Effectiveness of DTGPCR for Biased DNA Algorithms Ji

Effectiveness of DTG-PCR for Biased DNA Algorithms Ji Youn Lee
Goal of DNA Computing Computer Science Physics Smart DNA word design Mathematics Engineering Development of efficient and smart algorithms Chemistry 9/15/2020 Development of new biological tools Biology 2

Polymerase Chain Reaction (PCR) • In vitro amplification of nucleic acids • Thermus aquaticus (Taq) DNA polymerase • Consists of three reactions at different temperatures – Denaturation/ annealing/ extension • Cyclic reaction • Essential tool in molecular biology/DNA computing 9/15/2020 3
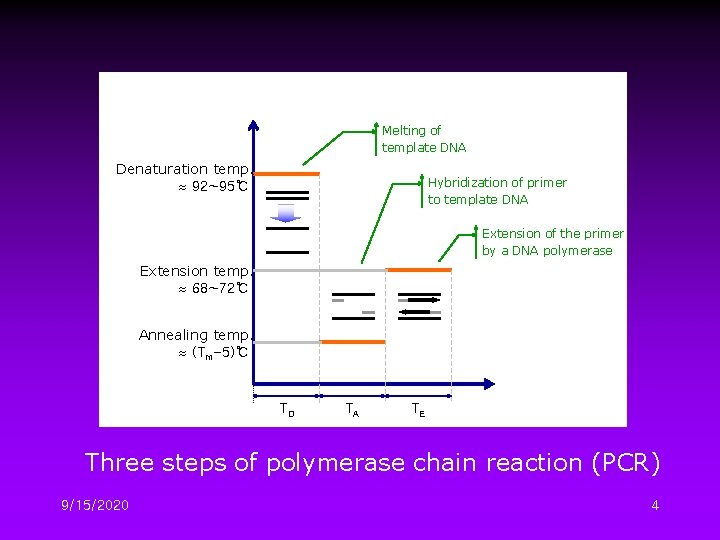
Melting of template DNA Denaturation temp. 92~95℃ Hybridization of primer to template DNA Extension of the primer by a DNA polymerase Extension temp. 68~72℃ Annealing temp. (Tm-5)℃ TD TA TE Three steps of polymerase chain reaction (PCR) 9/15/2020 4

DTG-PCR • Stands for… – Denaturation Temperature Gradient-Polymerase Chain Reaction • Temperature gradient in denaturation reaction with the cycle progress • Selectively more amplify the DNA strands of lower melting temperature 9/15/2020 5

Temperature Gradient Encoding • Traveling salesman problem • Weight (real number) representation • Temperature related tools – DTG-PCR – TGGE • The range and limit of DTG-PCR 9/15/2020 6

Theoretical Description

Necessity of Theoretical Description • Understanding of the reaction mechanism • The groundwork for modeling • For a prediction of experimental result 9/15/2020 8

Formal Approaches I • Efficiency-based expression: efficiency of each cycle Anal. Biol. Chem. 214 (1993) pp 582 -585 "Quantitative PCR: Theoretical considerations with practical implications" 9/15/2020 9

Formal Approaches II • Enzymological considerations J. theor. Biol. 184 (1997) pp 433 -440 "Enzymological Considerations for a Theoretical Description of the Quantitative Competitive Polymerase Chain Reaction" 9/15/2020 10

9/15/2020 11

9/15/2020 12

9/15/2020 13

9/15/2020 14

9/15/2020 15

Formal Approaches III • Probabilistic consideration • Markov process model for primer extension J. theor. Biol. 201 (1999) pp 239 -249 "Polymerase Chain Reaction: A Morkov Process Approach" 9/15/2020 16

Rate or Efficiency of the PCR • Influenced by the factors… – Concentrations of template DNA, DNA polymerase, d. NTPs, Mg. Cl 2 and primers – Denaturation, annealing and extension temperatures – Time and number of cycles – Ramping rates (of the machine) – The presence of contaminating DNA and inhibitors in the sample – Etc… Appropriate assumptions are necessary!! 9/15/2020 17

Causes of Plateau • Increase of template – Decrease of the ratio of enzyme to template – Renaturation between templates • Inactivation of polymerase – Binding or extension efficiency • Exhaustion of d. NTPs, primers – Efficient concentrations 9/15/2020 18

