Elongation and prem RNA processing MBV 4230 Introduction

Elongation and pre-m. RNA processing

MBV 4230 Introduction n Regulation of elongation most transcription units are probably regulated during elongation because the elongation machinery must coordinate with so many other nuclear processes while navigating a nucleoprotein template. ¨ Regulation can be general, applying to many genes, or selective. ¨ n Themes RNA polymerase II ¨ Elongation factors - Specific proteins affecting elongation ¨ Chromatin and elongation ¨ Pre-m. RNA processing ¨ n Capping, Splicing and Termination/3´-end formation

The elongating RNAPII
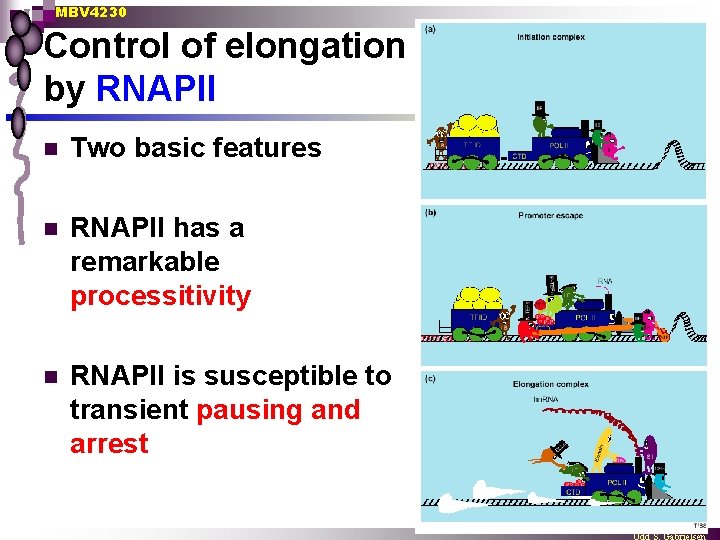
MBV 4230 Control of elongation by RNAPII n Two basic features n RNAPII has a remarkable processitivity n RNAPII is susceptible to transient pausing and arrest

MBV 4230 The Elongation phase - RNAPII n n Processivity explained by 3 D of RNAPII RNAP closes upon”promoter clearance” and transition to TEC (trx elongation complex) Contacts to PIC disrupted and new contacts with elongation factors formed ¨ CTD phosphorylated ¨ Conformational change to a ternary complex of high stability ¨ Closed chanel around DNA-RNA hybrid in the active site ¨ Encouter obstacles that lead to to pausing or arrest

MBV 4230 Early evidence for elongation factors n Early evidence for general elongation factors ¨ Elongation rate of RNAPII in vitro << in vivo n In vitro: 100 -300 nt per min, frequent pauses, some times full arrest n In vivo: 1200 -2000 nt per min, probably because elongation-factors suppress pausing ¨ The DRB-inhibitor: nucleotide-analogue causing strong inhibition of hn. RNA synthesis, acts by enhanced arrest of RNAPII, but has no effect on purified RNAPII, targets probably an elongation factor n DRB = 5, 6 -dichloro-1 -ß-D-ribofuranosylbenzimidazole
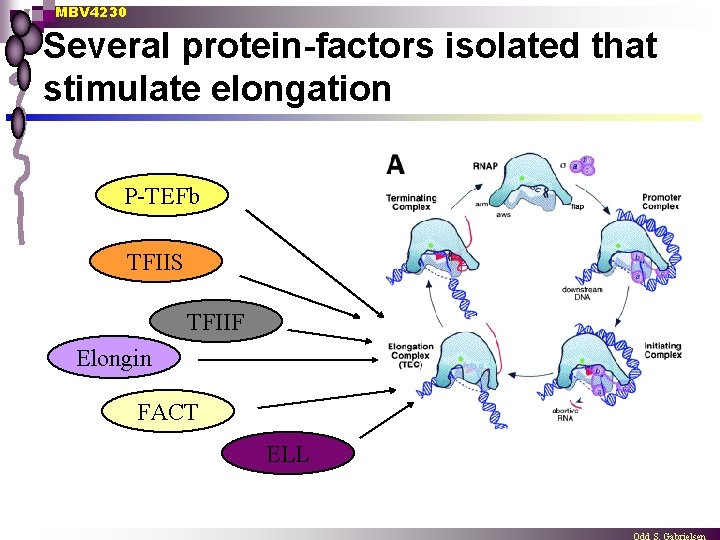
MBV 4230 Several protein-factors isolated that stimulate elongation P-TEFb TFIIS TFIIF Elongin FACT ELL
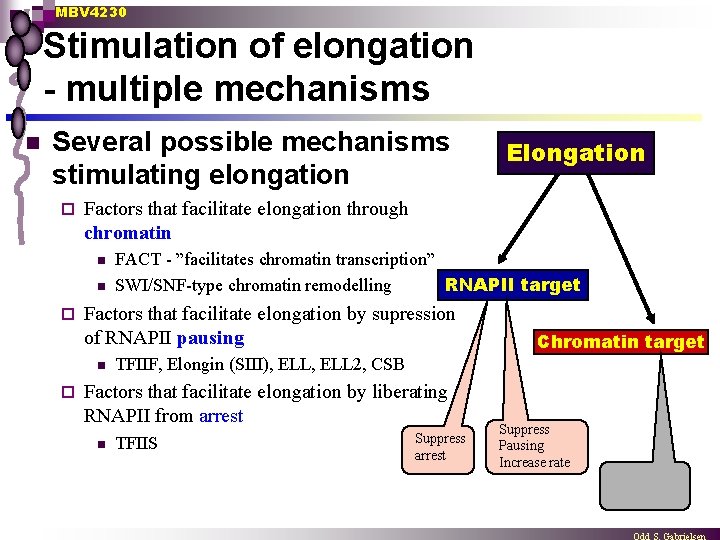
MBV 4230 Stimulation of elongation - multiple mechanisms n Several possible mechanisms stimulating elongation ¨ Factors that facilitate elongation through chromatin n n ¨ FACT - ”facilitates chromatin transcription” RNAPII target SWI/SNF-type chromatin remodelling Factors that facilitate elongation by supression of RNAPII pausing n ¨ Elongation TFIIF, Elongin (SIII), ELL 2, CSB Factors that facilitate elongation by liberating RNAPII from arrest n Chromatin target TFIIS Suppress arrest Suppress Pausing Increase rate

Elongation factors that target RNAPII Phosphorylation of CTD - early stages of elongation

MBV 4230 Elongation factors that target RNAPII: P-TEFb phosphorylates CTD n n The P-TEFb (positive transcription elongation factor) complex, which contains the cyclindependent kinase CDK 9 and cyclin T, couples RNA processing to transcription by phosphorylating Ser 2 of the RNAP II CTD Identified biochemically ¨ n n Based on its ability to protect RNAPII aginst arrest in a Drosophila tr. system structure: Heterodimer = 124 k. Da + 43 k. Da activity: a CTD-kinase Cdk 9 (også kalt PITALRE) + cyclin T 1, T 2 or K ¨ Kinase-inactive form without effect on elongation ¨ n Distinctive feature: inhibited by the nucleotide analogue DRB
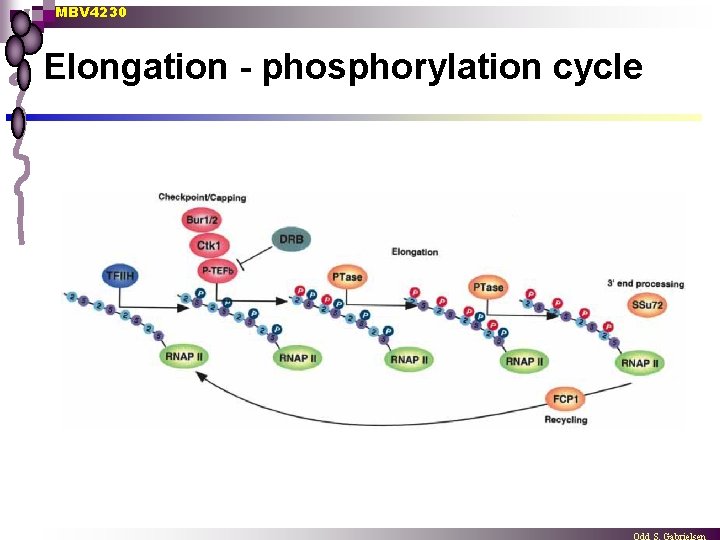
MBV 4230 Elongation - phosphorylation cycle

