Elementi ARE AU rich element AREbinding proteins Protein

Elementi ARE (AU rich element)
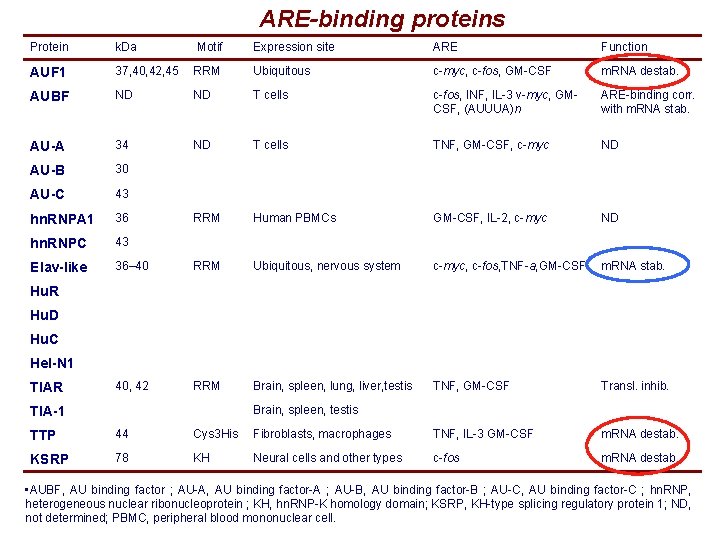
ARE-binding proteins Protein k. Da Motif Expression site ARE Function AUF 1 37, 40, 42, 45 RRM Ubiquitous c-myc, c-fos, GM-CSF m. RNA destab. AUBF ND ND T cells c-fos, INF, IL-3 v-myc, GMCSF, (AUUUA)n ARE-binding corr. with m. RNA stab. AU-A 34 ND T cells TNF, GM-CSF, c-myc ND AU-B 30 AU-C 43 hn. RNPA 1 36 RRM Human PBMCs GM-CSF, IL-2, c-myc ND hn. RNPC 43 Elav-like 36– 40 RRM Ubiquitous, nervous system c-myc, c-fos, TNF-a, GM-CSF m. RNA stab. 40, 42 RRM Brain, spleen, lung, liver, testis TNF, GM-CSF Transl. inhib. Hu. R Hu. D Hu. C Hel-N 1 TIAR Brain, spleen, testis TIA-1 TTP 44 Cys 3 His Fibroblasts, macrophages TNF, IL-3 GM-CSF m. RNA destab. KSRP 78 KH Neural cells and other types c-fos m. RNA destab. • AUBF, AU binding factor ; AU-A, AU binding factor-A ; AU-B, AU binding factor-B ; AU-C, AU