agents v. Element. Setter. Agent Control activity of a list of element agents Database of experimental quantities v. But also… Evaluate biomass o Sum of the quantities of all element agents Identify compartments within the cell o If the system is able to reorganise o Manage user’s constraints Engineering Self-Modelling Systems: Application to Biology 16

Nominal Behaviour of Agents v. Element agents Manage related element quantity depending on feedback from reaction agents Linked to a compartment v. Reaction agents Consume/product element agents depending on: o Stoichiometry o Contextual reaction speed (possible regulations) v. Viewer agents Access data of functional agents Store these data Compute error related to experimental data Engineering Self-Modelling Systems: Application to Biology 17
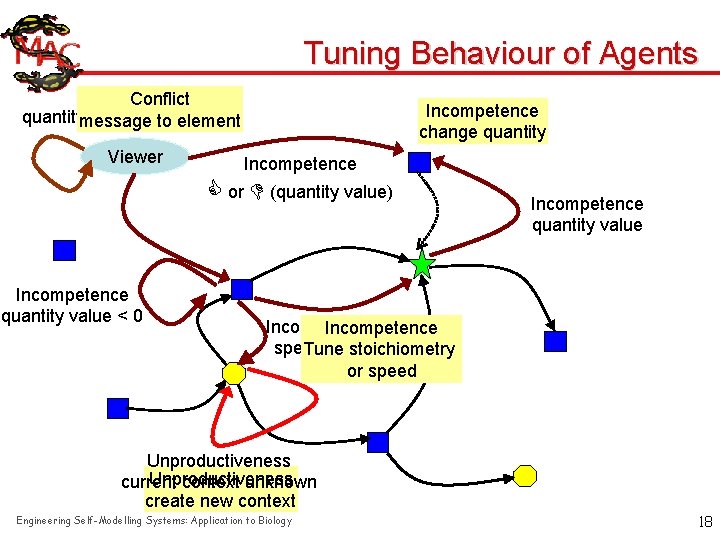
Tuning Behaviour of Agents Conflict quantitymessage error detected to element Viewer Incompetence change quantity Incompetence or (quantity value) Incompetence quantity value < 0 Incompetence quantity value Incompetence speed value Tune stoichiometry or speed Unproductiveness current context unknown create new context Engineering Self-Modelling Systems: Application to Biology 18
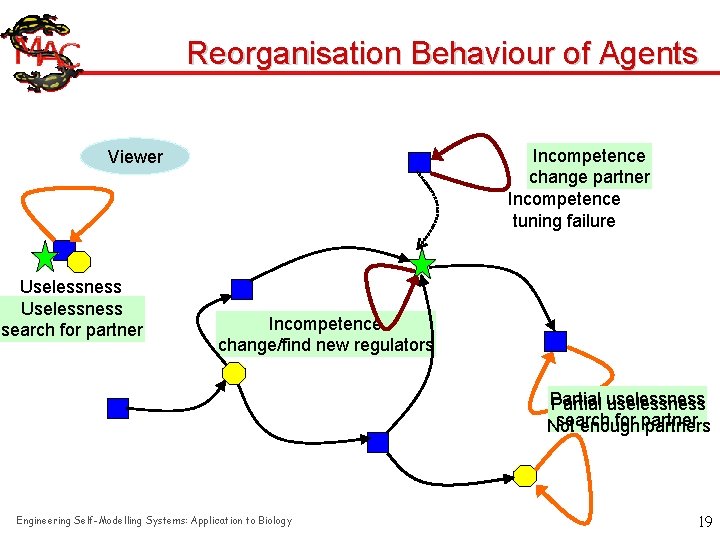
Reorganisation Behaviour of Agents Incompetence change partner Incompetence tuning failure Viewer Uselessness no partner Uselessness search for partner Incompetence tuningnew failure change/find regulators Partial uselessness search for partner Not enough partners Engineering Self-Modelling Systems: Application to Biology 19
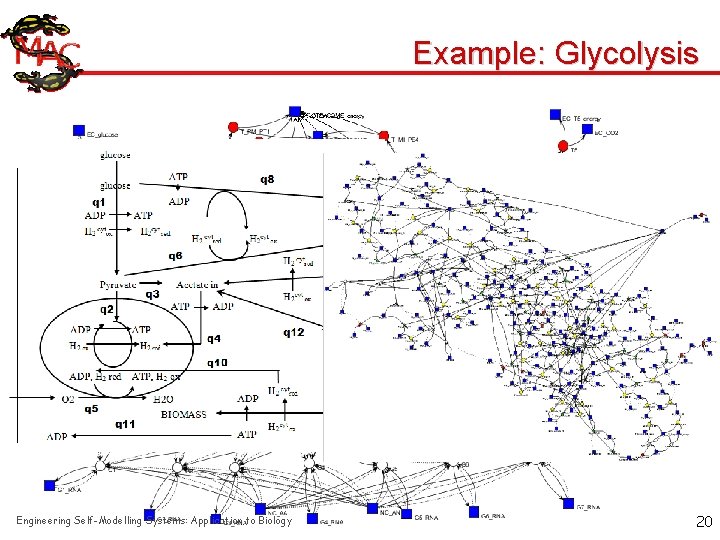
Example: Glycolysis Engineering Self-Modelling Systems: Application to Biology 20
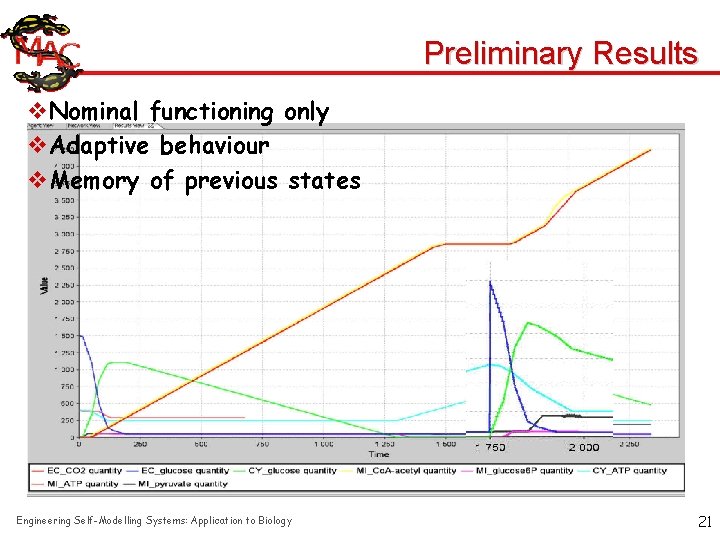
Preliminary Results v. Nominal functioning only v. Adaptive behaviour v. Memory of previous states Engineering Self-Modelling Systems: Application to Biology 21

Outline v. Making complex systems self-build Self-organisation by cooperation Four-layer model v. A domain of application: Biology micro. Mega specific case Agents and Biology v. Model applied to micro. Mega Architecture o Agents o Behaviours Preliminary results v. Conclusion Engineering Self-Modelling Systems: Application to Biology 22

Conclusion - Prospects v. Feasibility demonstration Self-building model Self-tuning model v. Model still incomplete v. Exhibits adaptation abilities v. Self-building = key for managing complexity v. Emergence = key for this self-building v. Finalise cooperative layers v. Overcome problems related to noise (forget) v. Validate models obtained on different experimental data Engineering Self-Modelling Systems: Application to Biology 23

Engineering Self-Modelling Systems: Application to Biology Thank you for your attention SMAC Team (Cooperative Multi-Agent Systems) Institut de Recherche en Informatique de Toulouse UPEtec www. irit. fr/SMAC - www. upetec. fr 20 janvier 2005
![References v. References related to SMAC team [Besse 05] C. Besse, Recherche de conformation References v. References related to SMAC team [Besse 05] C. Besse, Recherche de conformation](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-25.jpg)