MBV 4230 P-TEFb action - release from DSIF/NELF induced pause n Mechanism of action Ser 2 phosphorylation of CTD with P-TEFb blocks binding of the elongation -inhibitors NELF and DSIF (DRB-sensitivity inducing factor) ¨ NELF = Negative ELongation Factor, a multiprotein complex (5 polypeptides 46 -66 k. Da) ¨ DSIF = DRB Sensitivity Inducing Factor, a heterodimer (14 + 160 k. Da) = Spt 4 + Spt 5 in yeast ¨ Stop and wait for capping: DSIF interacts with hypophosphorylated CTD. NELF recognizes the RNAP II–DSIF complex and halts elongation. This pause allows the recruitment of the capping enzyme by the CTD and DSIF (Spt 5 subunit), which adds a 5 -cap to the nascent transcript. ¨
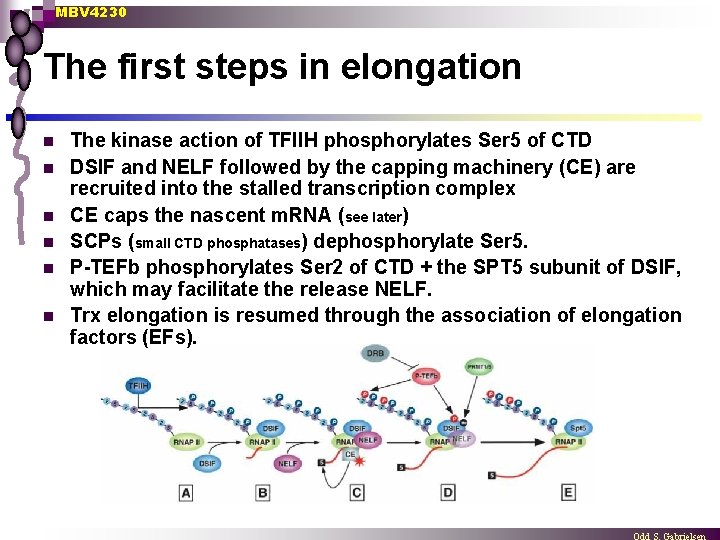
MBV 4230 The first steps in elongation n n n The kinase action of TFIIH phosphorylates Ser 5 of CTD DSIF and NELF followed by the capping machinery (CE) are recruited into the stalled transcription complex CE caps the nascent m. RNA (see later) SCPs (small CTD phosphatases) dephosphorylate Ser 5. P-TEFb phosphorylates Ser 2 of CTD + the SPT 5 subunit of DSIF, which may facilitate the release NELF. Trx elongation is resumed through the association of elongation factors (EFs).
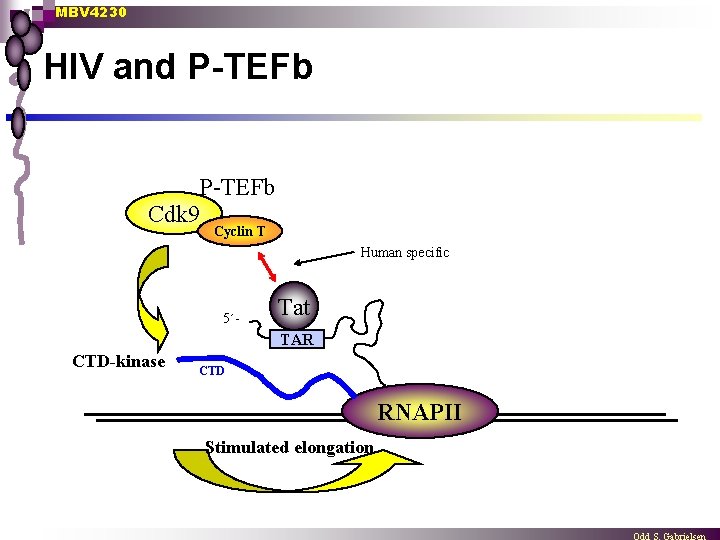
MBV 4230 HIV and P-TEFb Cdk 9 Cyclin T Human specific 5´- Tat TAR CTD-kinase CTD RNAPII Stimulated elongation

MBV 4230 HIV and P-TEFb n HIV-1 produces its own”elongation factor” Tat is a sequence-specific RNA-binding protein encoded by HIV ¨ Tat binds to a sequence-element TAR (transactivation response element) in the 5´-end of HIV-transcripts ¨ Tat+TAR promote effective elongation of HIV transcripts ¨ n P-TEFb + CTD is required for Tat-function Ternary complex formed with human cyclin T 1+ Tat + TAR (human, not murine T 1) ¨ Murine cells become HIV-infectable after transfection with human cyclin T 1 ¨ Mechanism: Tat facilitates HIV expression by recruiting P-TEFb to TAR, which improves specifically elongation of HIV-transcripts ¨

Elongation factors Helping arrested RNAPII

MBV 4230 Pausing and arrest n n n RNAPII encounters obstacles during elongation leading to pausing or arrest This stage of trx is subject to control and several genes may be regulated also on the level of elongation Pausing and arrest ¨ n n result from aberrant backward movement of RNAPII, leading to displacement of the 3´-OH end of the growing RNA from the catalytic site Pausing = reversible Arrest = not reversible

MBV 4230 Elongation factors that suppress RNAPII arrest : TFIIS n n structure: monomer = 38 k. Da TFIIS binds arrested RNAPII and TFIIS strongly enhances a weak intrinsic nuclease activity of RNAPII TFIIS induces the polymerase to cleave its nascent transcript, repositioning the new RNA 3´-end within the polymerase catalytic center
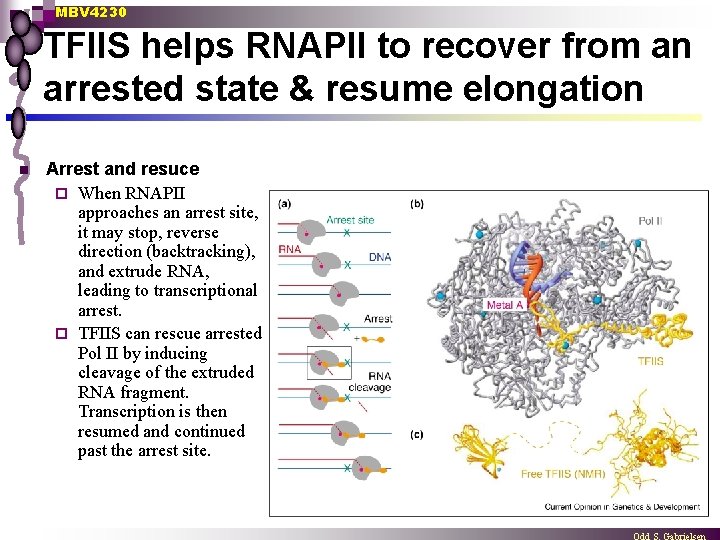
MBV 4230 TFIIS helps RNAPII to recover from an arrested state & resume elongation n Arrest and resuce ¨ When RNAPII approaches an arrest site, it may stop, reverse direction (backtracking), and extrude RNA, leading to transcriptional arrest. ¨ TFIIS can rescue arrested Pol II by inducing cleavage of the extruded RNA fragment. Transcription is then resumed and continued past the arrest site.
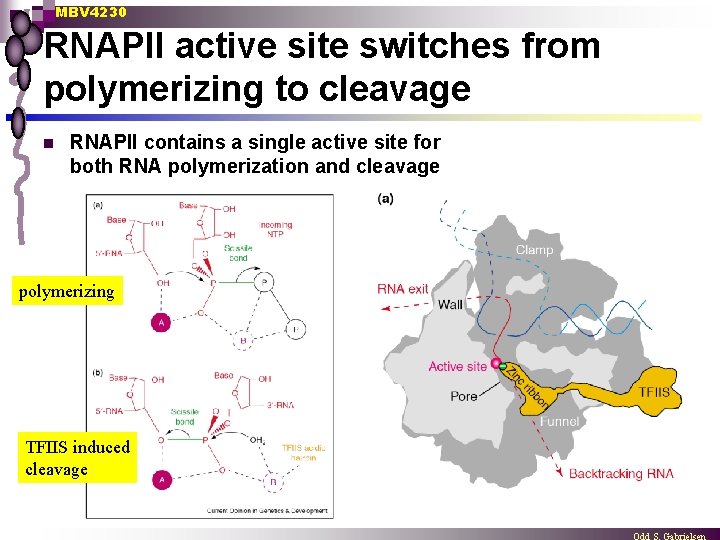
MBV 4230 RNAPII active site switches from polymerizing to cleavage n RNAPII contains a single active site for both RNA polymerization and cleavage polymerizing TFIIS induced cleavage