Basic Assumptions • Discrete reactions – One reaction in one step • Optimized reaction condition – – Optimized annealing temperature Optimized buffer concentration Plentiful reaction time No existence of intermediate molecules of insufficient length (assumption of complete extension) 9/15/2020 19

Optimization Conditions • Mg. Cl 2 concentration: 3. 0~6. 0 m. M • d. NTP concentration: 200~600 m. M each • Taq DNA polymerase: 1. 25~4. 5 U/50 ml reaction • Primer concentration: 100~500 n. M • An symmetric primer concentration • Probe or dye concentration 9/15/2020 20

Notation Summary Nds number of double-stranded DNA or template Nss number of single-stranded DNA or template eds molar extinction coefficient of double-stranded DNA ess molar extinction coefficient of single-stranded DNA Np number of primer Nhd number of heteroduplex molecule Q number of polymerase Q’ effective number of polymeraes Ncomplex Nd. NTP Nextended(i) Ncomplex’ 9/15/2020 number of heteroduplex combined with polymerase number of d. NTPs number of extended heteroduplex by i mer number of fully extended molecule combined with polymerase 21

Denaturation Step • Dissociation of Watson-Crick complement • Can be considered the reverse compliment to denaturating • Denaturation efficiency calculation Two possible approaches • Melting curve-based approach • Kinetics-based approach 9/15/2020 22

Melting Curve-Based Approach
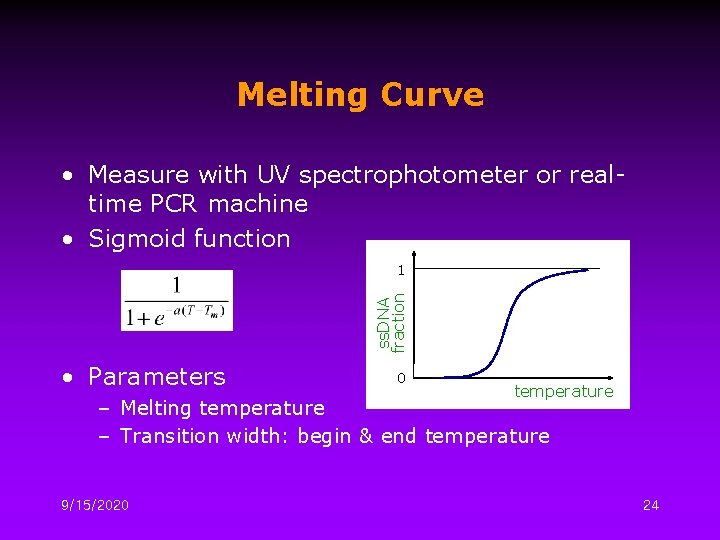
Melting Curve • Measure with UV spectrophotometer or realtime PCR machine • Sigmoid function ss. DNA fraction 1 • Parameters 0 temperature – Melting temperature – Transition width: begin & end temperature 9/15/2020 24

