ENGENHARIA METABLICA EM SACCHAROMYCES CEREVISIAE wesleygenerosousp br Saccharomyces

ENGENHARIA METABÓLICA EM SACCHAROMYCES CEREVISIAE wesleygeneroso@usp. br

Saccharomyces cerevisiae GRAS Glicólise em fluxo intenso Utilização industrial histórica Organismo modelo Alta eficiência em recombinação homóloga Preferência por reprodução assexuada Fácil manipulação genética
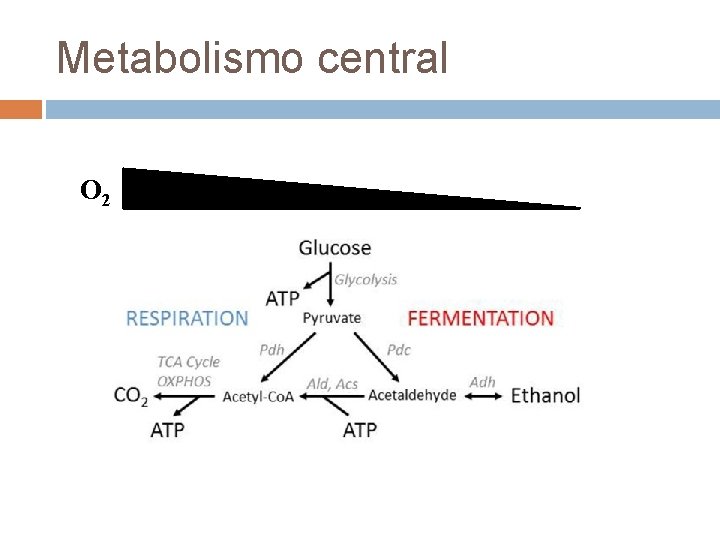
Metabolismo central O 2
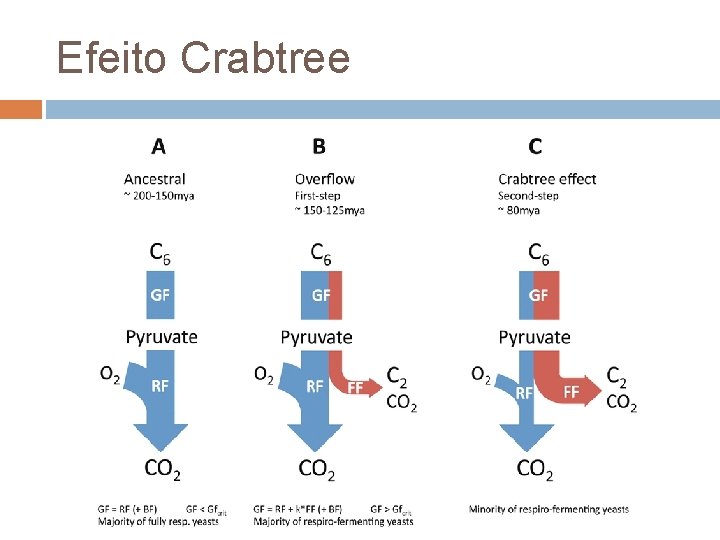
Efeito Crabtree

Características industriais Regulação fisiológica e populacional sob: � Estresse industrial � Estresse osmótico � Estresse ácido � Toxicidades (alcoóis e ácidos) � Contaminações Fácil separação do produto. Substratos para fermentação.