binding factor-C ; hn. RNP, heterogeneous nuclear ribonucleoprotein ; KH, hn. RNP-K homology domain; KSRP, KH-type splicing regulatory protein 1; ND, not determined; PBMC, peripheral blood mononuclear cell.

ARE-binding proteins
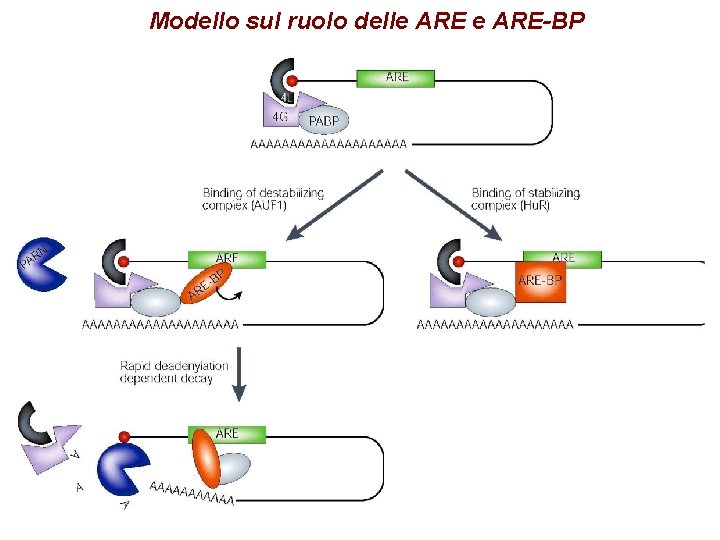
Modello sul ruolo delle ARE-BP
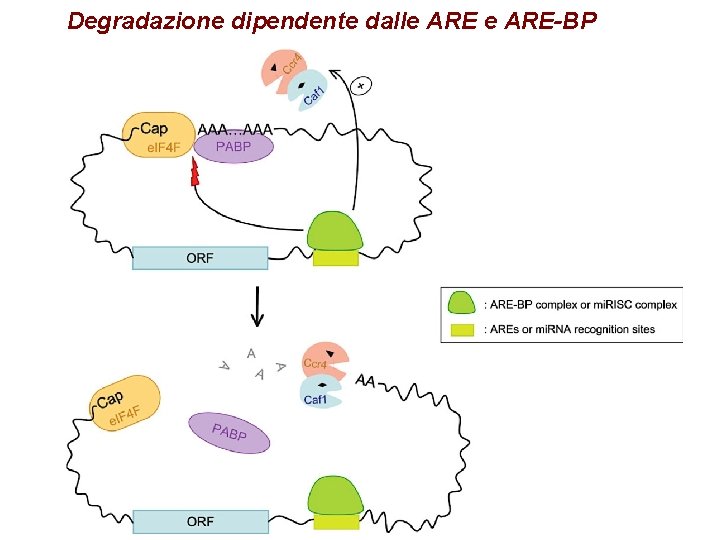
Degradazione dipendente dalle ARE-BP
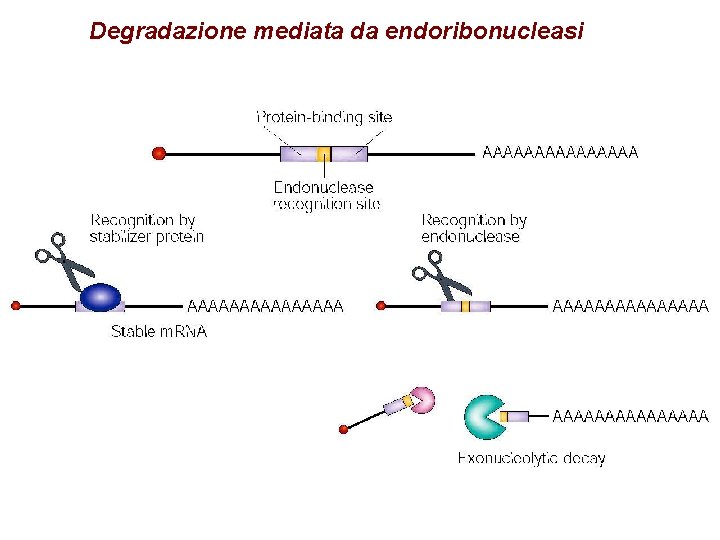
Degradazione mediata da endoribonucleasi
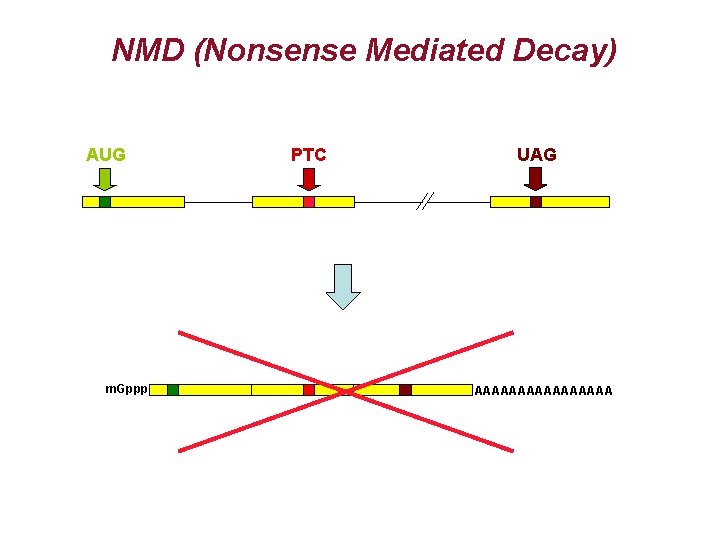
NMD (Nonsense Mediated Decay) AUG m. Gppp PTC UAG AAAAAAAA
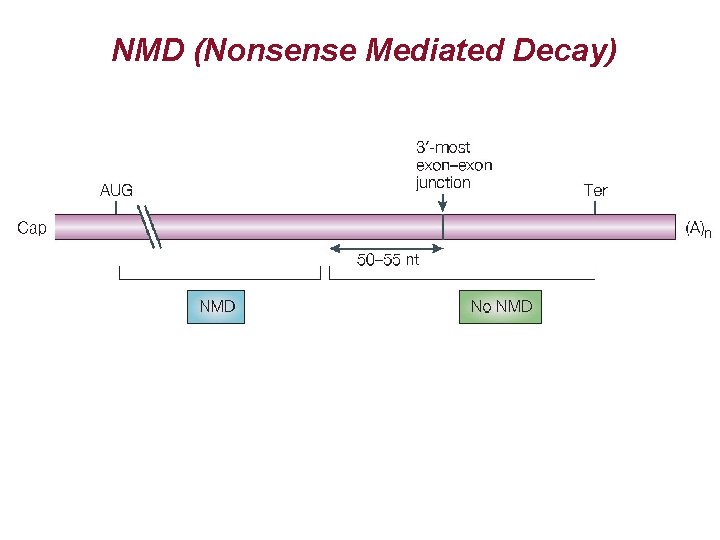
NMD (Nonsense Mediated Decay)
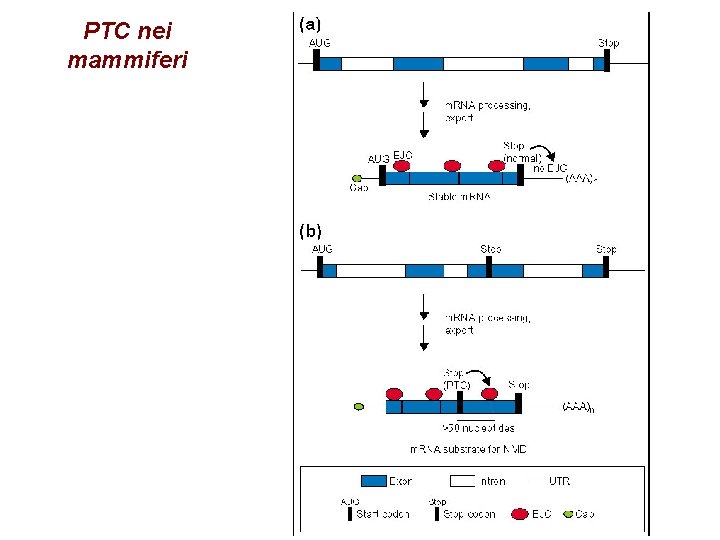
PTC nei mammiferi
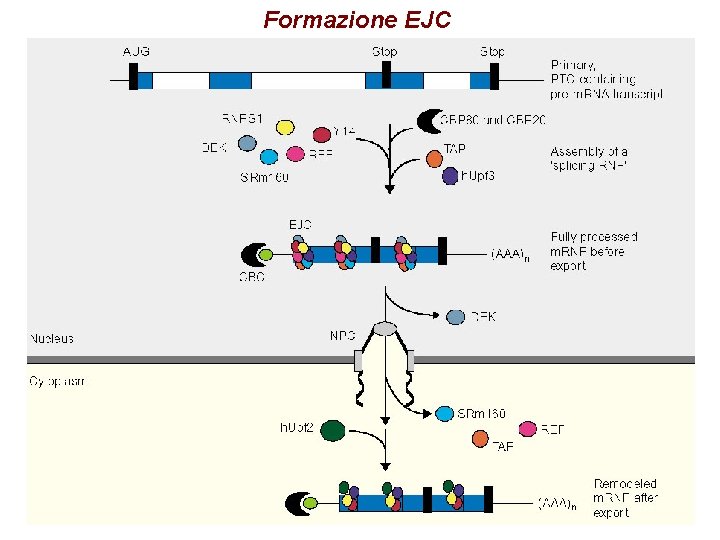
Formazione EJC
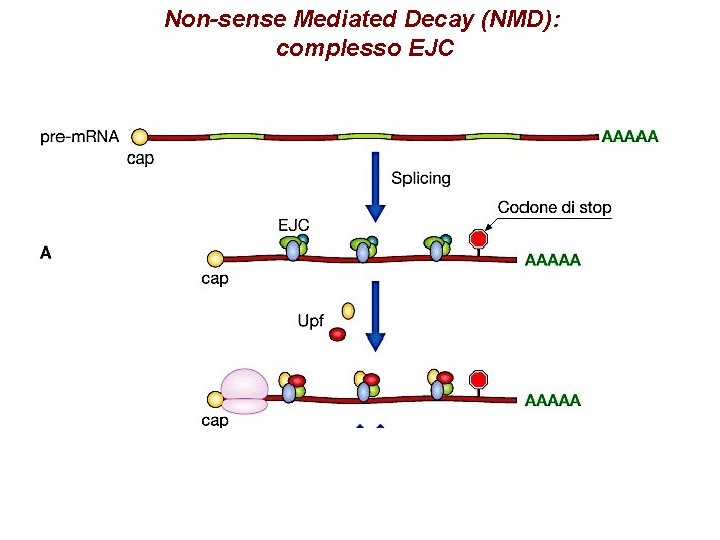
Non-sense Mediated Decay (NMD): complesso EJC
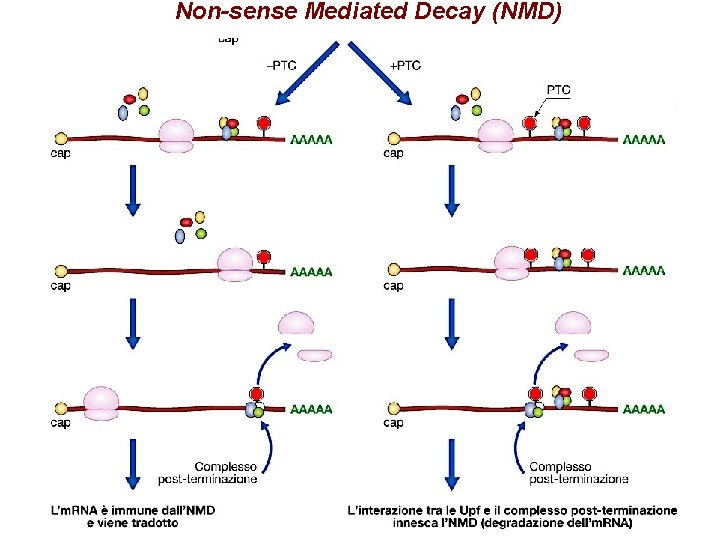
Non-sense Mediated Decay (NMD)
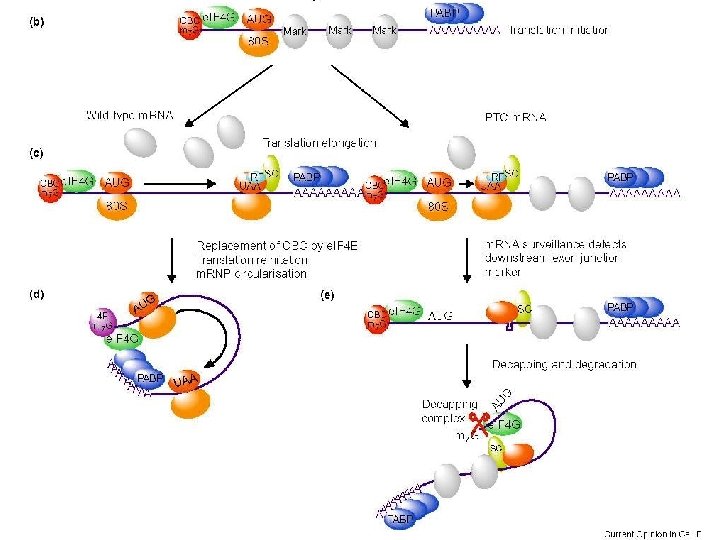