Elongation factors Helping paused RNAPII

MBV 4230 Pausing and arrest n Pausing = rate-limiting step during elongation RNAPII susceptible to pausing at each step ¨ RNAPII cycles between active and inactive (paused) conformations ¨
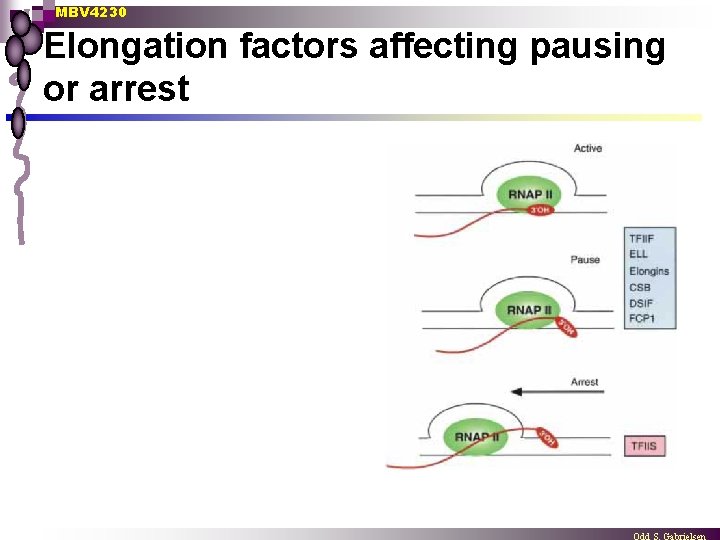
MBV 4230 Elongation factors affecting pausing or arrest

MBV 4230 Elongation factors that suppress pausing: TFIIF, Elongin (SIII), and ELL n TFIIF Protects the elongation complex against pausing ¨ Acts probably by a direct but transient interaction with the elongating RNAPII ¨ ¨ phosphorylation of RAP 74 stabilizes binding to RNAPII and stimulates elongation

MBV 4230 Elongation factors that suppress pausing: TFIIF, Elongin (SIII), and ELL n Elongin Heterotrimer of subunits A, B and C where A is active, B and C regulatory ¨ Elongins activity can probably be regulated by the von-Hippel-Lindau (VHL) tumor supressor protein which binds Elongin BC and blocks their binding to Elongin A. ¨ Genetic disease VHL dispose for several cancers, where mutated VHL binds Elongin BC less avidly ¨ n ELL ¨ 80 k. Da elongation factor found fused with MLL (mixed lineage leukemia) in certain leukemias with translocation between chromosome 11 and 19 (ELevennineteen Leukemia)

MBV 4230 Mechanims of action n Pausing = rate-limiting step during elongation RNAPII susceptible to pausing at each step ¨ RNAPII cycles between active and inactive (paused) conformations ¨ n Elongation factors that suppress pausing, probably act by decreasing the fraction of time RNAPII spends in an inactive paused conformation ¨ For many factors supressing pausing and increasing rate of trx, our understanding of mechanism is incomplete

Elongation factors helping RNAPII through chromatin

MBV 4230 Elongation through chromatin n Chromatin is not only an obstacle to TFIID binding and PIC assembly, but also for the elongating RNAPII
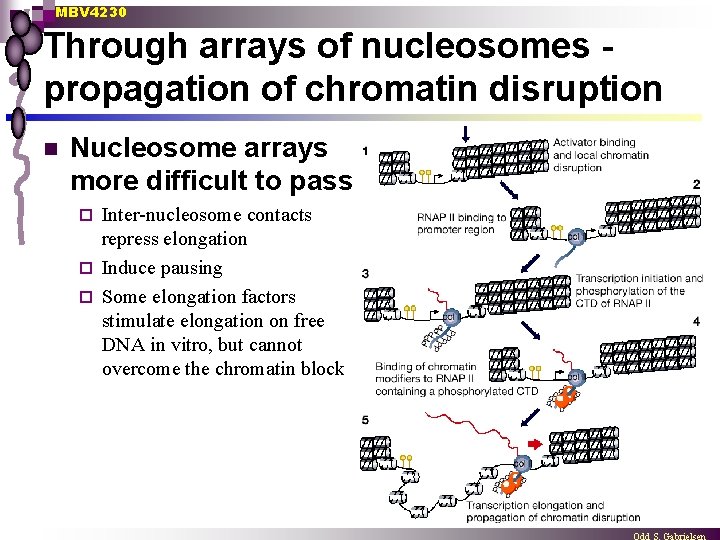
MBV 4230 Through arrays of nucleosomes propagation of chromatin disruption n Nucleosome arrays more difficult to pass Inter-nucleosome contacts repress elongation ¨ Induce pausing ¨ Some elongation factors stimulate elongation on free DNA in vitro, but cannot overcome the chromatin block ¨

MBV 4230 Through arrays of nucleosomes propagation of chromatin disruption n In vivo cellular factors helps to disrupt the chromatin block to elongation