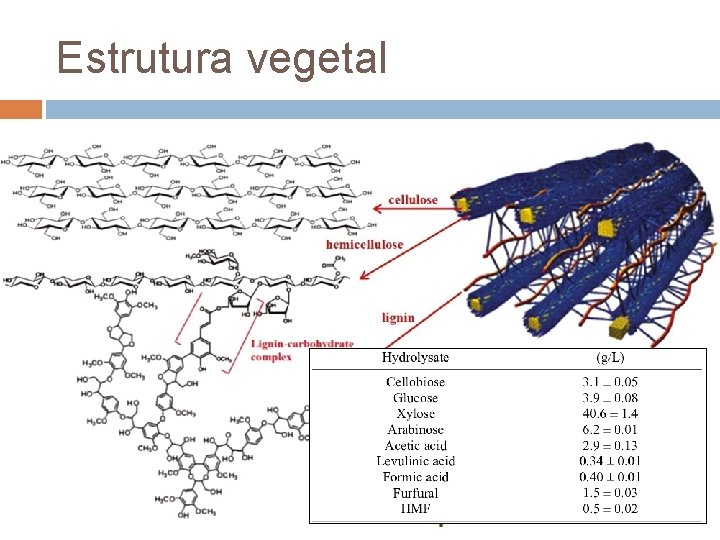
Estrutura vegetal
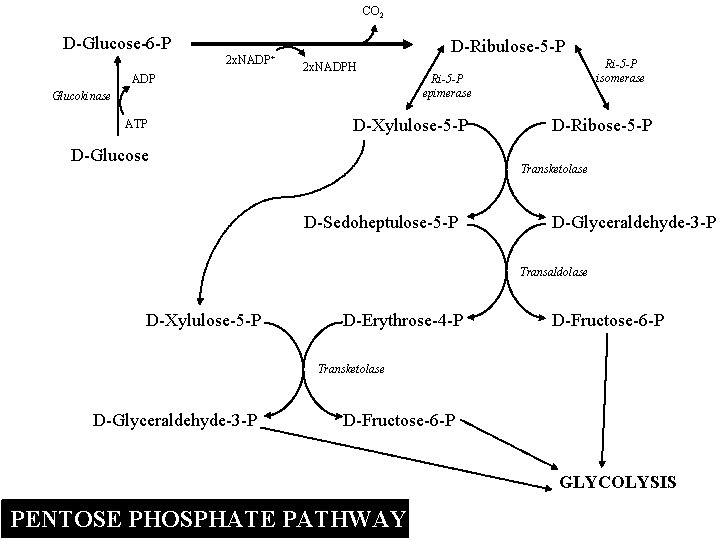
CO 2 D-Glucose-6 -P 2 x. NADP+ ADP D-Ribulose-5 -P 2 x. NADPH Glucokinase ATP Ri-5 -P isomerase Ri-5 -P epimerase D-Xylulose-5 -P D-Glucose D-Ribose-5 -P Transketolase D-Sedoheptulose-5 -P D-Glyceraldehyde-3 -P Transaldolase D-Xylulose-5 -P D-Erythrose-4 -P D-Fructose-6 -P Transketolase D-Glyceraldehyde-3 -P D-Fructose-6 -P GLYCOLYSIS PENTOSE PHOSPHATE PATHWAY
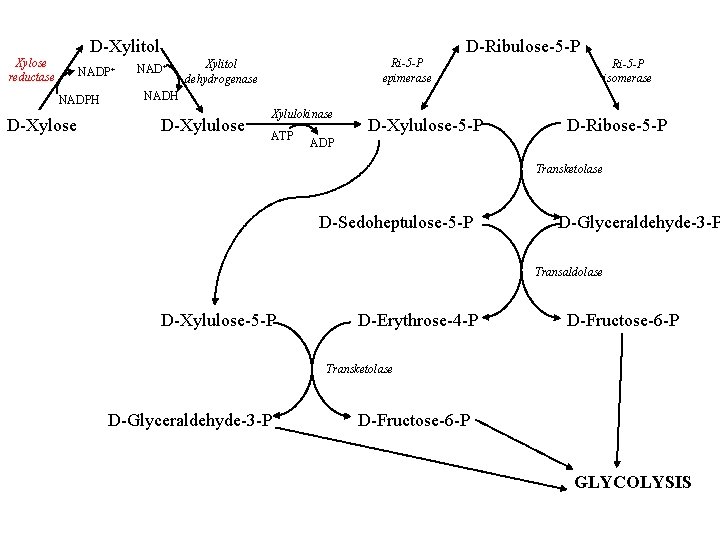
D-Xylitol Xylose reductase NADP+ NADPH D-Xylose D-Ribulose-5 -P NAD+ Ri-5 -P epimerase Xylitol dehydrogenase Ri-5 -P isomerase NADH D-Xylulose Xylulokinase ATP D-Xylulose-5 -P D-Ribose-5 -P ADP Transketolase D-Sedoheptulose-5 -P D-Glyceraldehyde-3 -P Transaldolase D-Xylulose-5 -P D-Erythrose-4 -P D-Fructose-6 -P Transketolase D-Glyceraldehyde-3 -P D-Fructose-6 -P GLYCOLYSIS


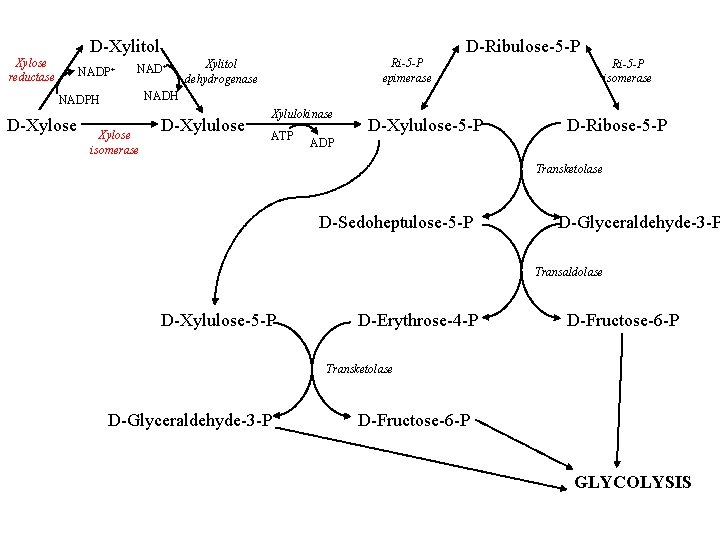
D-Xylitol Xylose reductase NADP+ NAD+ Xylitol dehydrogenase Ri-5 -P isomerase NADH NADPH D-Xylose D-Ribulose-5 -P Ri-5 -P epimerase Xylose isomerase D-Xylulose Xylulokinase ATP D-Xylulose-5 -P D-Ribose-5 -P ADP Transketolase D-Sedoheptulose-5 -P D-Glyceraldehyde-3 -P Transaldolase D-Xylulose-5 -P D-Erythrose-4 -P D-Fructose-6 -P Transketolase D-Glyceraldehyde-3 -P D-Fructose-6 -P GLYCOLYSIS

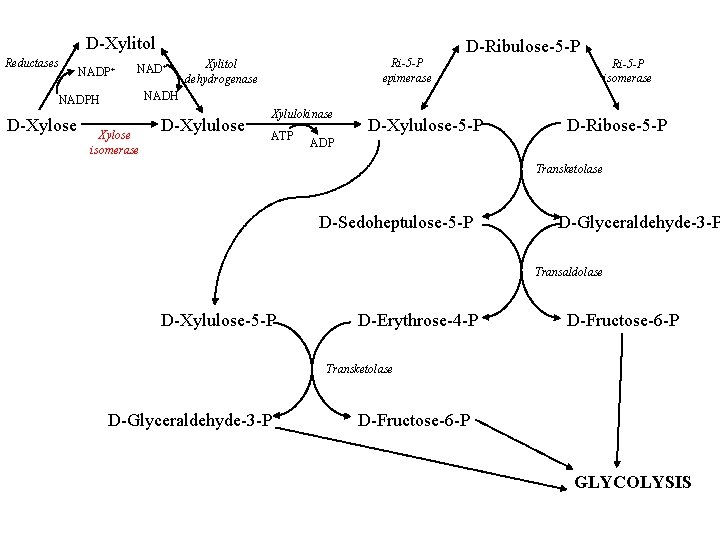
D-Xylitol Reductases NADP+ NAD+ Xylitol dehydrogenase Ri-5 -P isomerase NADH NADPH D-Xylose D-Ribulose-5 -P Ri-5 -P epimerase