Effect of Single Nucleotide Polymorphism in Affymetrix probes

Effect of Single Nucleotide Polymorphism in Affymetrix probes Olivia Sanchez-Graillet Departments of Biological Sciences and Mathematical Sciences University of Essex (UK) December 2008
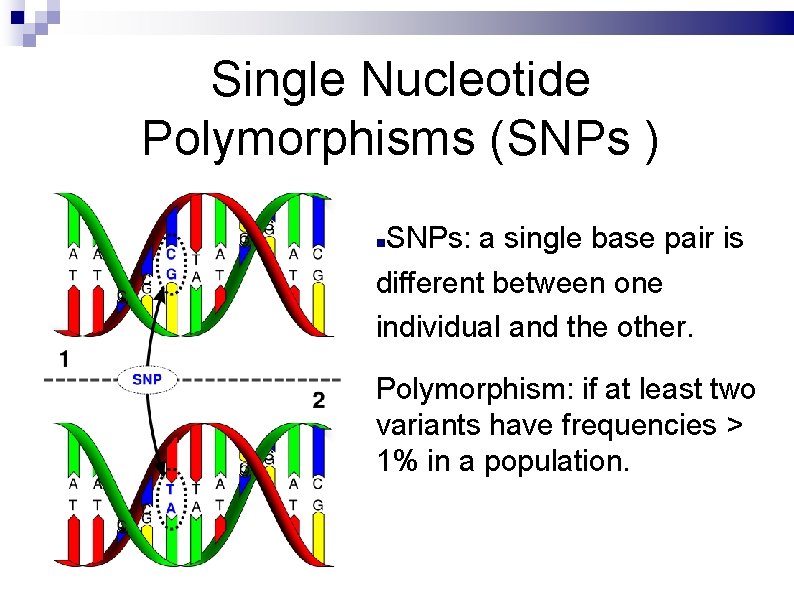
Single Nucleotide Polymorphisms (SNPs ) SNPs: a single base pair is different between one individual and the other. Polymorphism: if at least two variants have frequencies > 1% in a population.

SNPs are the most common type of sequence variation between individuals. SNPs are markers of phenotypes and diseases. SNPs may alter the gene expression and may change or not the amino acid sequence.

Other common variations: DIP: deletion/insertion polymorphism : STR: short tandem repeat (microsatellite) polymorphism (CA)19/20/21/22/23/24/25/26 MIXED: cluster containing submissions from 2 or more alleleic classes -/T , C/- -/AAAAA/AAAACCAAAAAAAA MNP: multiple nucleotide polymorphism with alleles of common length > 1 AAA/CCC

We are studying the relationships between probes intensities on Affymetrix Gene. Chips. Affymetrix Gene chips contain thousands of probes
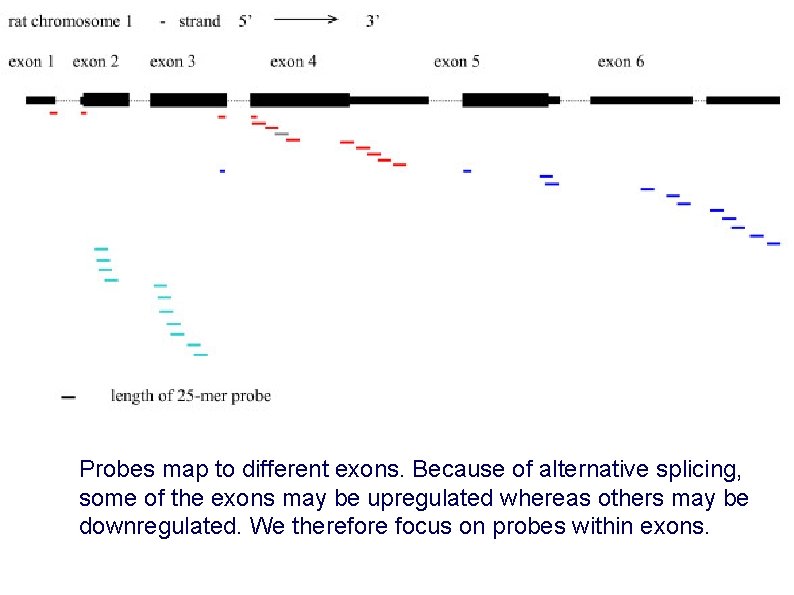
Probes map to different exons. Because of alternative splicing, some of the exons may be upregulated whereas others may be downregulated. We therefore focus on probes within exons.