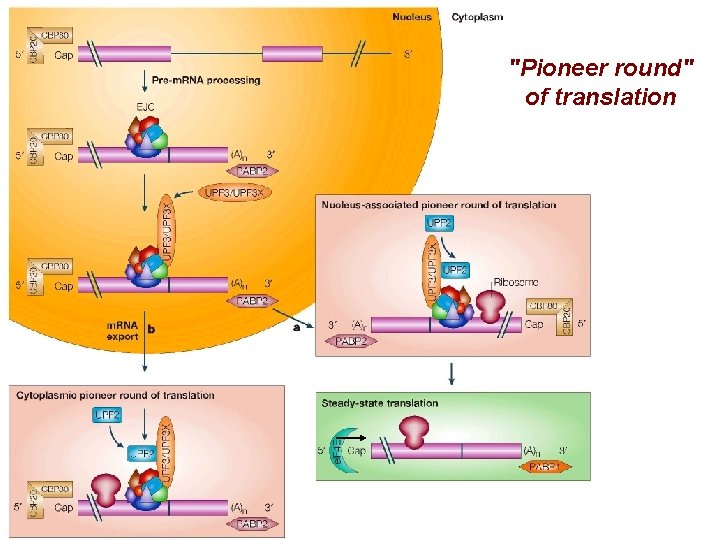
"Pioneer round" of translation
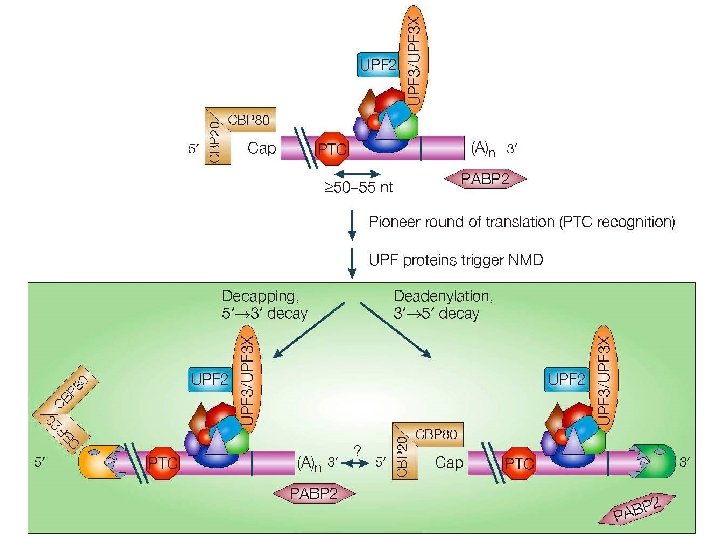
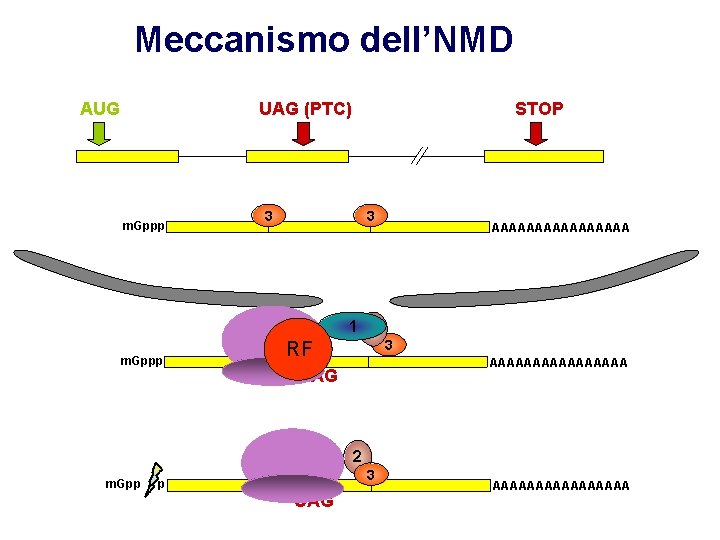
Meccanismo dell’NMD AUG UAG (PTC) m. Gppp 3 22 m. Gppp STOP 3 3 3 AAAAAAAA 1 2 RF 3 AAAAAAAA UAG 2 m. Gpp p 3 UAG AAAAAAAA
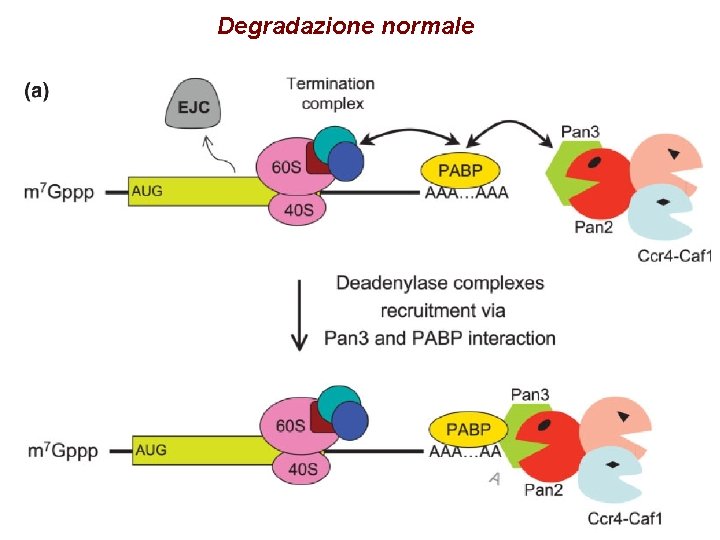
Degradazione normale
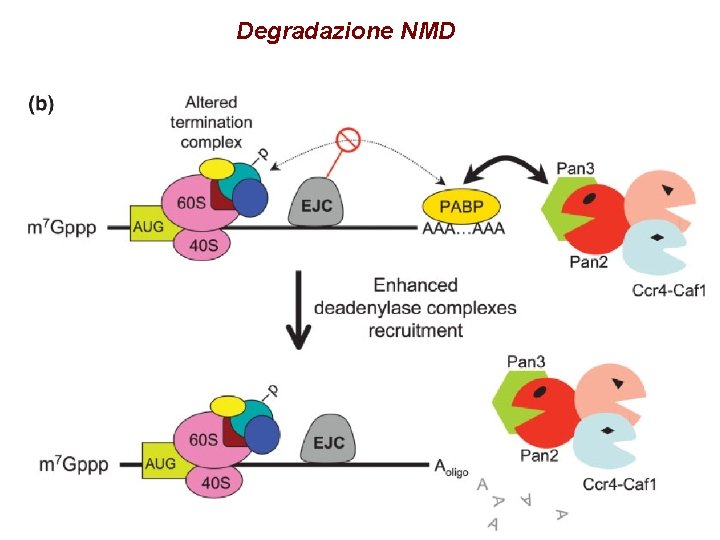
Degradazione NMD
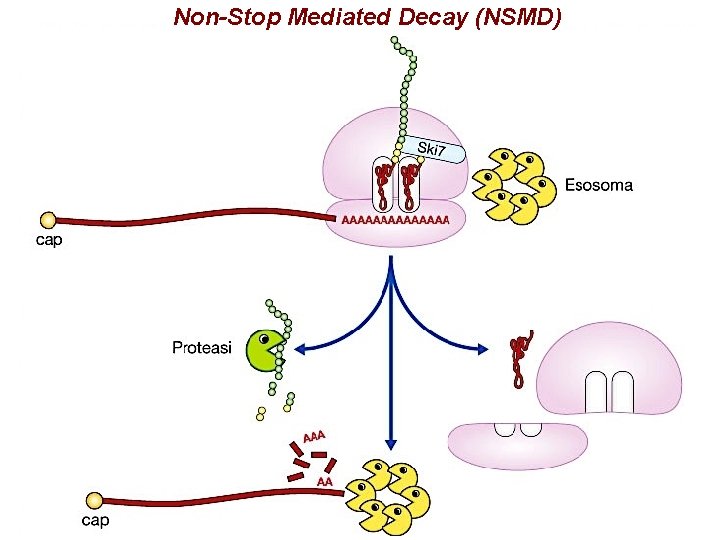
Non-Stop Mediated Decay (NSMD)
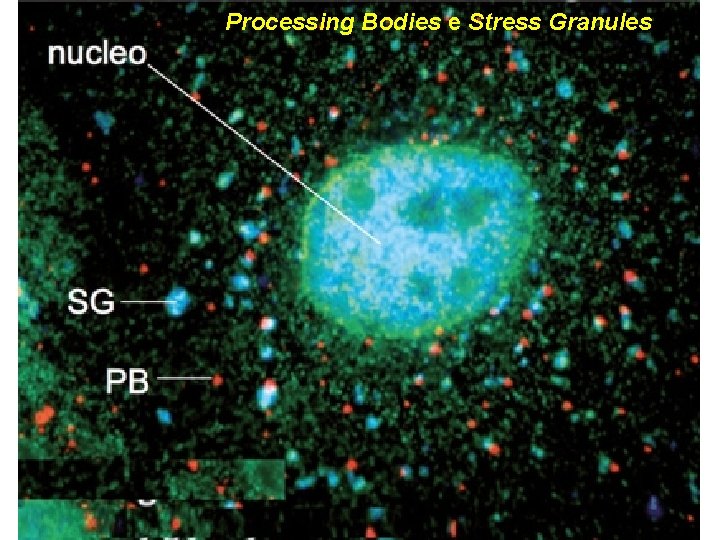
Processing Bodies e Stress Granules
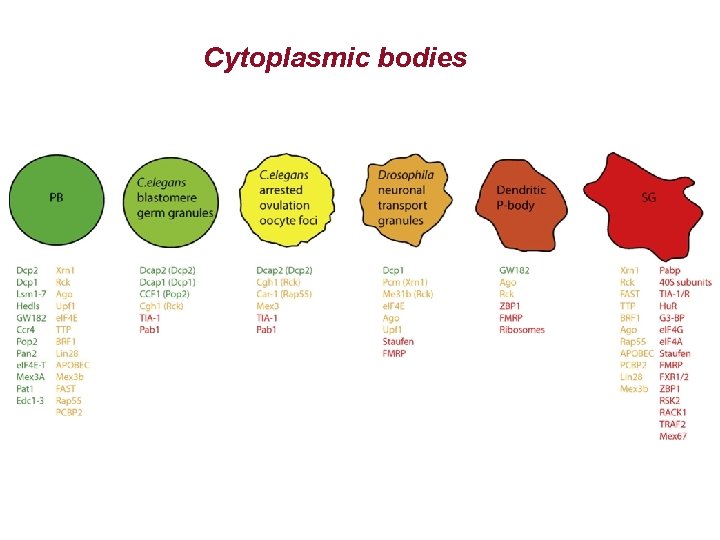
Cytoplasmic bodies
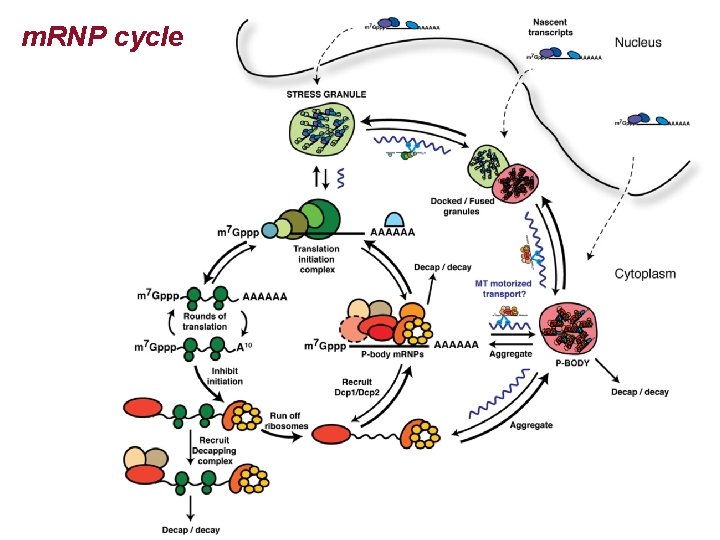
m. RNP cycle

RNA interference • Meccanismo • Significato biologico • Strumento di analisi della funzione dei geni

Co-soppressione Introduzione di copie transgeniche di un gene risulta nella ridotta espressione del transgene e del gene endogeno RNA interference Introduzione di RNA a doppio filamento (ds. RNA) induce silenziamento genico