Melting Temperature, Tm • The Tm is the temperature at which half the DNA is present in a single-stranded (denatured) form. • Primarily influenced by four parameters – – Temperature p. H Concentration of monovalent cations Presence of organic solvents 9/15/2020 25
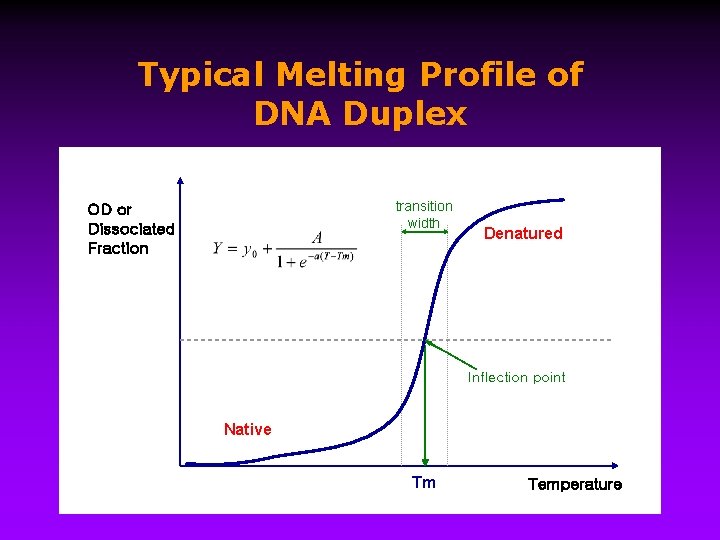
Typical Melting Profile of DNA Duplex transition width OD or Dissociated Fraction Denatured Inflection point Native Tm 9/15/2020 Temperature 26
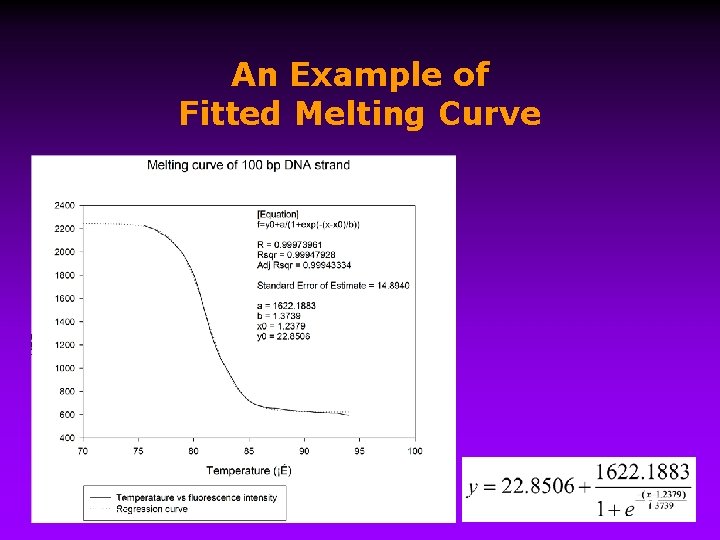
An Example of Fitted Melting Curve 9/15/2020 27

Hyperchromic Effect • Calculation of the ss. DNA fraction from the optical density and extinction coefficients ( ) 9/15/2020 28

Denaturation Efficiency, hd • Denaturation efficiency can be calculated from the melting curve. 9/15/2020 29

Annealing Step • Hybridization of single-stranded DNA template and primer • Factors – Single-stranded DNA template concentration – Primer concentration Cp/Css (r) and Css 9/15/2020 30

Can Be Considered as … • Competition between… – Renaturation of Watson strand Crick strand – Hybridization of single-stranded DNA template and primer 9/15/2020 31

vs Competition between ka and kd-1 9/15/2020 32

Denaturation of ds. DNA to ss. DNA Formation rate of ss. DNA Temperature dependency of reaction constant Arrhenius equation 9/15/2020 33

9/15/2020 34
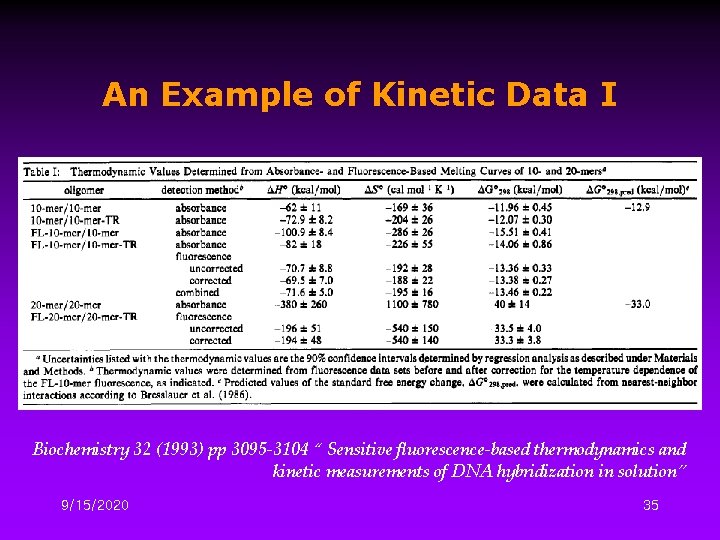
An Example of Kinetic Data I Biochemistry 32 (1993) pp 3095 -3104 “ Sensitive fluorescence-based thermodynamics and kinetic measurements of DNA hybridization in solution” 9/15/2020 35

An Example of Kinetic Data II Proc. Natl. Acad. Sci. USA 95 (1998) pp 1460 -1465 “A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics” 9/15/2020 36

NACST/seq Calculation of DH and DS from NN data Data from papers of Santa Lucia Calculation of equilibrium constant Calculation of ss. DNA concentration 9/15/2020 37