Probes mapping to the same exon should behave similarly. What causes Affymetrix probes to behave as outliers with respect to other probes within a single exon? Objective: Study the impact of SNPs and other common variation upon Affymetrix probes on Gene. Chips. Explore whether the existence of a SNP causes a probe to behave differently to other probes which map uniquely to a single exon.

Previous research on how SNPs might affect gene expression: Allele A is over-expressed compared to allele B or vs or both alleles are equally expressed (Kumari et al. , 2007). Hybridization resulted from variation might mislead the interpretation of data from individual genes, even if a single probe is affected (Alberts et al. , 2007). In 15 of 25 probesets, SNPs caused a difference in hybridization. Not every SNP causes a difference in hybridization (Alberts et al. , 2007). When the SNPs located at the very beginning or end of a probe, it might have little or not effect on hybridization (Hughes et al. , 2001).

Method: A) Generation of exon heatmaps B) Identification of probes containing SNPs. C) Study of SNP-probes which are outliers.

(A) Generation of exon heatmaps 1. CEL files are downloaded from the GEO database. 2. Calibration of microarray data: 3. Quality control: detection of spatial flaws. Row Quantile Normalisation. Correlate the intensities for groups of probes, using many thousands of Gene. Chip experiments.
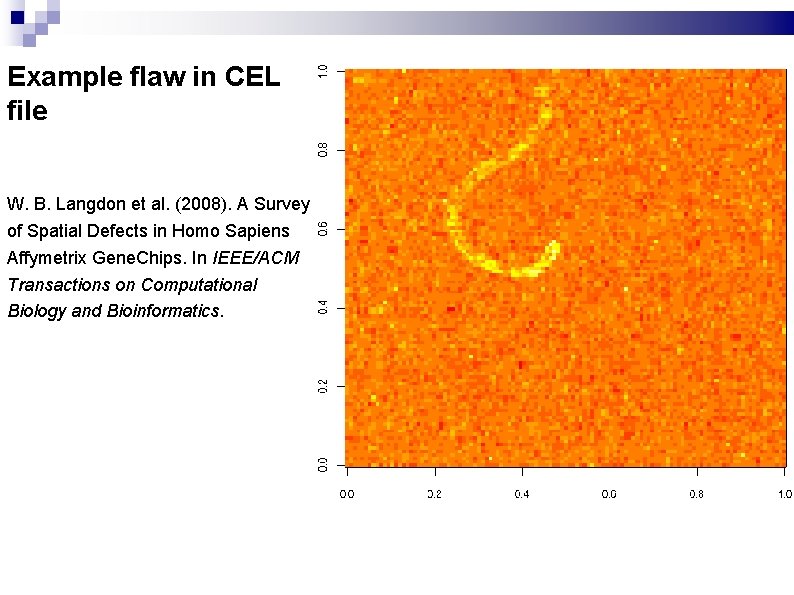
Example flaw in CEL file W. B. Langdon et al. (2008). A Survey of Spatial Defects in Homo Sapiens Affymetrix Gene. Chips. In IEEE/ACM Transactions on Computational Biology and Bioinformatics.
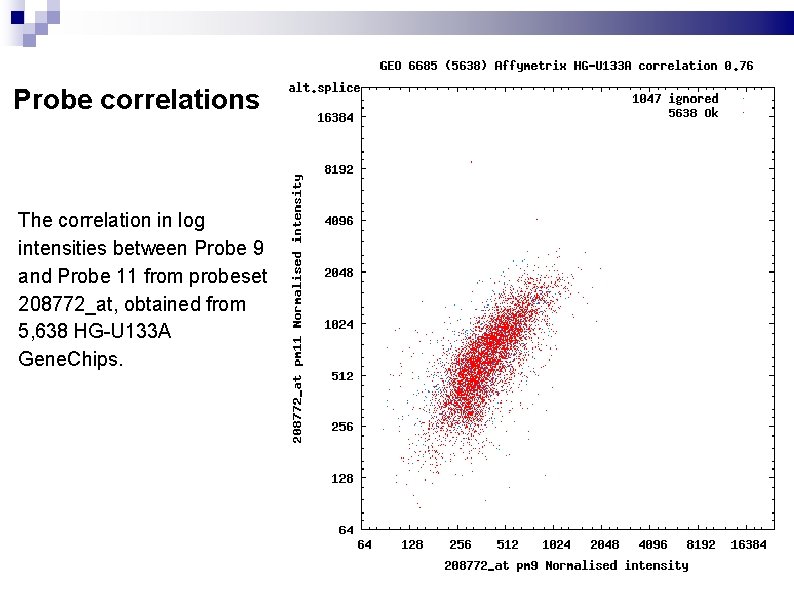
Probe correlations The correlation in log intensities between Probe 9 and Probe 11 from probeset 208772_at, obtained from 5, 638 HG-U 133 A Gene. Chips.
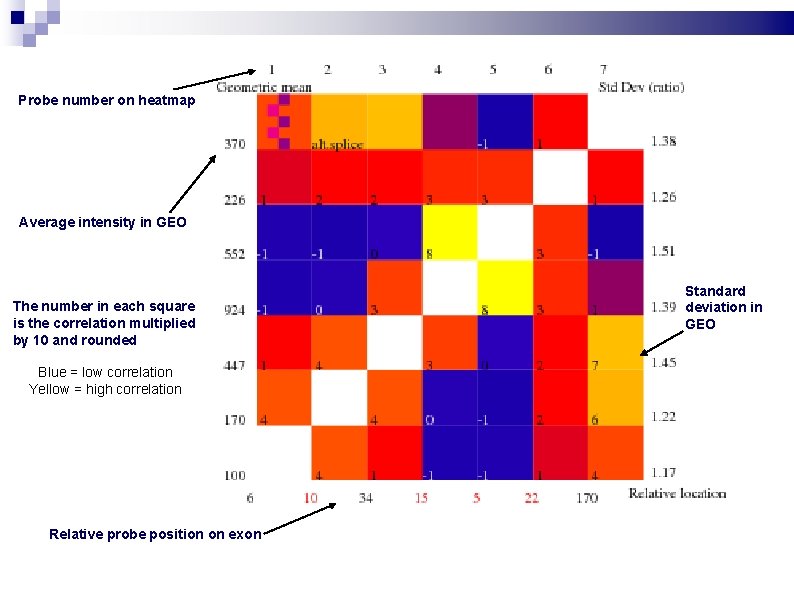
Probe number on heatmap Average intensity in GEO The number in each square is the correlation multiplied by 10 and rounded Blue = low correlation Yellow = high correlation Relative probe position on exon Standard deviation in GEO
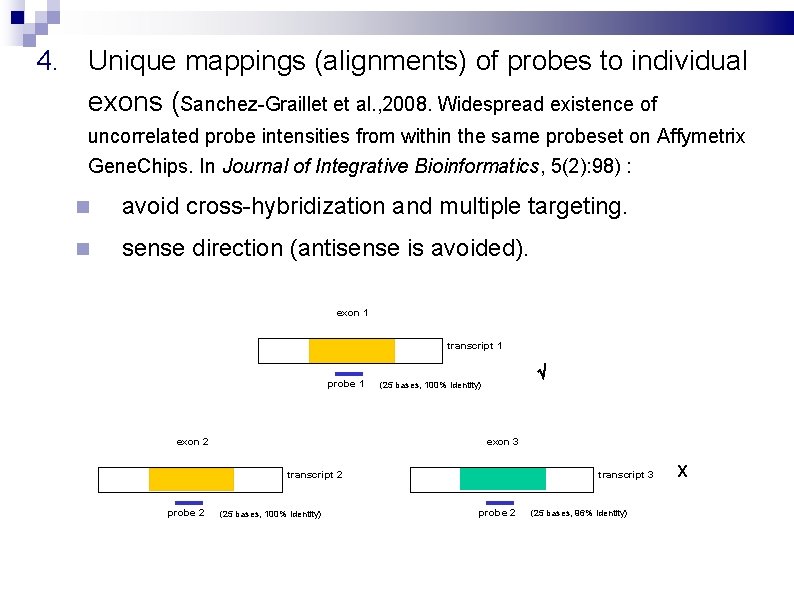
4. Unique mappings (alignments) of probes to individual exons (Sanchez-Graillet et al. , 2008. Widespread existence of uncorrelated probe intensities from within the same probeset on Affymetrix Gene. Chips. In Journal of Integrative Bioinformatics, 5(2): 98) : avoid cross-hybridization and multiple targeting. sense direction (antisense is avoided). exon 1 transcript 1 probe 1 exon 2 (25 bases, 100% identity) exon 3 transcript 2 probe 2 (25 bases, 100% identity) transcript 3 probe 2 (25 bases, 96% identity) X