MBV 4230 Elongation factors acting on chromatin n Factors that facilitate elongation through chromatin ¨ SWI/SNF-type chromatin remodellering through ATPdependent mechanisms n Swi-Snf and Chd 1 remodel nucleosomes Proteins that acetylate (e. g. Gcn 5 and Elp 3) or methylate histones ¨ FACT - ”facilitates chromatin transcription”- can bind to and destabilize nucleosomes ¨ n n n ¨ n a heterodimer where SPT 16 encodes the large subunit HMG 1 -like factor SSRP 1 Proposed that FACT transiently binds and removes H 2 A+H 2 B Spt 4+Spt 5 (DSIF) and SPT 6 proteins Reassembly of chromatin after passage of RNAPII important To suppress trx initiation from cryptic initiation site (noise) ¨ FACT and SPT 6 probably acts by enabling chromatin structure to be disrupted and then reestablished during trx ¨
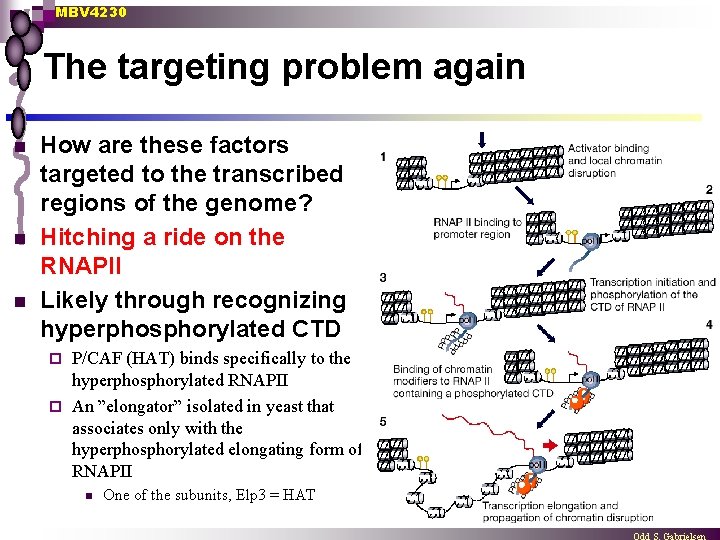
MBV 4230 The targeting problem again n How are these factors targeted to the transcribed regions of the genome? Hitching a ride on the RNAPII Likely through recognizing hyperphosphorylated CTD P/CAF (HAT) binds specifically to the hyperphosphorylated RNAPII ¨ An ”elongator” isolated in yeast that associates only with the hyperphosphorylated elongating form of RNAPII ¨ n One of the subunits, Elp 3 = HAT
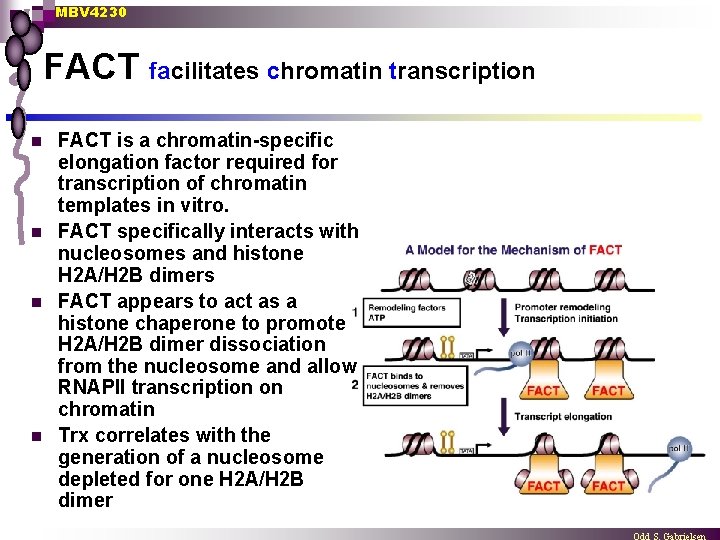
MBV 4230 FACT facilitates chromatin transcription n n FACT is a chromatin-specific elongation factor required for transcription of chromatin templates in vitro. FACT specifically interacts with nucleosomes and histone H 2 A/H 2 B dimers FACT appears to act as a histone chaperone to promote H 2 A/H 2 B dimer dissociation from the nucleosome and allow RNAPII transcription on chromatin Trx correlates with the generation of a nucleosome depleted for one H 2 A/H 2 B dimer
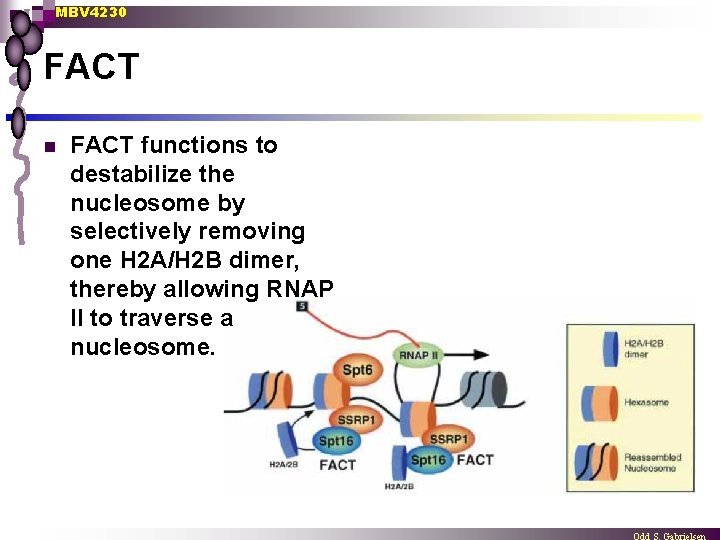
MBV 4230 FACT n FACT functions to destabilize the nucleosome by selectively removing one H 2 A/H 2 B dimer, thereby allowing RNAP II to traverse a nucleosome.
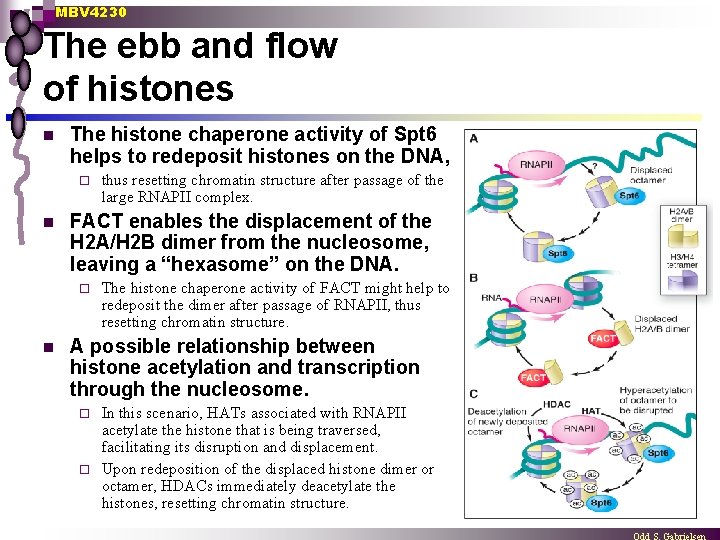
MBV 4230 The ebb and flow of histones n The histone chaperone activity of Spt 6 helps to redeposit histones on the DNA, ¨ n FACT enables the displacement of the H 2 A/H 2 B dimer from the nucleosome, leaving a “hexasome” on the DNA. ¨ n thus resetting chromatin structure after passage of the large RNAPII complex. The histone chaperone activity of FACT might help to redeposit the dimer after passage of RNAPII, thus resetting chromatin structure. A possible relationship between histone acetylation and transcription through the nucleosome. In this scenario, HATs associated with RNAPII acetylate the histone that is being traversed, facilitating its disruption and displacement. ¨ Upon redeposition of the displaced histone dimer or octamer, HDACs immediately deacetylate the histones, resetting chromatin structure. ¨
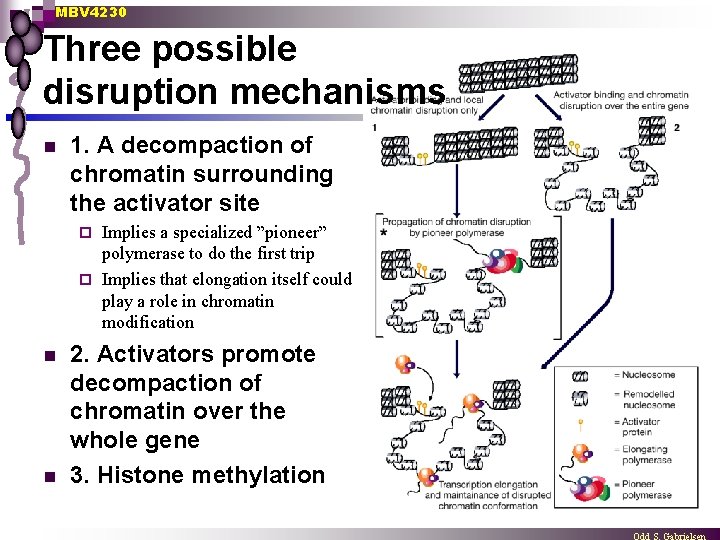
MBV 4230 Three possible disruption mechanisms n 1. A decompaction of chromatin surrounding the activator site Implies a specialized ”pioneer” polymerase to do the first trip ¨ Implies that elongation itself could play a role in chromatin modification ¨ n n 2. Activators promote decompaction of chromatin over the whole gene 3. Histone methylation
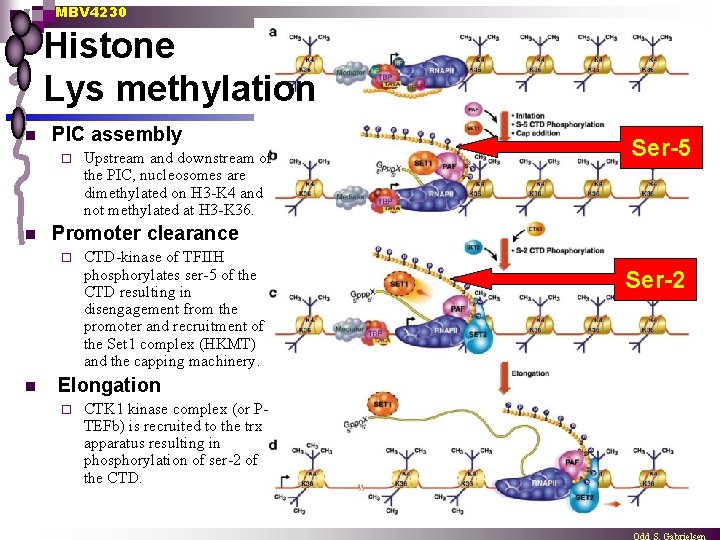
MBV 4230 Histone Lys methylation n PIC assembly ¨ n Ser-5 Promoter clearance ¨ n Upstream and downstream of the PIC, nucleosomes are dimethylated on H 3 -K 4 and not methylated at H 3 -K 36. CTD-kinase of TFIIH phosphorylates ser-5 of the CTD resulting in disengagement from the promoter and recruitment of the Set 1 complex (HKMT) and the capping machinery. Elongation ¨ CTK 1 kinase complex (or PTEFb) is recruited to the trx apparatus resulting in phosphorylation of ser-2 of the CTD. Ser-2
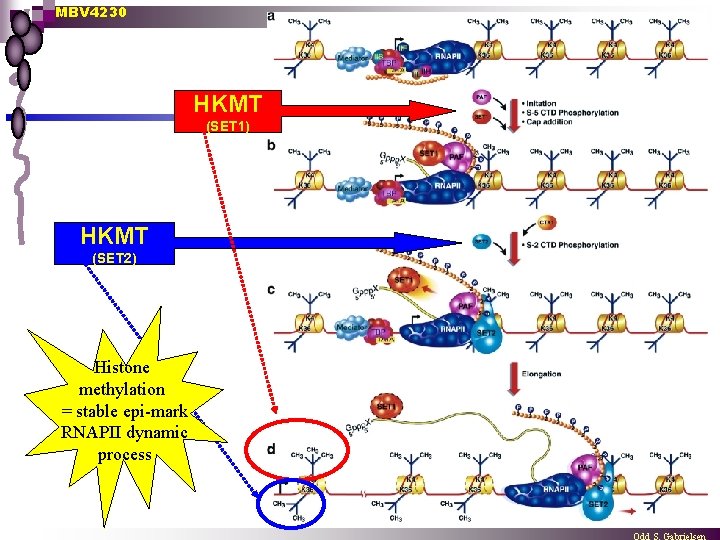
MBV 4230 HKMT (SET 1) HKMT (SET 2) Histone methylation = stable epi-mark RNAPII dynamic process
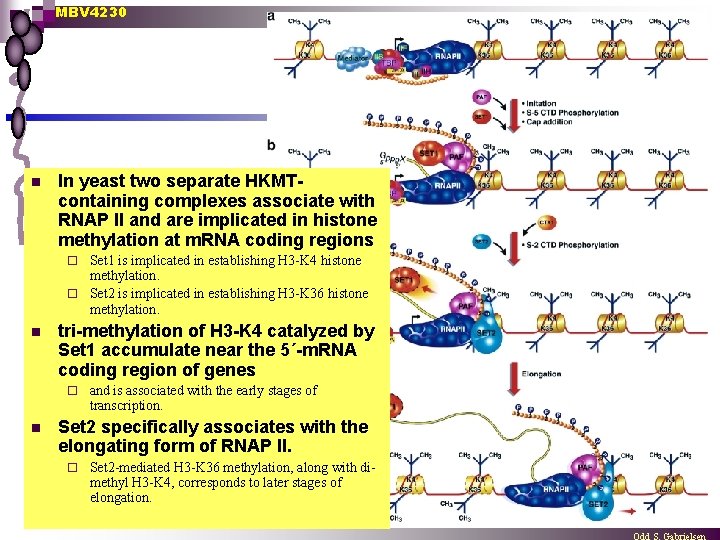
MBV 4230 n In yeast two separate HKMTcontaining complexes associate with RNAP II and are implicated in histone methylation at m. RNA coding regions Set 1 is implicated in establishing H 3 -K 4 histone methylation. ¨ Set 2 is implicated in establishing H 3 -K 36 histone methylation. ¨ n tri-methylation of H 3 -K 4 catalyzed by Set 1 accumulate near the 5´-m. RNA coding region of genes ¨ n and is associated with the early stages of transcription. Set 2 specifically associates with the elongating form of RNAP II. ¨ Set 2 -mediated H 3 -K 36 methylation, along with dimethyl H 3 -K 4, corresponds to later stages of elongation.
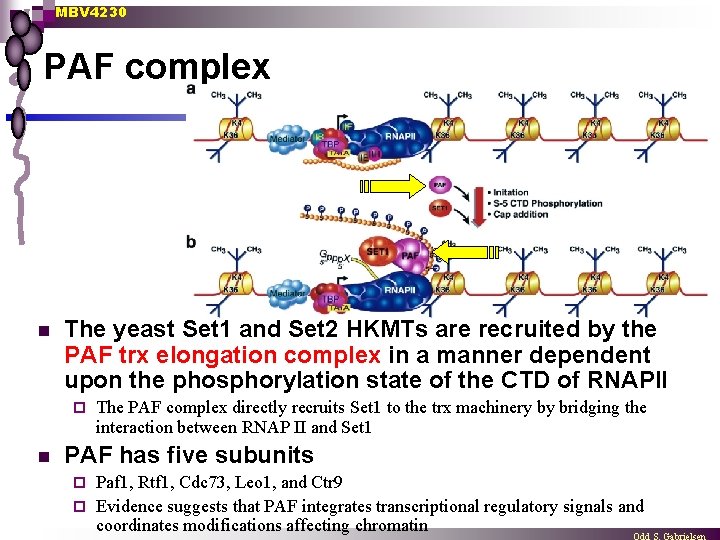
MBV 4230 PAF complex n The yeast Set 1 and Set 2 HKMTs are recruited by the PAF trx elongation complex in a manner dependent upon the phosphorylation state of the CTD of RNAPII ¨ n The PAF complex directly recruits Set 1 to the trx machinery by bridging the interaction between RNAP II and Set 1 PAF has five subunits Paf 1, Rtf 1, Cdc 73, Leo 1, and Ctr 9 ¨ Evidence suggests that PAF integrates transcriptional regulatory signals and coordinates modifications affecting chromatin ¨
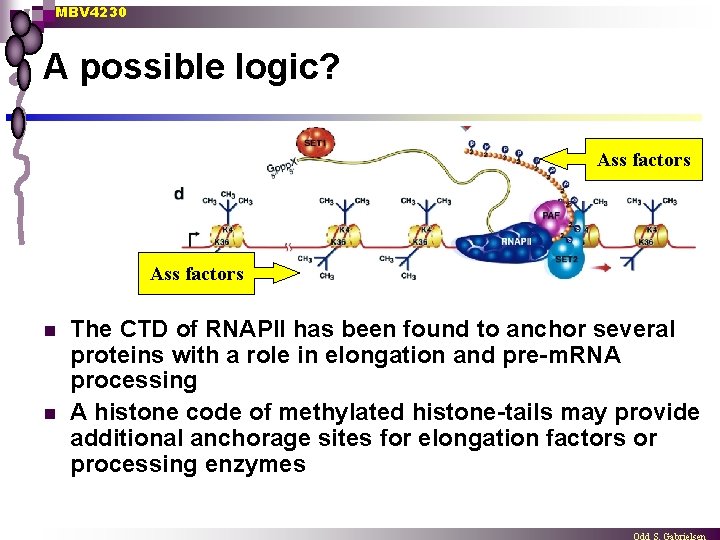
MBV 4230 A possible logic? Ass factors n n The CTD of RNAPII has been found to anchor several proteins with a role in elongation and pre-m. RNA processing A histone code of methylated histone-tails may provide additional anchorage sites for elongation factors or processing enzymes