Componenti dell’RNAi • si. RNA (small interfering RNA) = frammenti di RNA 21 -25 nt • Dicer = endoribonucleasi specifica per ds. RNA (tipo RNasi III) • RISC (RNA-induced silencing complex) = complesso ribonucleoproteico • Rd. RP (RNA-dependent RNA polymerase)
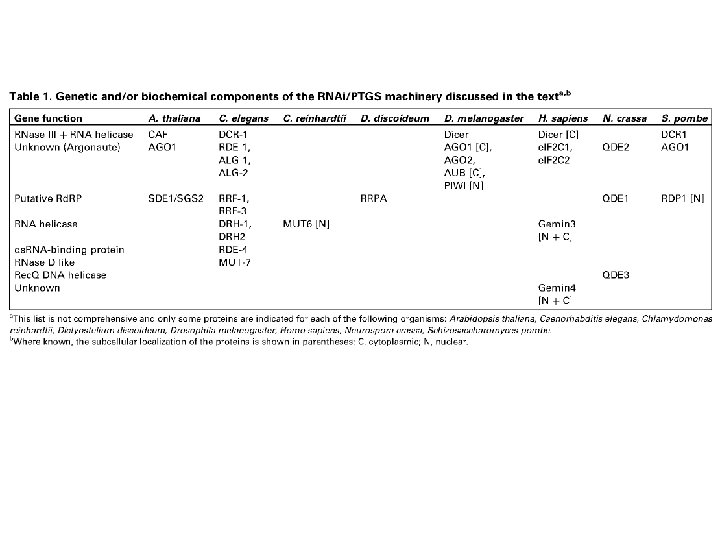

Inizio dell’RNAi • ds. RNA (80 -100 nt) in piante, C. elegans, Drosophila • si. RNA in mammiferi
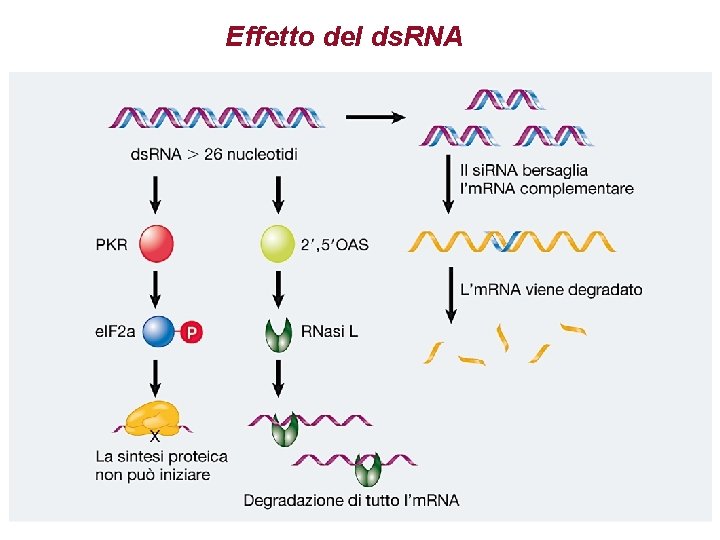
Effetto del ds. RNA
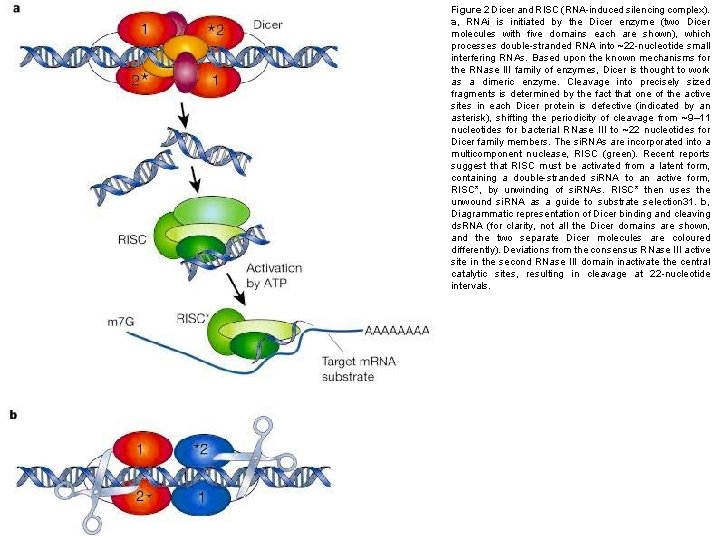
Figure 2 Dicer and RISC (RNA-induced silencing complex). a, RNAi is initiated by the Dicer enzyme (two Dicer molecules with five domains each are shown), which processes double-stranded RNA into ~22 -nucleotide small interfering RNAs. Based upon the known mechanisms for the RNase III family of enzymes, Dicer is thought to work as a dimeric enzyme. Cleavage