References v. References related to SMAC team [Besse 05] C. Besse, Recherche de conformation de molécules et apprentissage du potentiel de Lennard. Jones par systèmes multi-agents adaptatifs, Research Master IARCL Report, Université Paul Sabatier, June 2005. [Camps 97] V. Camps, M. P. Gleizes, S. Trouilhet, Properties Analysis of a Learning Algorithm for Adaptive Systems, First International Conference on Computing Anticipatory Systems, Liège, Belgium, August 1997. [Camps 98] V. Camps, Vers une théorie de l'auto-organisation dans les systèmes multi-agents basée sur la coopération : application à la recherche d'information dans un système d'information répartie , Ph. D thesis, Université Paul Sabatier N° 2890, IRIT, Toulouse, January 1998. [Capera 05] D. Capera, Systèmes multi-agents adaptatifs pour la résolution de problèmes : Application à la conception de mécanismes, Ph. D thesis, Université Paul Sabatier, IRIT, Toulouse III, 23 June 2005. [Cornet 06] F. Cornet, Etude d'un problème d'allocation de fréquences par systèmes multi-agents adaptatifs, Research Master IARCL Report, Université Paul Sabatier, June 2006. [Dotto 99] F. Dotto, L. Trave-Massuyes, P. Glize, Acheminement du trafic d'un réseau téléphonique commuté par une approche multi agent adaptative, Congrès CCIA, Girona. [Georgé 04] J. P. Georgé, Résolution de problèmes par émergence, Etude d'un Environnement de Programmation Emergente, Ph. D thesis, Université Paul Sabatier, IRIT, Toulouse III, 6 July 2004. [Mano 06] J. P. Mano, Etude de l’émergence fonctionnelle au sein d’un réseau de neuro-agents coopératifs, Ph. D thesis, Université Paul Sabatier, IRIT, Toulouse III, 30 May 2006. 25
![References (2) v. References related to SMAC team (2) [Ottens 07] K. Ottens, Un References (2) v. References related to SMAC team (2) [Ottens 07] K. Ottens, Un](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-26.jpg)
References (2) v. References related to SMAC team (2) [Ottens 07] K. Ottens, Un système multi-agent adaptatif pour la construction d'ontologies à partir de textes, Ph. D thesis, Université Paul Sabatier, IRIT, Toulouse III, 2 October 2007. [Pesquet 99] B. Pesquet, M. P. Gleizes, P. Glize, Une équipe de robots footballeurs auto-organisée : les SMACkers, Intelligence artificielle située, cerveau, corps et environnement, A. Drogoul & J. A. Meyer coordonnateurs, Editions Hermès, 1999. [Picard 04] G. Picard, Cooperative Agent Model Instantiation to Collective Robotics, In: 5 th International Workshop on Engineering Societies in the Agents World (ESAW 2004), Toulouse, France, M. P. Gleizes, A. Omicini, F. Zambonelli (Eds), Springer Verlag, LNCS 3451, 209 -221. [Sontheimer 99] T. Sontheimer, Modèle adaptatif de prévision de crues par systèmes multi-agents autoorganisateurs, Institut Universitaire Professionnalisé Report, 1999, Diren. [TFGSO 04] Agent. Link. III TFG “Self-organisation in Multi-Agent Systems” report. [Topin 99] X. Topin, V. Fourcassie, M. P. Gleizes, G. Theraulaz, C. Régis, P. Glize, Theories and Experiments on Emergent Behaviour: From Natural to Artificial Systems and Back , In: European Conference on Cognitive Science, Siena, 1999. [Welcomme 08] J. B. Welcomme, MASCODE : un système multi-agent adaptatif pour concevoir des produits complexes. Application à la conception préliminaire avion , Ph. D thesis, Université de Toulouse, 31 March 2008. 26