Pre-m. RNA processing Processes tightly linked to elongation
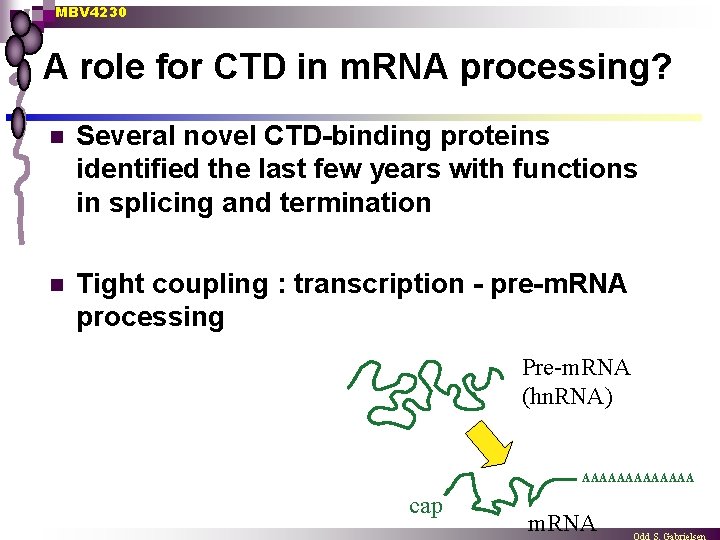
MBV 4230 A role for CTD in m. RNA processing? n Several novel CTD-binding proteins identified the last few years with functions in splicing and termination n Tight coupling : transcription - pre-m. RNA processing Pre-m. RNA (hn. RNA) AAAAAAA cap m. RNA