Extension Step • Divided into three reactions (1) Binding of polymerase to the heteroduplex (2) Successive incorporation of d. NTPs by polymerase (3) Detachment of polymerase from fully extended DNA 9/15/2020 38

9/15/2020 39

Assumptions in Extension Step 1), 3) are dependent on the binding and release activity of polymerase 2) is dependent on the incorporation activity of polymerase • Assumption – Binding efficiency and detachment efficiency is unity – Therefore, the efficiency of the extension reaction only rely on the d. NTP incorporation rate. – The incorporation rate of each d. NTPs is same. 9/15/2020 40

Factors Affecting the Extension Efficiency • Polymerase activity • Heteroduplex concentration 9/15/2020 41
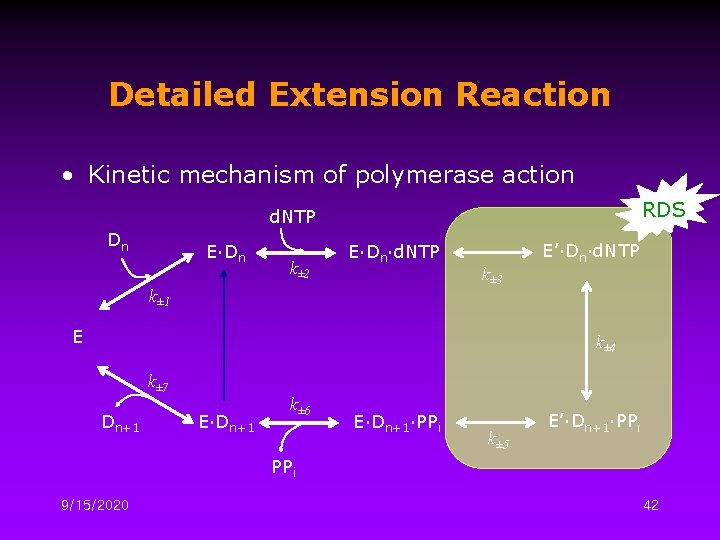
Detailed Extension Reaction • Kinetic mechanism of polymerase action RDS d. NTP Dn E∙Dn k± 2 E∙Dn∙d. NTP k± 1 k± 3 E E’∙Dn∙d. NTP k± 4 k± 7 Dn+1 E∙Dn+1 k± 6 E∙Dn+1∙PPi k± 5 E’∙Dn+1∙PPi 9/15/2020 42

Parameter Value Kd 1 5 n. M Binding of template with polymerase Kd 2 5 m. M d. NTP incorporation to form a ternary complex K 4 2. 5 Chemical catalysis Kd 6 100 μM Release of PPi k 3 50 sec-1 k-3 0. 5 sec-1 k 4 1000 sec-1 k-4 400 sec-1 k 5 15 sec-1 k-5 sec-1 k 7 k-7 9/15/2020 15 M-1 Chemical catalysis Second conformational change 0. 06 sec-1 1. 2 107 Conformational change of polymerase sec-1 Dissociation of polymerase 43

Enzyme Inactivation • Causes – Enzyme aggregation, or enzyme heterogeneity, or formation of partially deactivated forms of the enzyme • Thermal inactivation • Reversible vs irreversible • Due either – to the loss of the native conformation (without rupture or formation of covalent bonds) – or to chemical modification of functional groups of the active site 9/15/2020 44

Models of Enzyme Inactivation • • First-order model Bi-exponential models Isozyme model Lumry-Eyring model 9/15/2020 45

Kinetics of Enzyme Inactivation • Common assumptions; – Assume that thermal inactivation mechanisms involve a single active enzyme form, i. e. , the native one. – Assume 1 st order kinetics with respect to the native enzyme concentration. An exponential activity decay is predicted However, there are exceptions… Due to the occurrence of multiple active forms 9/15/2020 46