Xylose isomerase D-Xylulose Xylulokinase ATP D-Xylulose-5 -P D-Ribose-5 -P ADP Transketolase D-Sedoheptulose-5 -P D-Glyceraldehyde-3 -P Transaldolase D-Xylulose-5 -P D-Erythrose-4 -P D-Fructose-6 -P Transketolase D-Glyceraldehyde-3 -P D-Fructose-6 -P GLYCOLYSIS
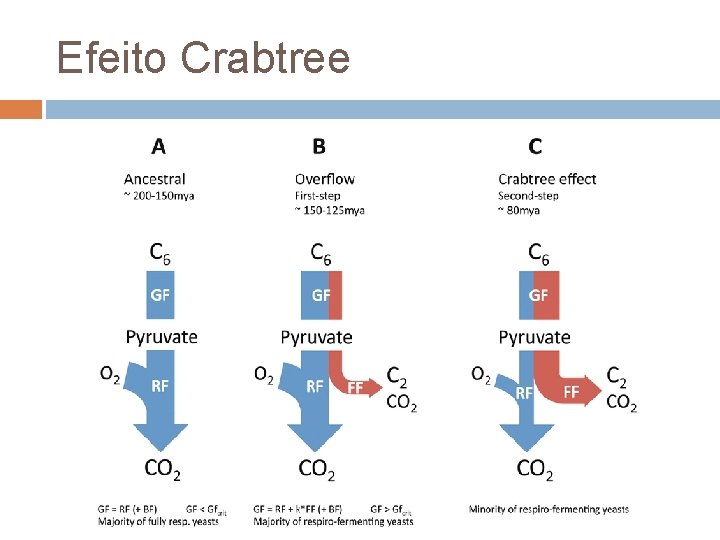
Efeito Crabtree
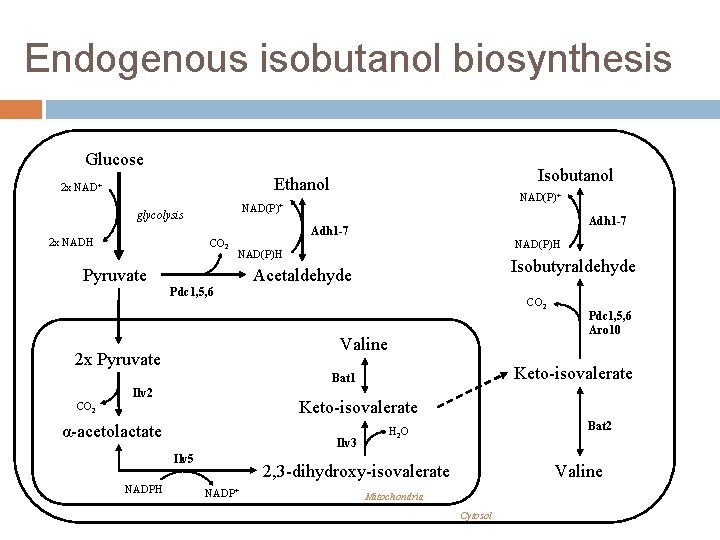
Endogenous isobutanol biosynthesis Glucose Isobutanol Ethanol 2 x NAD+ glycolysis 2 x NADH NAD(P)+ CO 2 Adh 1 -7 NAD(P)H Pyruvate Isobutyraldehyde Acetaldehyde Pdc 1, 5, 6 CO 2 Valine 2 x Pyruvate Keto-isovalerate Bat 1 Ilv 2 Keto-isovalerate CO 2 α-acetolactate Ilv 3 Ilv 5 NADPH Pdc 1, 5, 6 Aro 10 Bat 2 H 2 O 2, 3 -dihydroxy-isovalerate NADP+ Valine Mitochondria Cytosol