MBV 4230 CTD-mediated coupling : transcription - pre-m. RNA processing n Pre-m. RNA processing Capping ¨ Splicing ¨ Cleavage/polyadenylation ¨ n Physical contact between the machines for transcription and pre-m. RNA processing through CTD

Capping
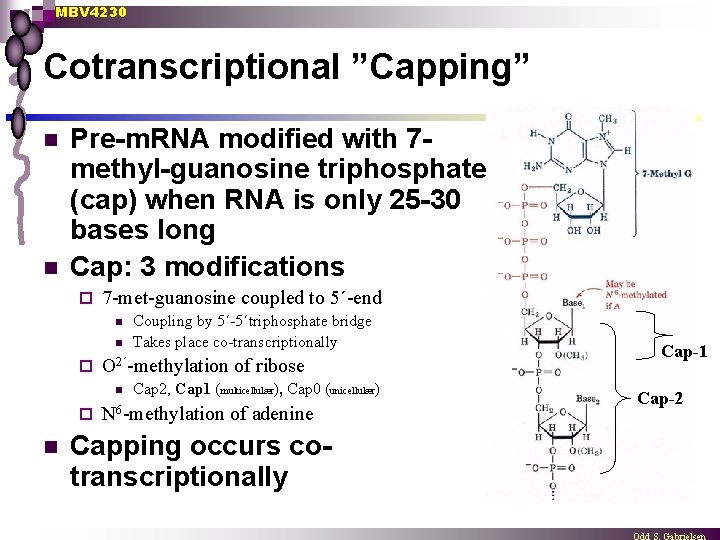
MBV 4230 Cotranscriptional ”Capping” n n Pre-m. RNA modified with 7 methyl-guanosine triphosphate (cap) when RNA is only 25 -30 bases long Cap: 3 modifications ¨ 7 -met-guanosine coupled to 5´-end n n ¨ O 2´-methylation n ¨ n Coupling by 5´-5´triphosphate bridge Takes place co-transcriptionally of ribose Cap 2, Cap 1 (multicellulær), Cap 0 (unicellulær) N 6 -methylation of adenine Capping occurs cotranscriptionally Cap-1 Cap-2
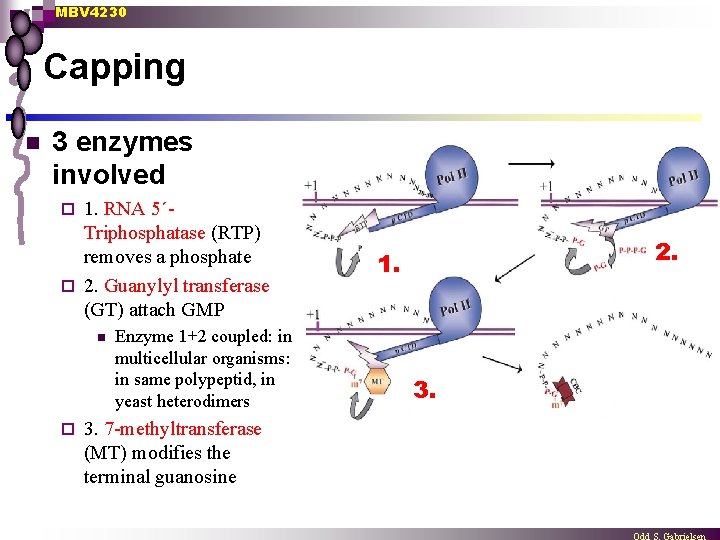
MBV 4230 Capping n 3 enzymes involved 1. RNA 5´Triphosphatase (RTP) removes a phosphate ¨ 2. Guanylyl transferase (GT) attach GMP ¨ n ¨ Enzyme 1+2 coupled: in multicellular organisms: in same polypeptid, in yeast heterodimers 3. 7 -methyltransferase (MT) modifies the terminal guanosine 2. 1. 3.

MBV 4230 Cotranscriptional ”Capping” n CTD recruits capping enzyme as soon as it is phosphorylated CTD required for effective capping ¨ Guanylyl transferase (mammalian + yeast) binds directly to phosphorylated CTD, not to non-phosphorylated ¨ 7 -methyltransferase (yeast) binds also phosphorylated CTD ¨ n n phosphorylated CTD may also regulate the activity of the enzymes Cap structure is recognized by CBC (Cap binding complex) Composed of two proteins CBP 20 and CBP 80 ¨ Major role in stabilization, block exonucleases ¨ CBC stimulates subsequent splicing and 3´-end processing ¨