– Irreversible reaction – Involve only two different enzyme forms: native (active) and denatured 9/15/2020 47
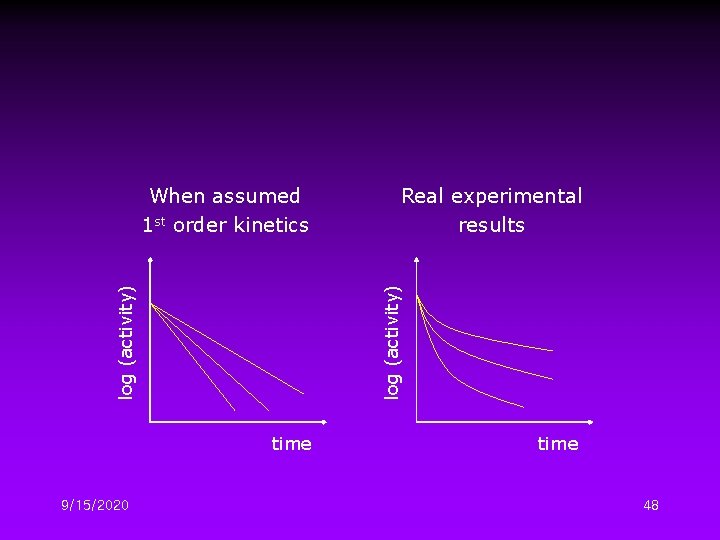
time 9/15/2020 Real experimental results log (activity) When assumed 1 st order kinetics time 48

An Example of the Half Life I • Half life of Ta. Ka. Ra Taq Polymerase 9/15/2020 Temperature Half life 97. 5℃ 7 min 95℃ 35 min 95℃ (+10% glycerol) > 60 min 95℃ (+5% glycerol) 40 min 92. 5℃ > 130 min 49

An Example of the Half Life II • Pierce Taq at 95℃ = 2. 5 hours (150 min) • Therefore, if we assume the half life of polymerase is 200 min… – The decay rate constant is about 6 10 -5 s-1 9/15/2020 50

Inactivation of Taq Polymerase • Assumption of 1 st order reaction • Assume the half life is … • The calculated rate constant is … • Therefore, the effective enzyme concentration is … 9/15/2020 51

Experimental Results
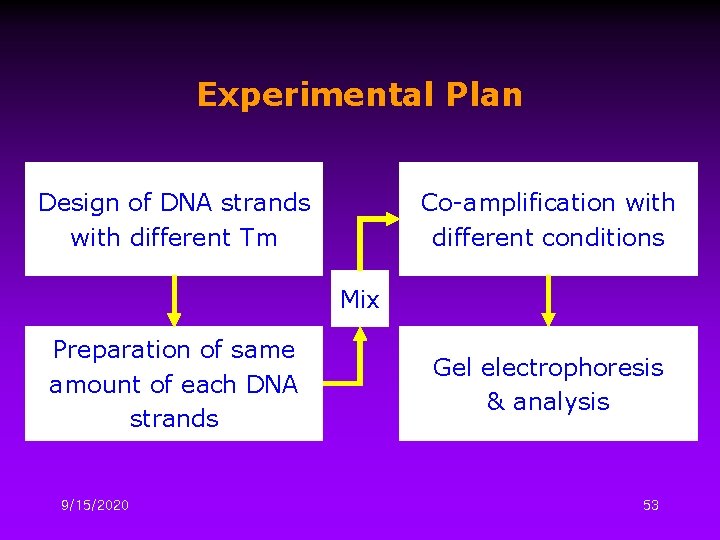
Experimental Plan Design of DNA strands with different Tm Co-amplification with different conditions Mix Preparation of same amount of each DNA strands 9/15/2020 Gel electrophoresis & analysis 53