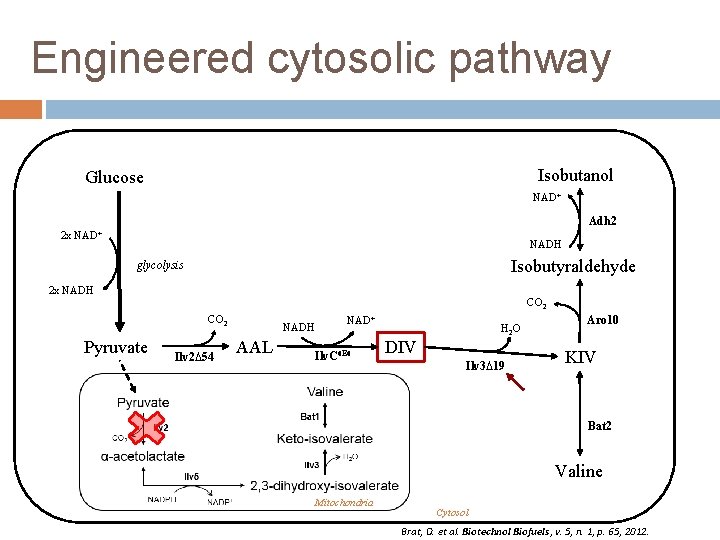
Engineered cytosolic pathway Isobutanol Glucose NAD+ Adh 2 2 x NAD+ NADH Isobutyraldehyde glycolysis 2 x NADH CO 2 Pyruvate Ilv 2∆54 NADH AAL NAD+ Ilv. C 6 E 6 H 2 O DIV Ilv 3∆19 Aro 10 KIV Bat 2 Valine Mitochondria Cytosol Brat, D. et al. Biotechnol Biofuels, v. 5, n. 1, p. 65, 2012.
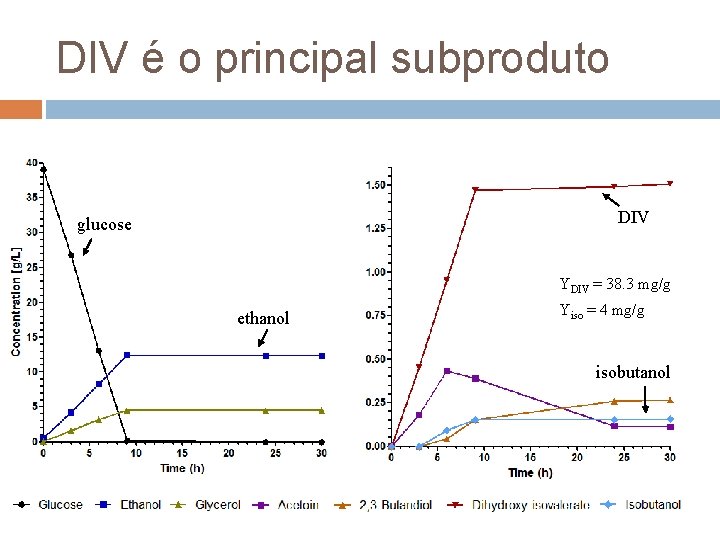
DIV é o principal subproduto DIV glucose YDIV = 38. 3 mg/g ethanol Yiso = 4 mg/g isobutanol
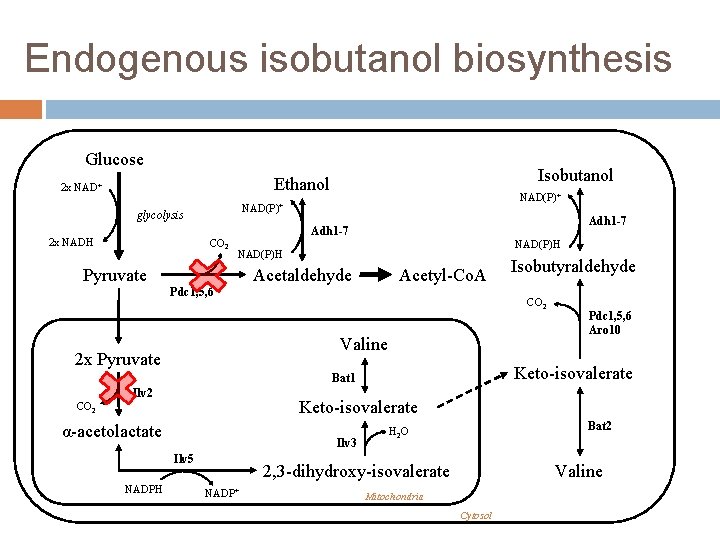
Endogenous isobutanol biosynthesis Glucose Isobutanol Ethanol 2 x NAD+ glycolysis 2 x NADH NAD(P)+ CO 2 Adh 1 -7 NAD(P)H Pyruvate Acetaldehyde Acetyl-Co. A Pdc 1, 5, 6 CO 2 Valine 2 x Pyruvate Keto-isovalerate CO 2 α-acetolactate Ilv 3 Ilv 5 