Splicing

MBV 4230 Splicing n Splicing of introns occurs cotranscriptionally EM evidence ¨ Half-life BR 1 intron only 2. 5 min ≈ 5 kbs elongation of RNAPII ¨ n Splicing depends on CTD Inhibited by CTD truncation ¨ In vitro splicing stimulated by added phosphorylated CTD ¨ n CTD binds probably splicing-factors Not fully characterized ¨ CTD associated with SR- and Sm-splicing factors ¨
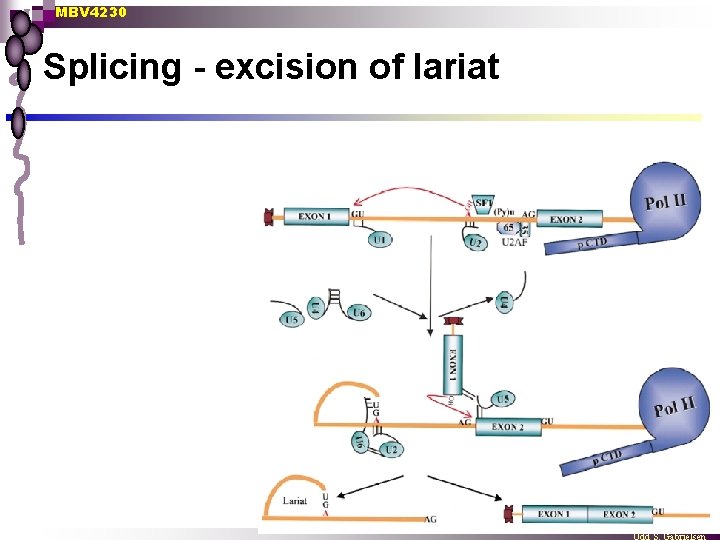
MBV 4230 Splicing - excision of lariat
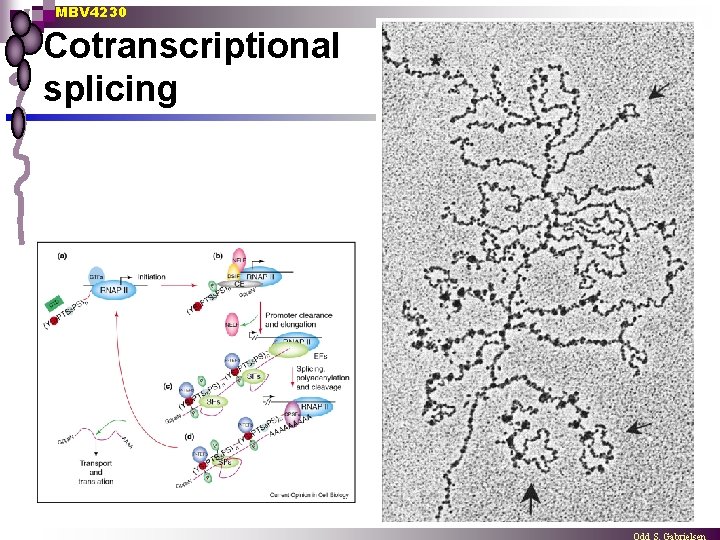
MBV 4230 Cotranscriptional splicing

MBV 4230 Association CTD-splicing factors n CTD binds probably splicing-factors CTD associated with SR- and Sm-splicing factors ¨ CASP (CTD-associated SR-like proteins) and SCAF (SR-like CTD-associated factors) ¨ RNA-binding proteins due to ¨ n n n RRM-domains target the factor to exon enhancer sequences RS-domains acting as ”glue” by forming RS-RS interactions Promoter-context can determine associated SR proteins and hence splicing Fibronectin: one intron included or excluded depending on the promoter ¨ Model: SR-CTD interaction set up during intiation, thus priming the elongation complex ¨ n Elongation rate can determine choice of alternative splice sites

3´-end formation

MBV 4230 Modification of 3´- end: poly-adenylation n Defined 3´-end is formed not by precise termination, but as a result of processing Pre-m. RNA heterogenous 3´-ends, ¨ m. RNA well defined 3´-ends ¨ n Poly(A) tails added in 3´-end ¨ Ca 200 x adenosines in a stretch of As added in a particular process n I. e. poly(A) not gene encoded AAAAAAA cap
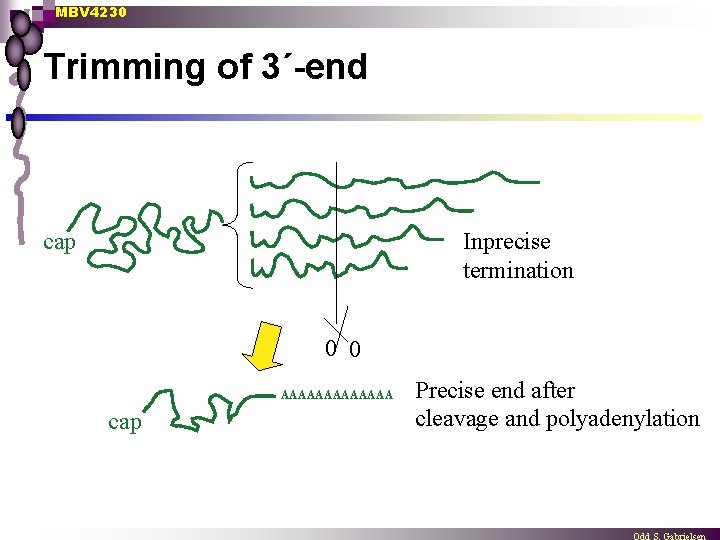
MBV 4230 Trimming of 3´-end cap Inprecise termination 0 0 AAAAAAA cap Precise end after cleavage and polyadenylation
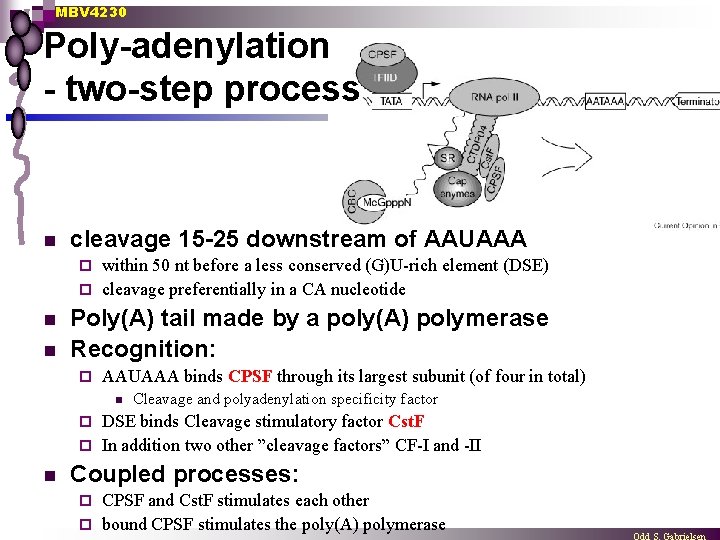
MBV 4230 Poly-adenylation - two-step process n cleavage 15 -25 downstream of AAUAAA within 50 nt before a less conserved (G)U-rich element (DSE) ¨ cleavage preferentially in a CA nucleotide ¨ n n Poly(A) tail made by a poly(A) polymerase Recognition: ¨ AAUAAA binds CPSF through its largest subunit (of four in total) n Cleavage and polyadenylation specificity factor DSE binds Cleavage stimulatory factor Cst. F ¨ In addition two other ”cleavage factors” CF-I and -II ¨ n Coupled processes: CPSF and Cst. F stimulates each other ¨ bound CPSF stimulates the poly(A) polymerase ¨
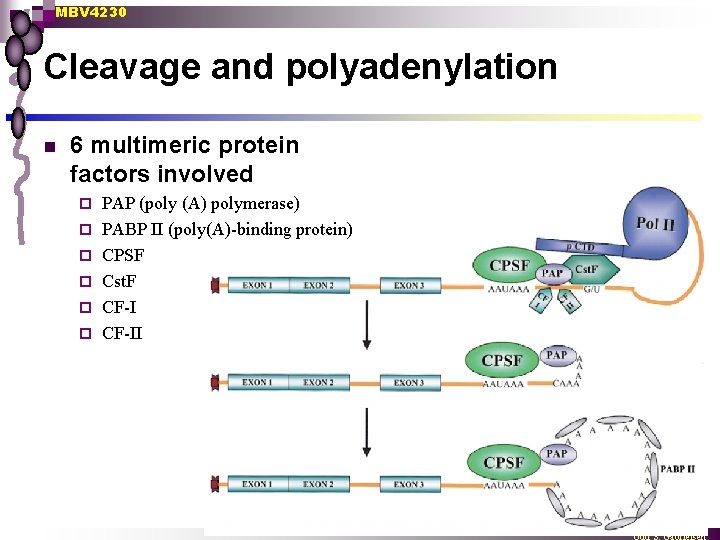
MBV 4230 Cleavage and polyadenylation n 6 multimeric protein factors involved ¨ ¨ ¨ PAP (poly (A) polymerase) PABP II (poly(A)-binding protein) CPSF Cst. F CF-II