into precisely sized fragments is determined by the fact that one of the active sites in each Dicer protein is defective (indicated by an asterisk), shifting the periodicity of cleavage from ~9– 11 nucleotides for bacterial RNase III to ~22 nucleotides for Dicer family members. The si. RNAs are incorporated into a multicomponent nuclease, RISC (green). Recent reports suggest that RISC must be activated from a latent form, containing a double-stranded si. RNA to an active form, RISC*, by unwinding of si. RNAs. RISC* then uses the unwound si. RNA as a guide to substrate selection 31. b, Diagrammatic representation of Dicer binding and cleaving ds. RNA (for clarity, not all the Dicer domains are shown, and the two separate Dicer molecules are coloured differently). Deviations from the consensus RNase III active site in the second RNase III domain inactivate the central catalytic sites, resulting in cleavage at 22 -nucleotide intervals.

Meccanismo 1. Produzione si. RNA 2. Formazione complesso RISC 3. Attivazione complesso RISC 4. Taglio dell’RNA bersaglio
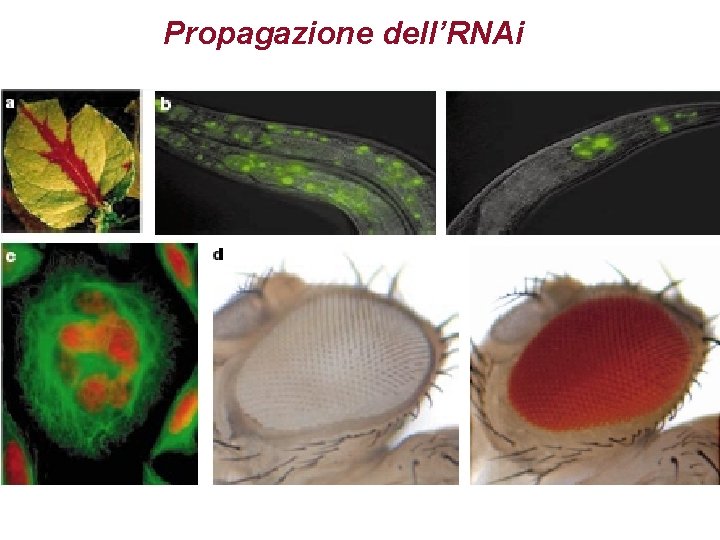
Propagazione dell’RNAi
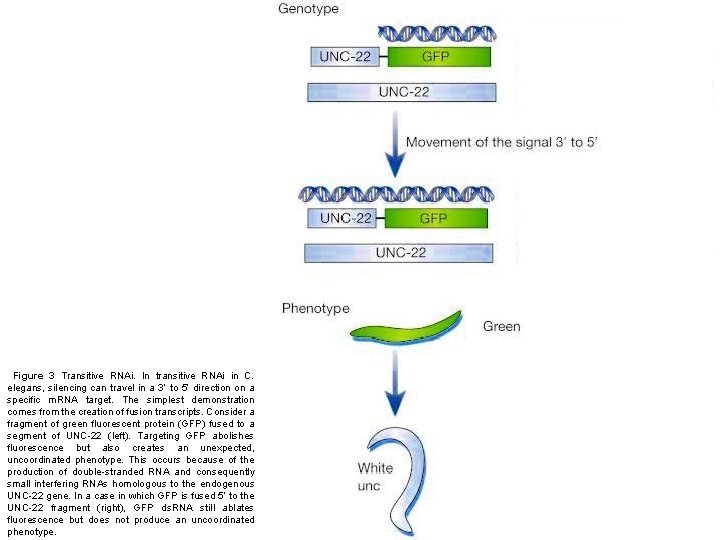
Figure 3 Transitive RNAi. In transitive RNAi in C. elegans, silencing can travel in a 3’ to 5’ direction on a specific m. RNA target. The simplest demonstration comes from the creation of fusion transcripts. Consider a fragment of green fluorescent protein (GFP) fused to a segment of UNC-22 (left). Targeting GFP abolishes fluorescence but also creates an unexpected, uncoordinated phenotype. This occurs because of the production of double-stranded RNA and consequently small interfering RNAs homologous to the endogenous UNC-22 gene. In a case in which GFP is fused 5’ to the UNC-22 fragment (right), GFP ds. RNA still ablates fluorescence but does not produce an uncoordinated phenotype.