(B, C) Identification of probes containing SNPs and outlier SNP probes
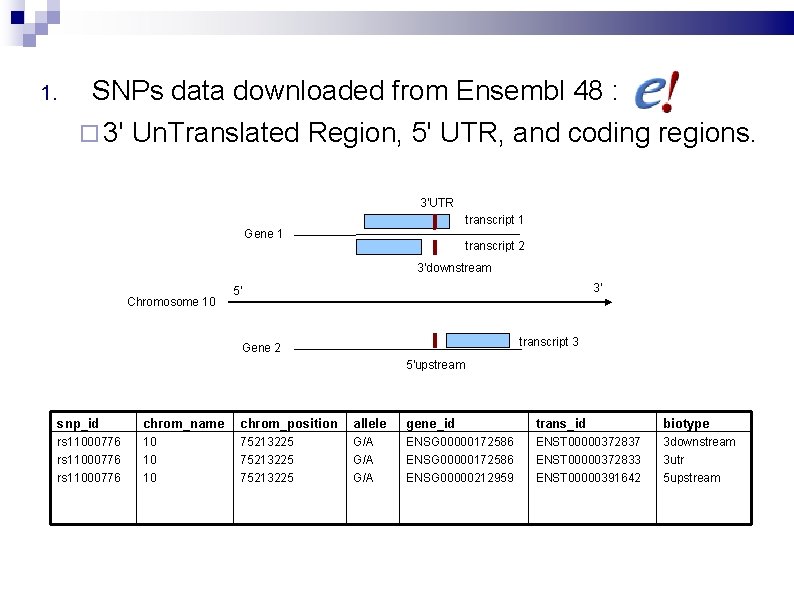
1. SNPs data downloaded from Ensembl 48 : 3' Un. Translated Region, 5' UTR, and coding regions. 3'UTR transcript 1 Gene 1 transcript 2 3'downstream Chromosome 10 3' 5' transcript 3 Gene 2 5'upstream snp_id chrom_name chrom_position allele gene_id trans_id biotype rs 11000776 10 10 10 75213225 G/A G/A ENSG 00000172586 ENSG 00000212959 ENST 00000372837 ENST 00000372833 ENST 00000391642 3 downstream 3 utr 5 upstream

2. Identification of exons with SNPs by using transcript information and chromosomic positions. 3. Selection of unique exons and probes: Only unique exons with more than 4 probes. SNP positions on the probes uniquely mapping to exons are obtained.

4. Identification of SNP-probes which are outliers: The overall correlation matrix median (OMM) is compared with each SNP-probe median (SPM). If OMM – SPM >= 0. 15

OMM SPM_8 SPM_9 0. 87 0. 84 0. 21 Difference 0. 03<0. 15 0. 66>0. 15 SNP in an no-outlier probe SNP in an outlier probe