NADPH Pdc 1, 5, 6 Aro 10 Keto-isovalerate Bat 1 Ilv 2 Isobutyraldehyde Bat 2 H 2 O 2, 3 -dihydroxy-isovalerate NADP+ Valine Mitochondria Cytosol

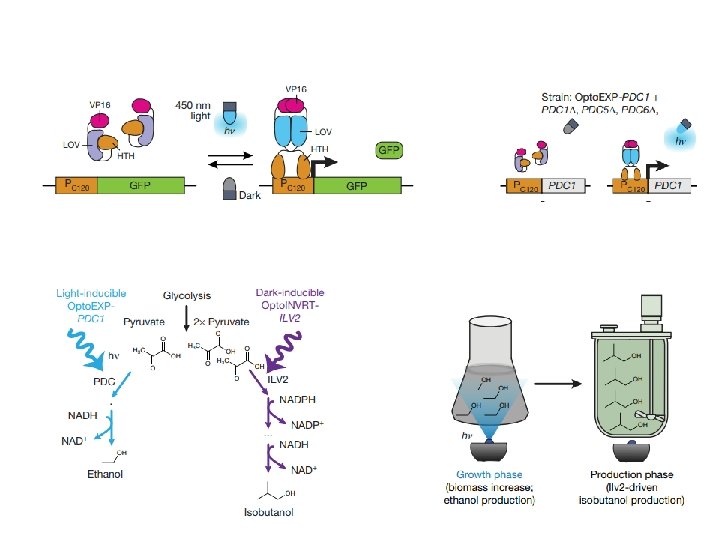
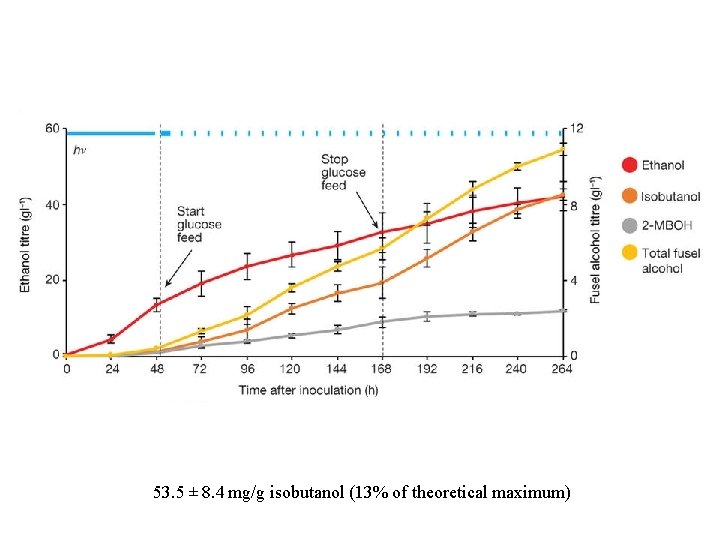
53. 5 ± 8. 4 mg/g isobutanol (13% of theoretical maximum)

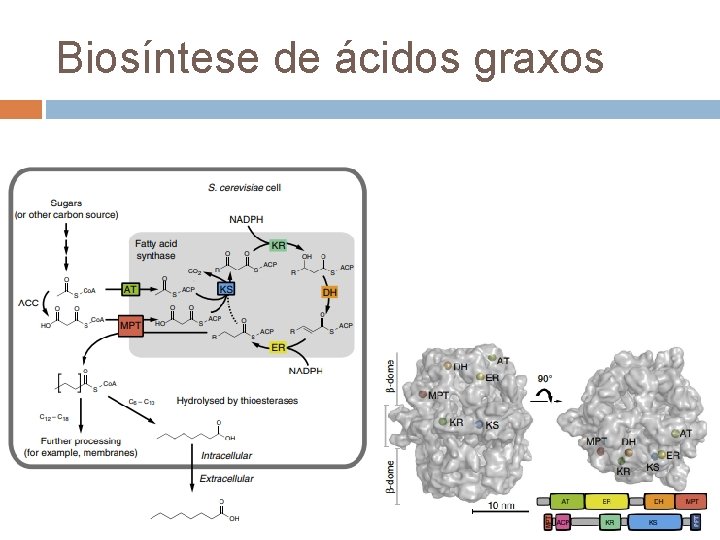
Biosíntese de ácidos graxos
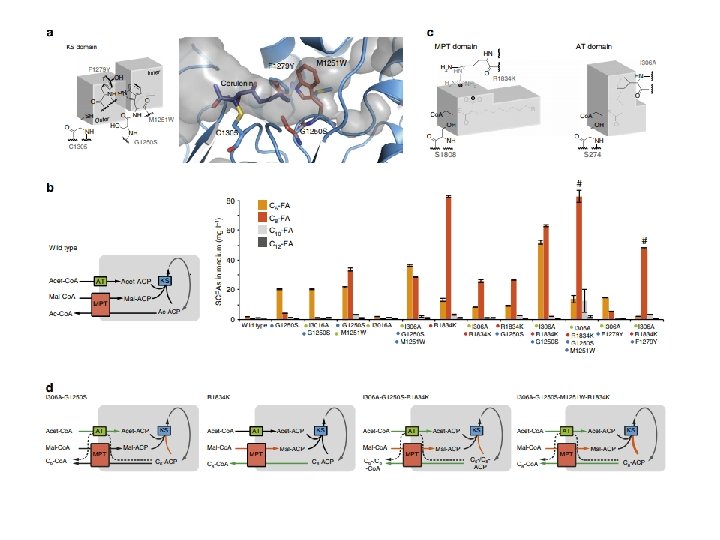

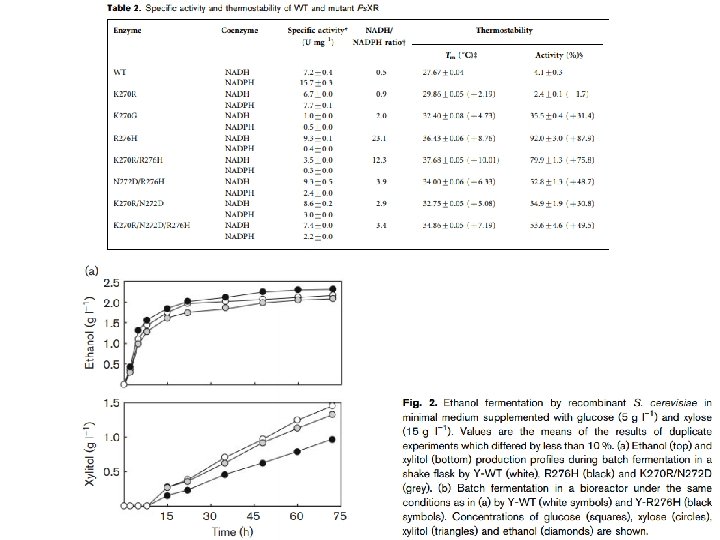
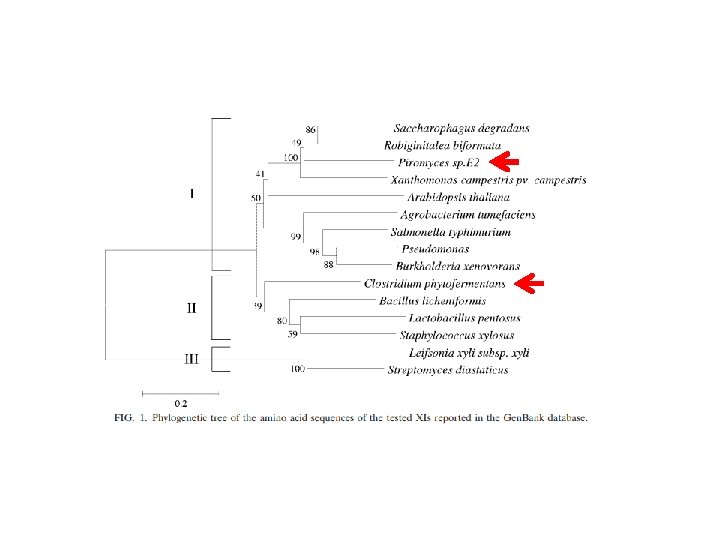
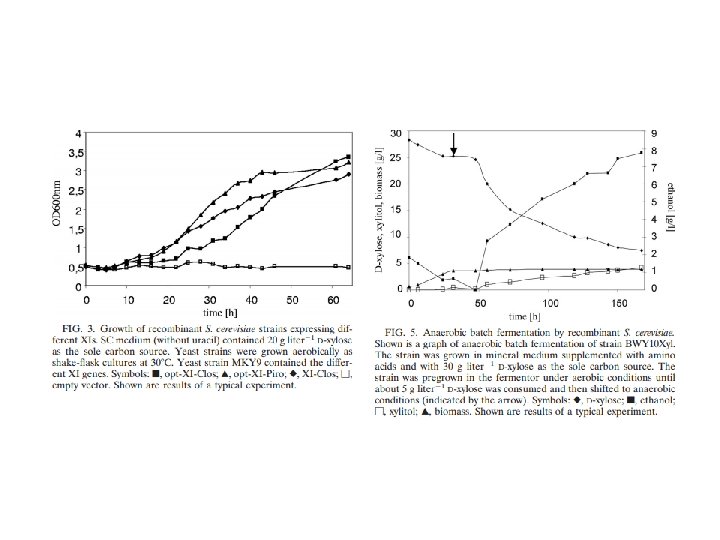
Reviews sobre eng. metabólica Moysés, Danuza Nogueira et al. “Xylose Fermentation by Saccharomyces Cerevisiae: Challenges and Prospects. ” International Journal of Molecular Sciences 17. 