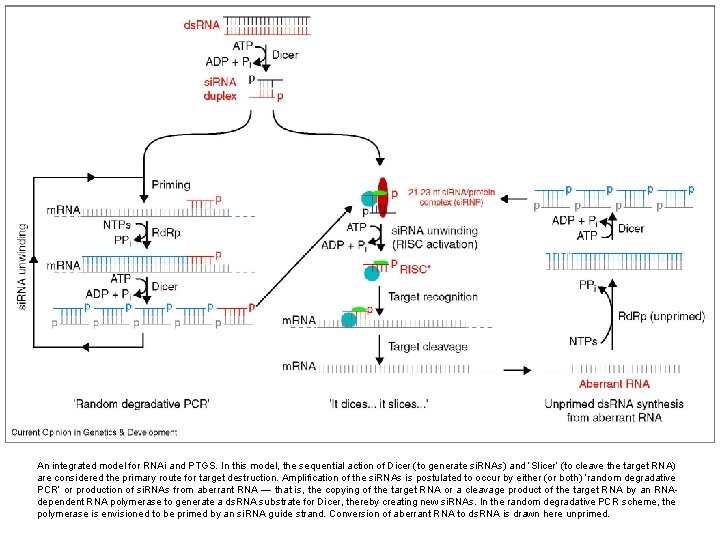
An integrated model for RNAi and PTGS. In this model, the sequential action of Dicer (to generate si. RNAs) and ‘Slicer’ (to cleave the target RNA) are considered the primary route for target destruction. Amplification of the si. RNAs is postulated to occur by either (or both) ‘random degradative PCR’ or production of si. RNAs from aberrant RNA — that is, the copying of the target RNA or a cleavage product of the target RNA by an RNAdependent RNA polymerase to generate a ds. RNA substrate for Dicer, thereby creating new si. RNAs. In the random degradative PCR scheme, the polymerase is envisioned to be primed by an si. RNA guide strand. Conversion of aberrant RNA to ds. RNA is drawn here unprimed.
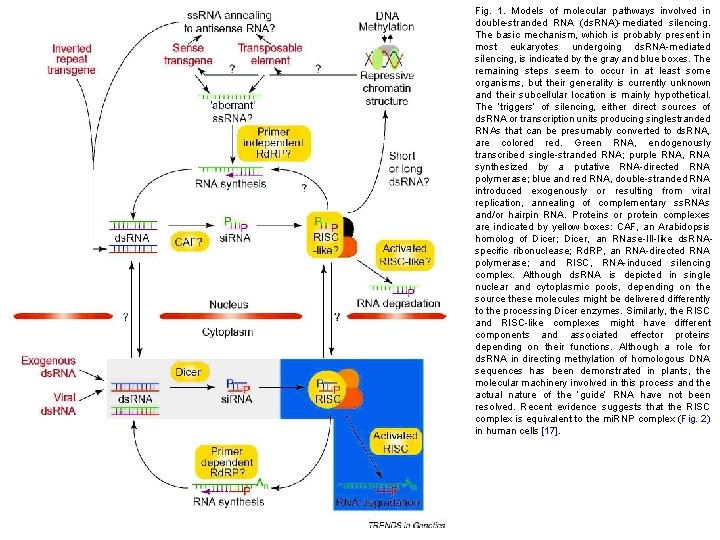
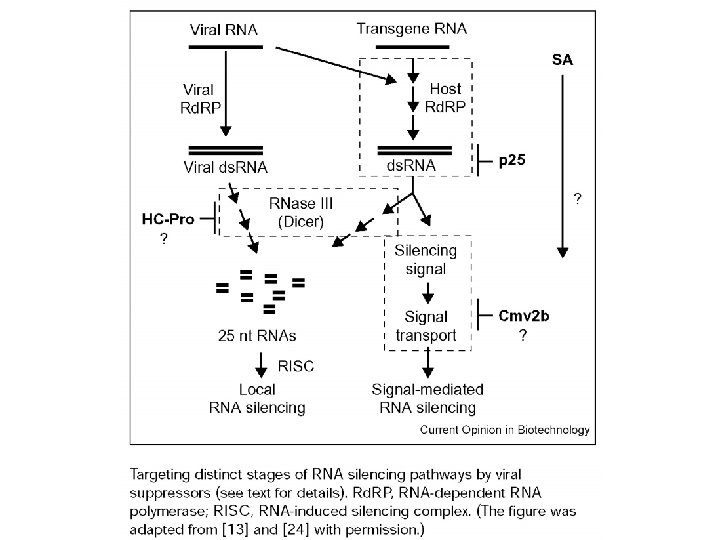
Fig. 1. Models of molecular pathways involved in double-stranded RNA (ds. RNA)-mediated silencing. The basic mechanism, which is probably present in most eukaryotes undergoing ds. RNA-mediated silencing, is indicated by the gray and blue boxes. The remaining steps seem to occur in at least some organisms, but their generality is currently unknown and their subcellular location is mainly hypothetical. The ‘triggers’ of silencing, either direct sources of ds. RNA or transcription units producing singlestranded RNAs that can be presumably converted to ds. RNA, are colored red. Green RNA, endogenously transcribed single-stranded RNA; purple RNA, RNA synthesized by a putative RNA-directed RNA polymerase; blue and red RNA, double-stranded RNA introduced exogenously or resulting from viral replication, annealing of complementary ss. RNAs and/or hairpin RNA. Proteins or protein complexes are indicated by yellow boxes: CAF, an Arabidopsis homolog of Dicer; Dicer, an RNase-III-like ds. RNAspecific ribonuclease; Rd. RP, an RNA-directed RNA polymerase; and RISC, RNA-induced silencing complex. Although ds. RNA is depicted in single nuclear and cytoplasmic pools, depending on the source these molecules might be delivered differently to the processing Dicer enzymes. Similarly, the RISC and RISC-like complexes might have different components and associated effector proteins depending on their functions. Although a role for ds. RNA in directing methylation of homologous DNA sequences has been demonstrated in plants, the molecular machinery involved in this process and the actual nature of the ‘guide’ RNA have not been resolved. Recent evidence suggests that the RISC complex is equivalent to the mi. RNP complex (Fig. 2) in human cells [17].