Design of DNA Strands • Preparation with existing oligomers Length (bp) GC % Tm (calculated by GC method) Tm (measured) 100 47 74. 17 82 A 5’-TCAGCCAACGTACACAGCAAAGGCGAGTATGGGGTATATCTTATGATTCC ACTGGATATCATCGTACTCATGGTCCCTACTCGCGACAGAGTGGATGAAA-3’ 100 B 9/15/2020 72. 53 80 5’-TCAGCCAACGTACACAGCAATTATGATTCCACTGGATATCAGGCGAGTAT GGGGTATATCATGATATGTAGATTCCTCGCGACAGAGTGGATGAAA-3’ 120 C 43 47. 5 75. 21 80. 5 5’-TCAGCCAACGTACACAGCAAAGGCGAGTATGGGGTATATCTTATGATTCCACTGGATATC ATCGTACTCATGGTCCCTACCGCTCCATCCTTGATCGTTTTCGCGACAGAGTGGATGAAA-3’ 54
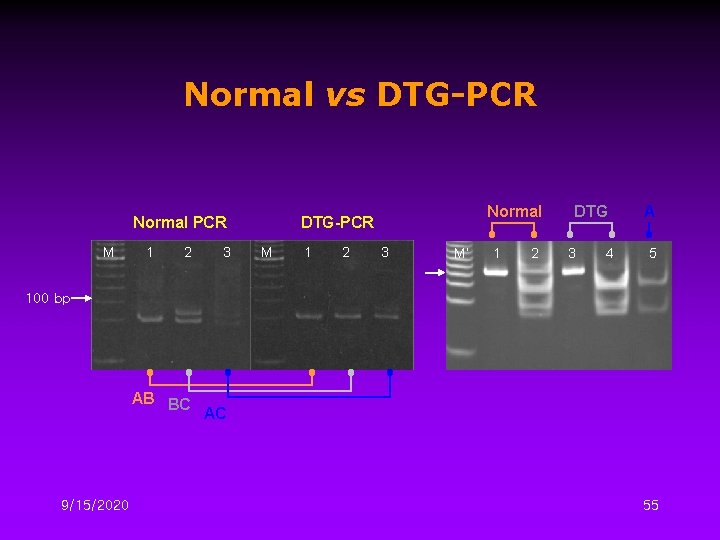
Normal vs DTG-PCR Normal PCR M 1 2 3 Normal DTG-PCR M 1 2 3 M’ 1 2 DTG 3 4 A 5 100 bp AB BC 9/15/2020 AC 55
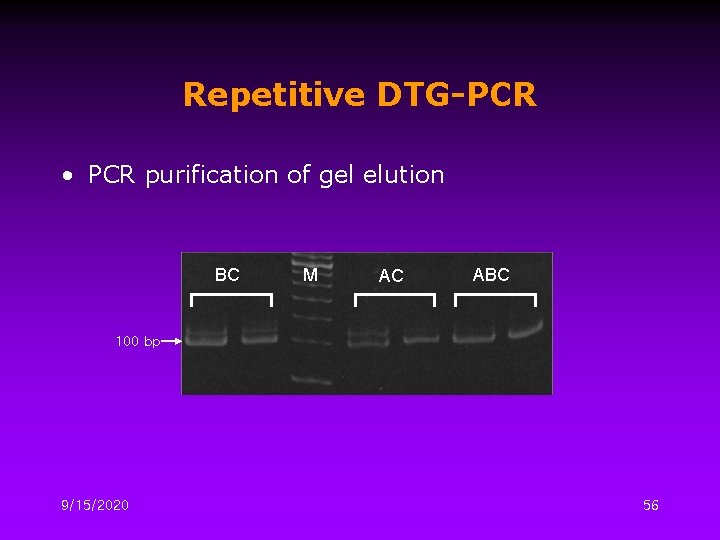
Repetitive DTG-PCR • PCR purification of gel elution BC M AC ABC 100 bp 9/15/2020 56
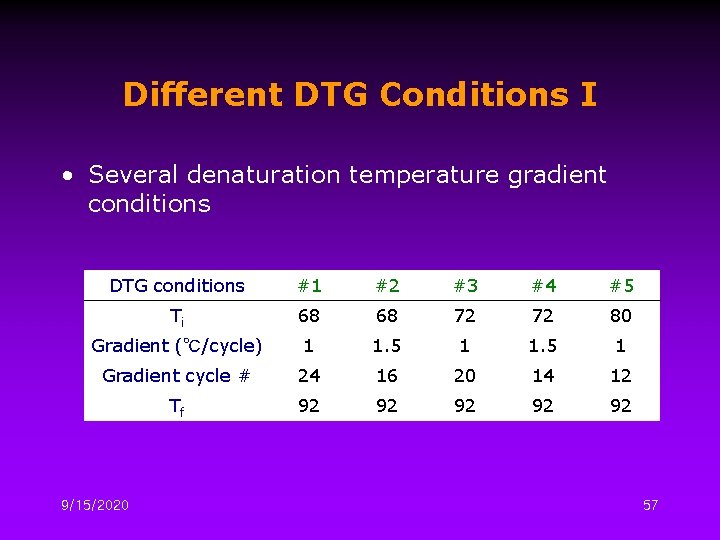
Different DTG Conditions I • Several denaturation temperature gradient conditions DTG conditions #1 #2 #3 #4 #5 Ti 68 68 72 72 80 Gradient (℃/cycle) 1 1. 5 1 Gradient cycle # 24 16 20 14 12 Tf 92 92 92 9/15/2020 57
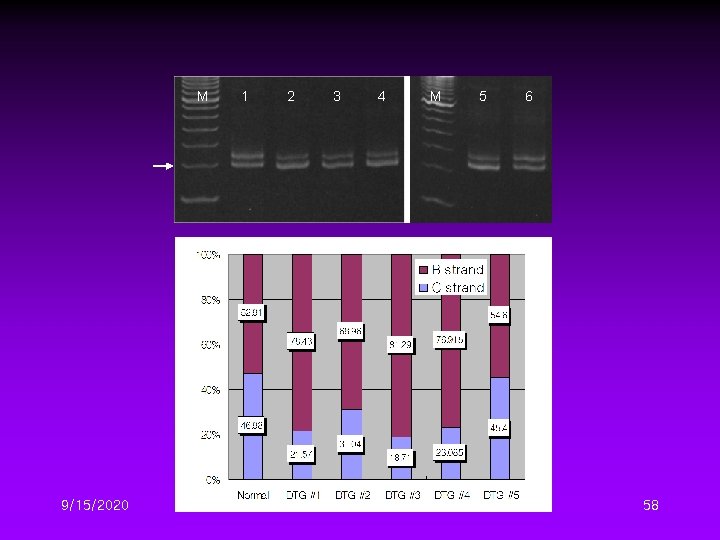
M 9/15/2020 1 2 3 4 M 5 6 58