MBV 4230 Processing of 3´-end: ”Cleavage/polyadenylation” n When RNAPII is approaching the 3´-end of the transcript, several coupled processes are taking place ¨ ¨ ¨ n Splicing of terminal intron cleavage at poly(A)-site, addition of poly(A) tail, termination downstream of poly(A)-site and liberation of RNAPII These av difficult to separate in time These processes depend on CTD Splicing, processing of 3´-end and termination downstream of poly(A) site are all inhibited by CTD truncations ¨ ”Cleavage-polyadenylation specificity factor” CPSF and ”cleavage stimulation factor” Cst. F bind specifically to CTD and are found associated with holo. RNAPII. ¨ Poly(A) polymerase is NOT associated with RNAPII ¨ CPSF is TBP-associated - becomes at some stage transferred from TFIID to CTD ¨
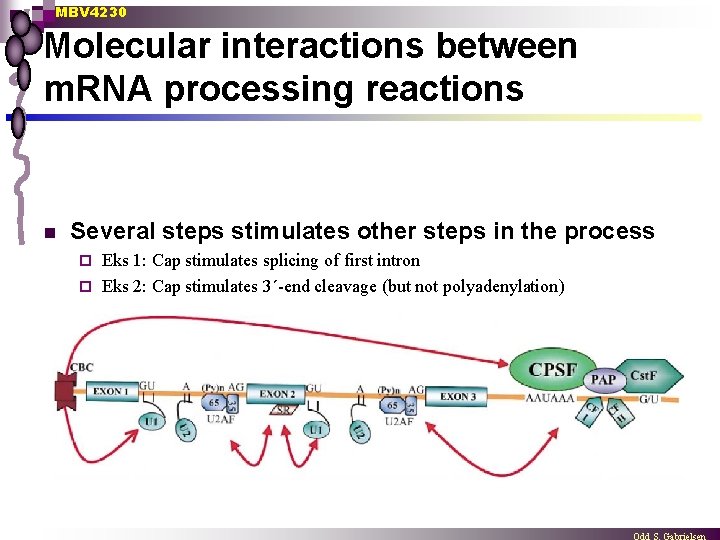
MBV 4230 Molecular interactions between m. RNA processing reactions n Several steps stimulates other steps in the process Eks 1: Cap stimulates splicing of first intron ¨ Eks 2: Cap stimulates 3´-end cleavage (but not polyadenylation) ¨
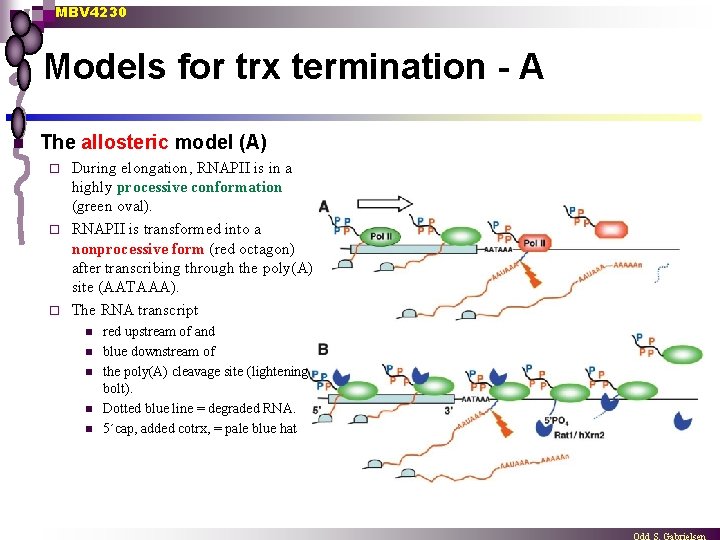
MBV 4230 Models for trx termination - A n The allosteric model (A) During elongation, RNAPII is in a highly processive conformation (green oval). ¨ RNAPII is transformed into a nonprocessive form (red octagon) after transcribing through the poly(A) site (AATAAA). ¨ The RNA transcript ¨ n n n red upstream of and blue downstream of the poly(A) cleavage site (lightening bolt). Dotted blue line = degraded RNA. 5´cap, added cotrx, = pale blue hat

MBV 4230 Models for trx termination - B n The torpedo model (B) RNA downstream of the poly(A) cleavage site (blue line) is digested by a 5´ 3´exonuclease (Rat 1 in yeast and h. Xrn 2 in humans (blue pacman), which tracks with RNAPII throughout the length of the gene. ¨ After poly(A) site cleavage, the exonuclease torpedo is guided along the RNA to its polymerase target and dissociates it from the DNA template. ¨
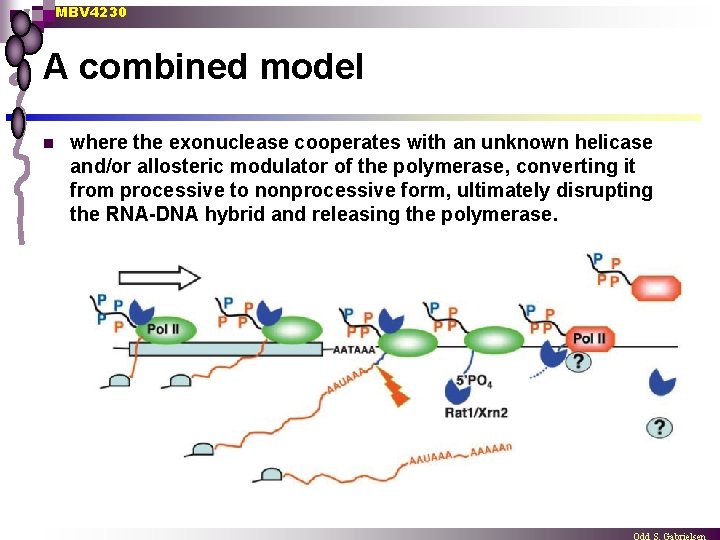
MBV 4230 A combined model n where the exonuclease cooperates with an unknown helicase and/or allosteric modulator of the polymerase, converting it from processive to nonprocessive form, ultimately disrupting the RNA-DNA hybrid and releasing the polymerase.
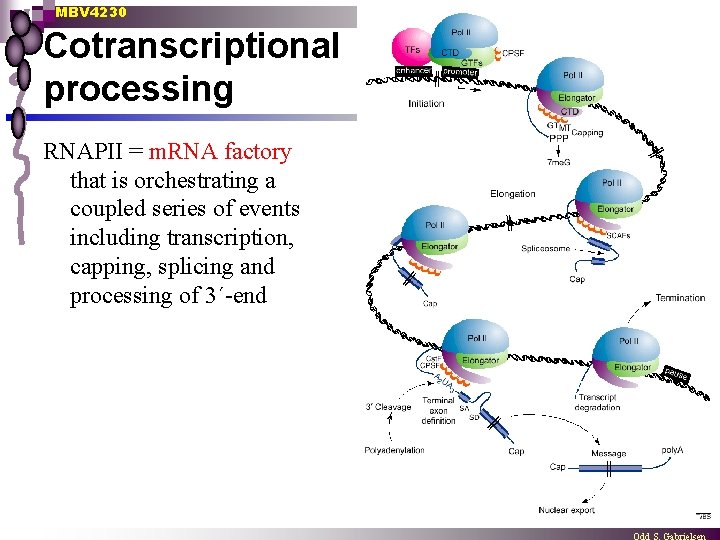
MBV 4230 Cotranscriptional processing RNAPII = m. RNA factory that is orchestrating a coupled series of events including transcription, capping, splicing and processing of 3´-end
- Slides: 64