Caratteristiche dell’RNAi • Può agire a diversi livelli: trascrizionale, posttrascrizionale, traduzionale • Può amplificarsi e diffondersi nell’organismo • Può coinvolgere sequenze adiacenti
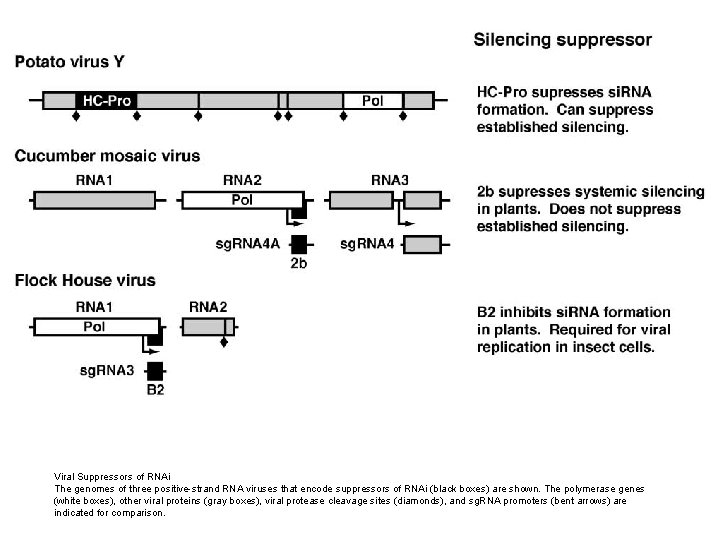
Viral Suppressors of RNAi The genomes of three positive-strand RNA viruses that encode suppressors of RNAi (black boxes) are shown. The polymerase genes (white boxes), other viral proteins (gray boxes), viral protease cleavage sites (diamonds), and sg. RNA promoters (bent arrows) are indicated for comparison.

Controllo di acidi nucleici “parassiti” • C. elegans senza RNAi hanno una aumentata mobilità di trasposoni endogeni • In alcuni sistemi i trasposoni “silenziati” si trovano organizzati in eterocromatina Regolazione dell’espressione genica • Mutazioni in componenti dell’RNAi causano alterazioni dello sviluppo (Arabidopsis, C. elegans, Drosophila) • Regolazione del gene Stellate in Drosophila

Ruolo biologico dell’RNAi • Difesa dai virus (nelle piante VISG) • Controllo dei trasposoni • Regolazione dell’espressione genica

Uso dell’RNAi • Studio sistematico della funzione dei geni • Terapia genica
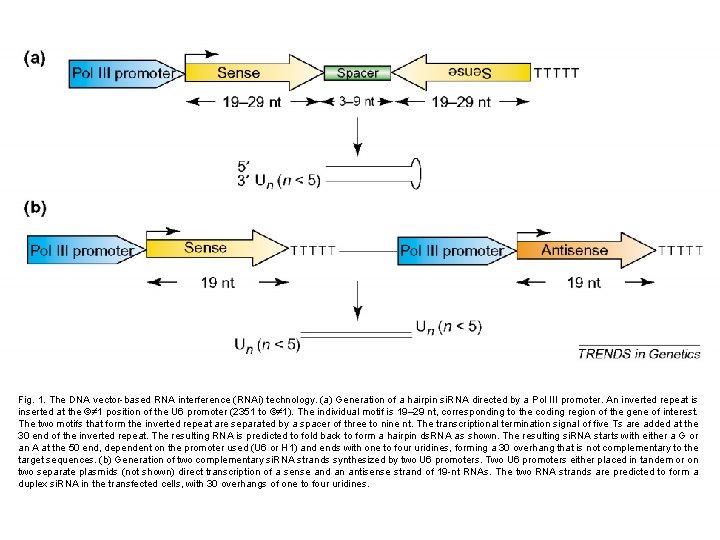
Fig. 1. The DNA vector-based RNA interference (RNAi) technology. (a) Generation of a hairpin si. RNA directed by a Pol III promoter. An inverted repeat is inserted at the ©≠ 1 position of the U 6 promoter (2351 to ©≠ 1). The individual motif is 19– 29 nt, corresponding to the coding region of the gene of interest. The two motifs that form the inverted repeat are separated by a spacer of three to nine nt. The transcriptional termination signal of five Ts are added at the 30 end of the inverted repeat. The resulting RNA is predicted to fold back to form a hairpin ds. RNA as shown. The resulting si. RNA starts with either a G or an A at the 50 end, dependent on the promoter used (U 6 or H 1) and ends with one to four uridines, forming a 30 overhang that is not complementary to the target sequences. (b) Generation of two complementary si. RNA strands synthesized by two U 6 promoters. Two U 6 promoters either placed in tandem or on two separate plasmids (not shown) direct transcription of a sense and an antisense strand of 19 -nt RNAs. The two RNA strands are predicted to form a duplex si. RNA in the transfected cells, with 30 overhangs of one to four uridines.

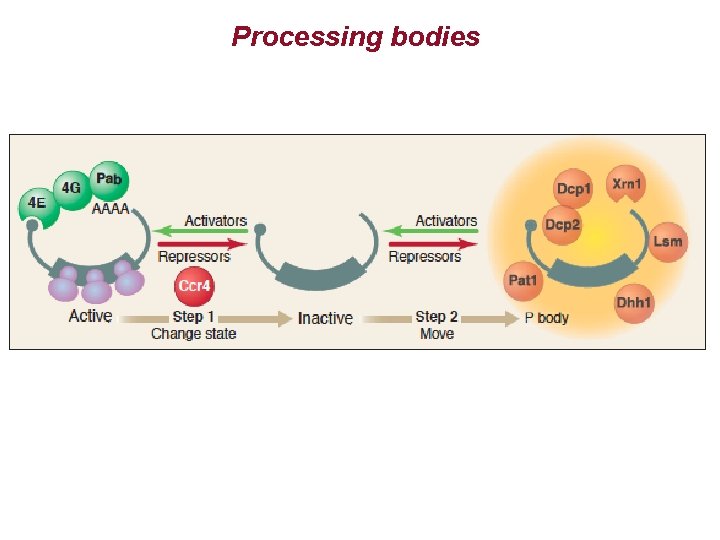
Processing bodies
- Slides: 43