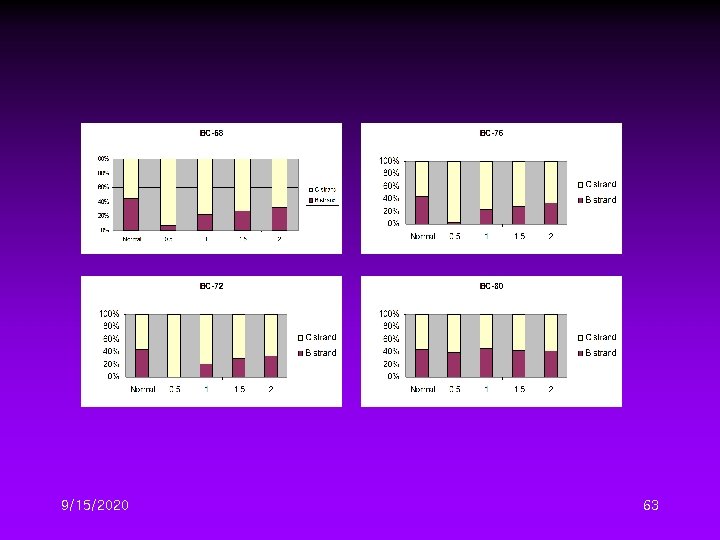
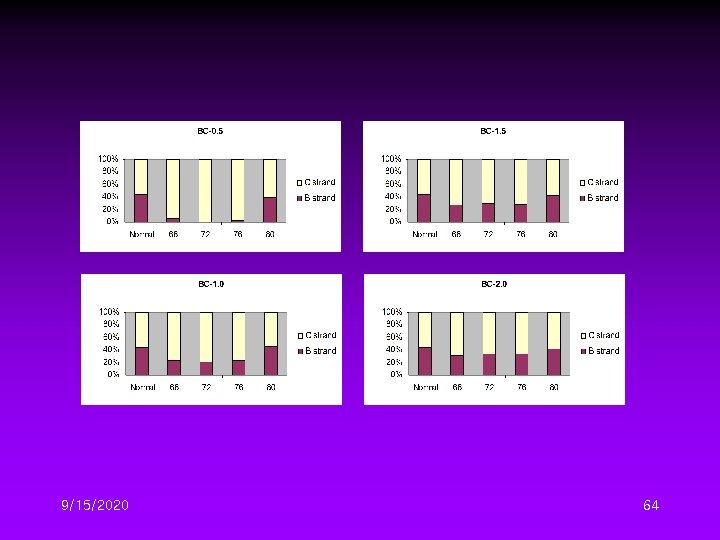
Different DTG Conditions II • Four initial denaturation temperature – 68, 72, 76, 80℃ • Four temperature gradient – 0. 5, 1. 0, 1. 5, 2. 0℃/cycle 9/15/2020 59