3 (2016): 207. Generoso et al. “Metabolic engineering of Saccharomyces cerevisiae for production of butanol isomers. ” Current Opinion in Biotechnology 33 (2015): 1 -7. Kavšček et al. “Yeast as a cell factory: current state and perspectives. ” Microbial Cell Factories 14 (2015): 94
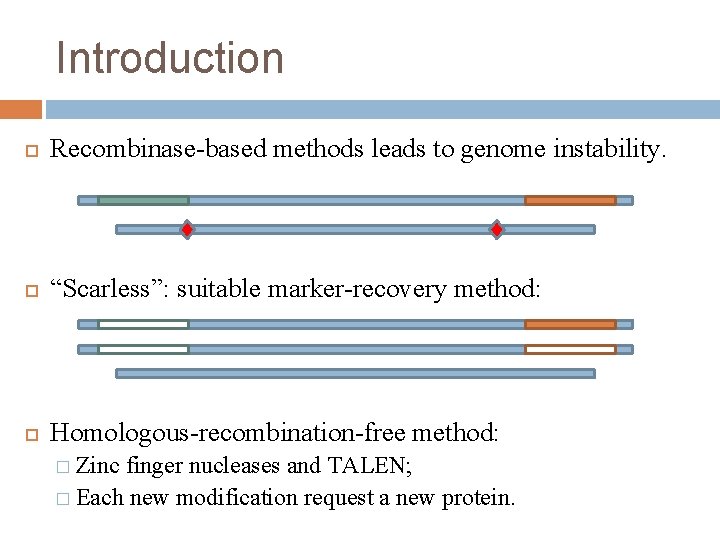
Introduction Recombinase-based methods leads to genome instability. “Scarless”: suitable marker-recovery method: Homologous-recombination-free method: � Zinc finger nucleases and TALEN; � Each new modification request a new protein.

Crispr/Cas 30 Prokaryotic immune system: � Confers resistance to foreign genetic elements. � CRISPR/Cas recognize and cut these exogenous genetic elements. � Analogous to RNAi in eukaryotic organisms. Gene truncations