![References (3) v. References external to SMAC team (1) [Amigoni 07] F. Amigoni, V. References (3) v. References external to SMAC team (1) [Amigoni 07] F. Amigoni, V.](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-27.jpg)
References (3) v. References external to SMAC team (1) [Amigoni 07] F. Amigoni, V. Schiaffonati, Multiagent-Based Simulation in Biology: A Critical Analysis , In: Model -Based Reasoning in Science, Technology, and Medicine, Springer, Studies in Computational Biology, 64, Lorenzo Magnani and Ping Li (Eds), 179 -191, 2007. [Armano 05] G. Armano, G. Mancosu, A. Orro, E. Vargiu, A Multi-agent System for Protein Secondary Structure Prediction, In: Transactions on Computational Systems Biology III, LNCS 3737, Springer, 14 -32, 2005. [Bortolussi 05] L. Bortolussi, A. Dovier, F. Fogolari, Multi-Agent Simulation of Protein Folding, In: First Workshop on Multi-Agent Systems for Medecine, Computational Biology, and Bioinformatics (MAS*BIOMED'05@AAMAS'05), 91 -106, 2005. [Bosse 06] T. Bosse, C. Jonker, J. Treur, Simulation and Analysis of Complex Biological Processes: an Organisation Modelling Perspective, In: 39 th Annual Simulation Symposium, 2006. [Camazine 01] S. Camazine, J. L. Deneubourg, N. Franks, J. Sneyd, G. Theraulaz G. , E. Bonabeau, Self. Organization in Biological Systems, Princeton University Press, 2001. [Conceicao 08] D. Conceição, M. Gatti, C. de Lucena, An Agent-based Framework for Stem Cell Behavior Modeling and Simulation, Research report 17/08, Department of Computer Sciences, Pontificia Universidade Catolico do Rio de Janeiro, April 2008. [D’Inverno 05] M. d’Inverno, R. Saunders, Agent-based Modelling of Stem Cell Organisation in a Niche, In: Engineering Self-Organising Systems: Methodologies and Applications , Springer, Brueckner S. , Di Marzo Serugendo G. , Karageorgos A. , Nagpal R. (Eds), LNCS 3464, Springer, 52 -68, 2005. [Emonet 05] T. Emonet, C. Macal, M. North, C. Wickersham, P. Cluzel, Agent. Cell: a Digital Single-cell Assay for Bacterial Chemotaxis, Bioinformatics Advance Access, In: Bioinformatics, 21, 2714 -2721, 2005. 27
![References (4) v. References external to SMAC team (2) [Querrec 03] G. Querrec, V. References (4) v. References external to SMAC team (2) [Querrec 03] G. Querrec, V.](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-28.jpg)