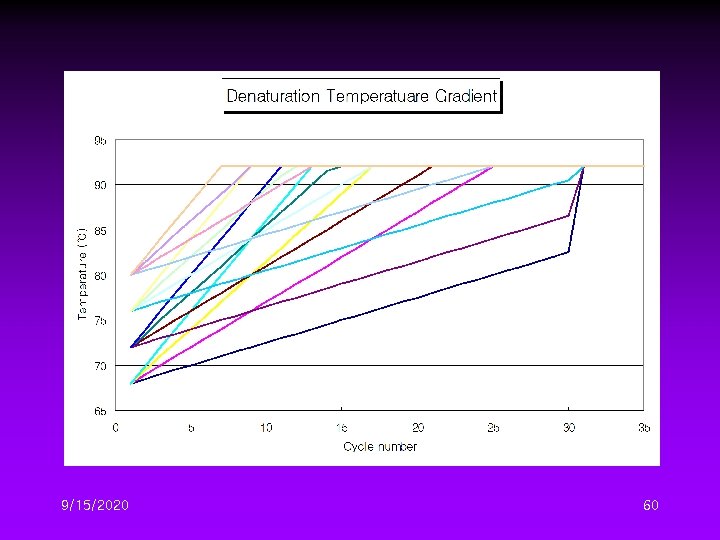
9/15/2020 60
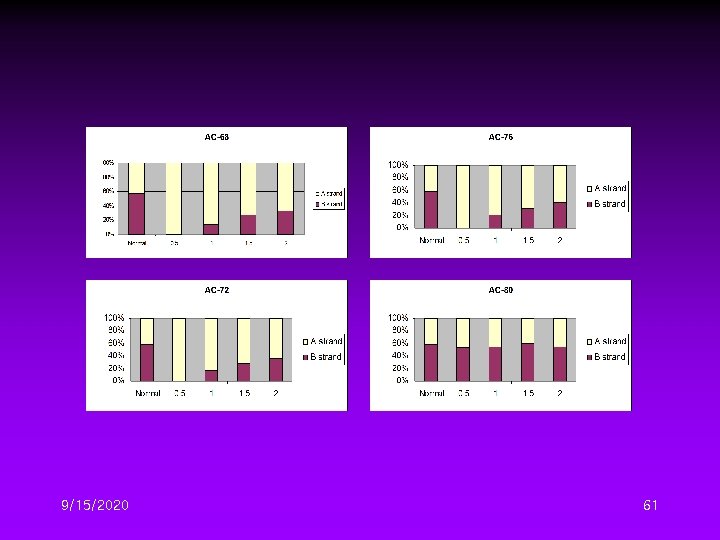
9/15/2020 61
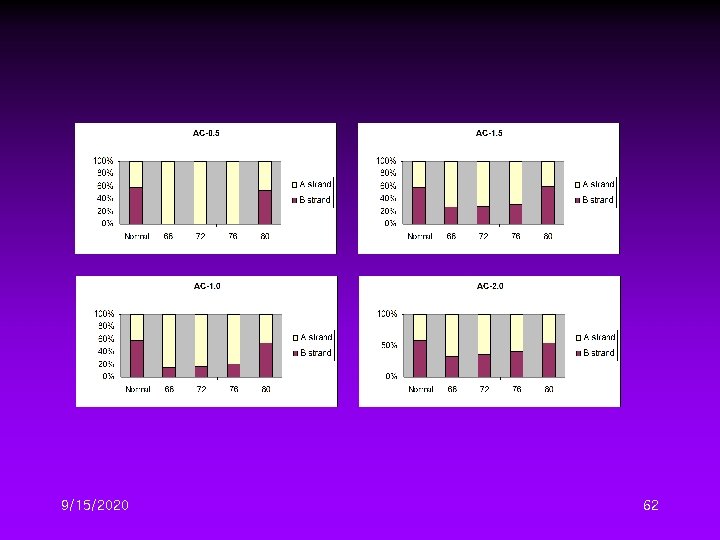
9/15/2020 62
9/15/2020 63
9/15/2020 64

Future Work 9/15/2020 65
- Slides: 65