or insertions, and whole gene deletions. � Double-strand break mediated homologous recombination. � DNA restriction enzyme guided by a g. RNA.
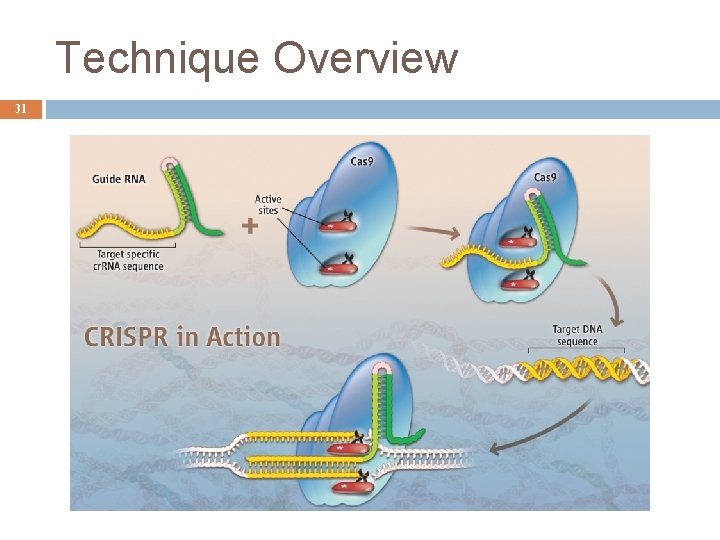
Technique Overview 31
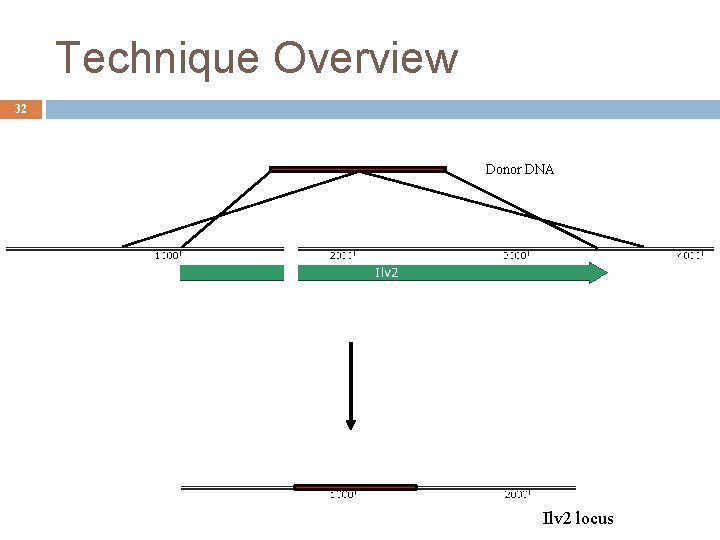
Technique Overview 32 Donor DNA Ilv 2 locus
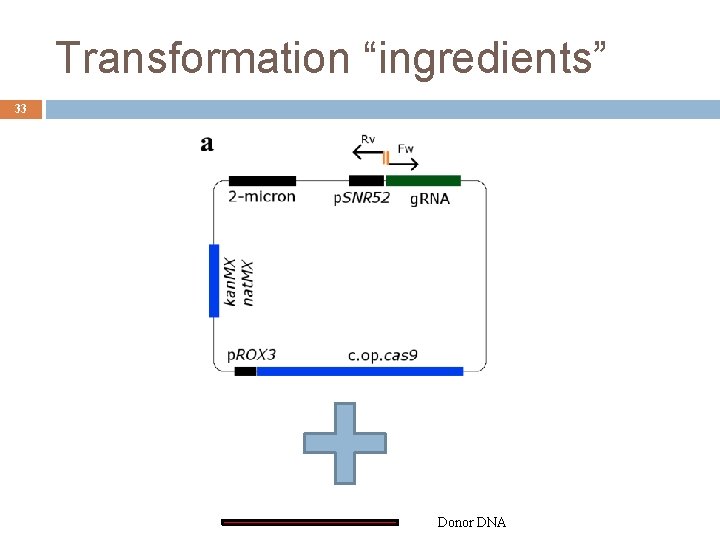
Transformation “ingredients” 33 Donor DNA
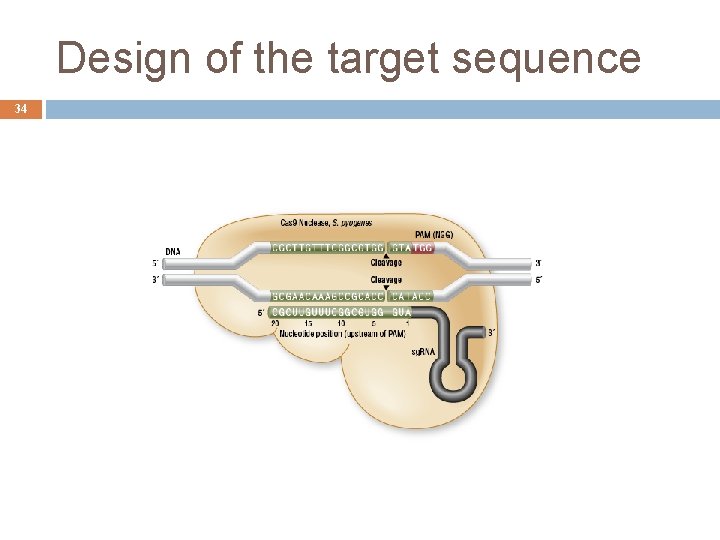
Design of the target sequence 34
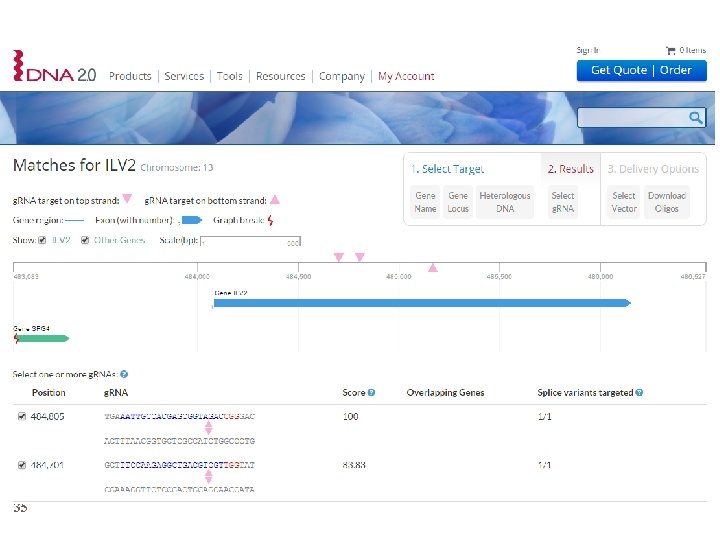
35

Considerations 36 No incorporation in the genome is needed. Deletion of two alleles or multiple genes at once. Fast and versatile for genome editing. “Complex plasmid building” for the g. RNA. You must have the system (plasmid).
- Slides: 36