References (4) v. References external to SMAC team (2) [Querrec 03] G. Querrec, V. Rodin, J. F. Abgrall, S. Kerdelo, J. Tisseau, Uses of Multiagent Systems for Simulation of MAPK Pathway, In: Third IEEE Symposium on Bioinformatics and Bioengineering (BIBE'03), 421 -425, 2003. [Gonzalez 03] P. González, M. Cárdenas, D. Camacho, A. Franyuti, O. Rosas, J. Lagúnez-Otero, Cellulat: an Agent-based Intracellular Signalling Model, In: Biosystems, 68(2 -3), 171 -185, 2003. [Guo 07] D. Guo, E. Santos, A. Singhal, E. Santos, Q. Zhao, Adaptivity Modeling for Complex Adaptive Systems with Application to Biology, In: IEEE International Conference on Systems, Man and Cybernetics, 272 -277, 2007. [Jonker 08] C. Jonker, J. Snoep, J. Treur, H. Westerhoff, W. Wijngaards, BDI-modelling of Complex Intracellular Dynamics, In: Journal of Theoretical Biology, 251, 1 -23, 2008. [Khan 03] S. Khan, R. Makkena, W. Gillis, C. Schmidt, A Multi-agent System for the Quantitative Simulation of Biological Networks, In: Second International Joint Conference on Autonomous Agents & Multiagent Systems (AAMAS’ 03), Melbourne, ACM, 385 -392, 2003. [Lales 05] C. Lales, N. Parisey, J-P. Mazat, M. Beurton-Aimar, Simulation of Mitochondrial Metabolism using Multi-agents System, In: First Workshop on Multi-Agent Systems for Medecine, Computational Biology, and Bioinformatics (MAS*BIOMED'05 at AAMAS'05), 137 -145, 2005. [Lints 05] T. Lints, Multiagent Modelling of a Bacterial Cell, a Dna. A Titration Model Based Agent Model as an Example, In: Ninth Symposium on Programming Languages and Software Tools, Tartu, Estonia, Vene V. , Meriste M. (Eds. ), 82 -96, 2005. [Luck 05] M. Luck, E. Merelli, TFG on Agents in Bioinformatics, In: Knowledge Engineering Review, 20(2), 117125, 2005. [Mendes 93] P. Mendes, GEPASI: A Software Package for Modelling the Dynamics, Steady States and Control of Biochemical and other Systems, In: Computer Applications in the Biosciences, 9(5), 563 -571, 1993. 28
![References (5) v. References external to SMAC team (3) [Merelli 06] E. Merelli, G. References (5) v. References external to SMAC team (3) [Merelli 06] E. Merelli, G.](http://slidetodoc.com/presentation_image_h2/18d52dad763b3280b1ffbf9b3bac061f/image-29.jpg)
References (5) v. References external to SMAC team (3) [Merelli 06] E. Merelli, G. Armano, N. Cannata, F. Corradini, M. d'Inverno, A. Doms, P. Lord, A. Martin, L. Milanesi, S. Moller, M. Schroeder, M. Luck, Agents in Bioinformatics, Computational and Systems Biology, In: Briefings in Bioinformatics, 8(1), 45 -59, 2006. [Troisi 05] A. Troisi, V. Wong, M. Ratner, An Agent-based Approach for Modeling Molecular Self-organization , In: Proceedings of the National Academy of Sciences of the USA (PNAS), 102(2), 255 -260, 2005. [Santos 04] E. Santos, D. Guo, E. Santos Jr. , W. Onesty, A Multi-Agent System Environment for Modelling Cell and Tissue Biology, In: International Conference on Parallel and Distributed Processing Techniques and Applications, Las Vegas, USA, CSREA Press, Arabnia H. R. (Eds), 3 -9, 2004. [Shafaei 08] S. Shafaei, N. Aghaee, Biological Network Simulation Using Holonic Multiagent Systems , In: Tenth International Conference on Computer Modeling and Simulation (UKSIM'08), 1 -3 April, 617 -622, 2008. [Webb 06] K. Webb, T. White, Cell Modeling with Reusable Agent-based Formalisms, In: Applied Intelligence, 24(2), 169 -181, 2006. 29
- Slides: 29