Results
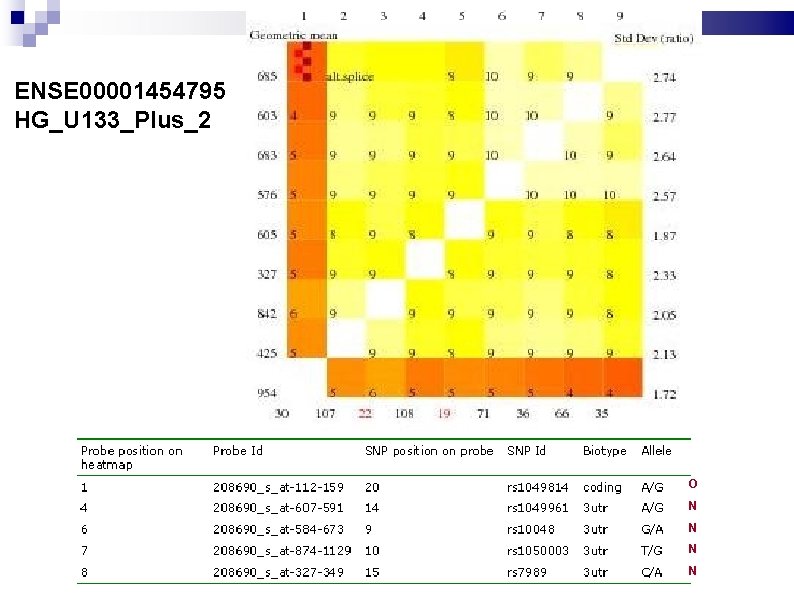
ENSE 00001454795 HG_U 133_Plus_2 O N N
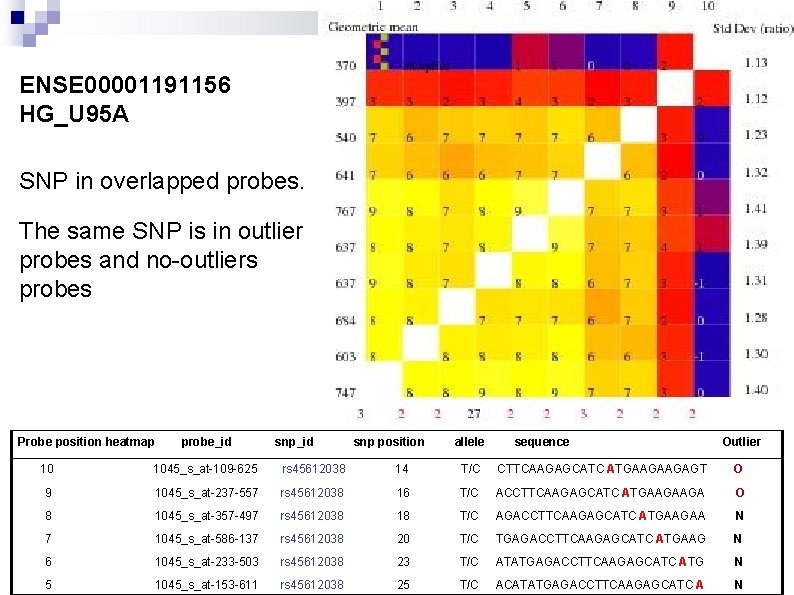
ENSE 00001191156 HG_U 95 A SNP in overlapped probes. The same SNP is in outlier probes and no-outliers probes Probe position heatmap probe_id snp position allele sequence Outlier 10 1045_s_at-109 -625 rs 45612038 14 T/C CTTCAAGAGCATC ATGAAGAAGAGT O 9 1045_s_at-237 -557 rs 45612038 16 T/C ACCTTCAAGAGCATC ATGAAGAAGA O 8 1045_s_at-357 -497 rs 45612038 18 T/C AGACCTTCAAGAGCATC ATGAAGAA N 7 1045_s_at-586 -137 rs 45612038 20 T/C TGAGACCTTCAAGAGCATC ATGAAG N 6 1045_s_at-233 -503 rs 45612038 23 T/C ATATGAGACCTTCAAGAGCATC ATG N 5 1045_s_at-153 -611 rs 45612038 25 T/C ACATATGAGACCTTCAAGAGCATC A N

ENSE 0000129003 HG_U 133 A SNPs in only no-outlier probes snp_id probe_position_heatmap snp_position_probe allele seq rs 11038 221667_s_at-512 -441 10 13 A/G GTTTATGATCTGACCTAGGTCCCCC N rs 6413487 221667_s_at-570 -641 9 7 C/G TAAGGACGCTGGGAGCCTGTCAGTT N
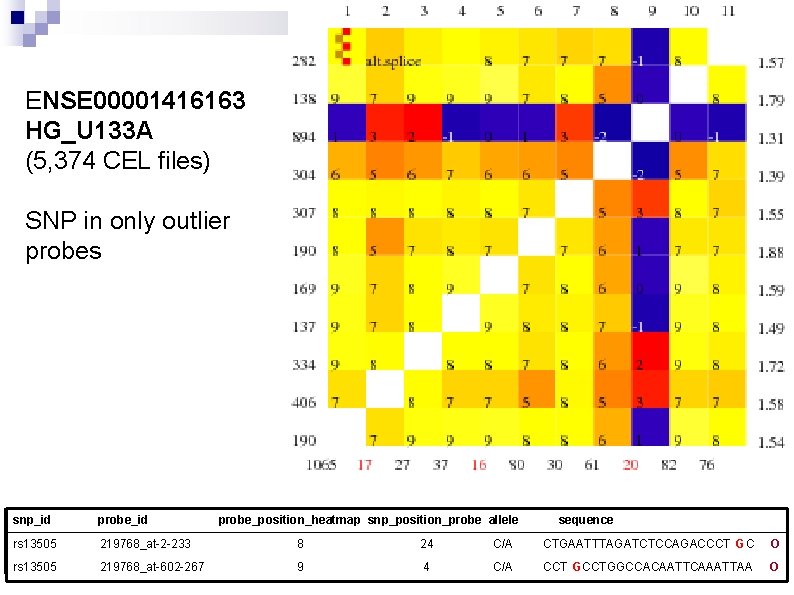
ENSE 00001416163 HG_U 133 A (5, 374 CEL files) SNP in only outlier probes snp_id probe_position_heatmap snp_position_probe allele sequence rs 13505 219768_at-2 -233 8 24 C/A CTGAATTTAGATCTCCAGACCCT GC O rs 13505 219768_at-602 -267 9 4 C/A CCT GCCTGGCCACAATTCAAATTAA O
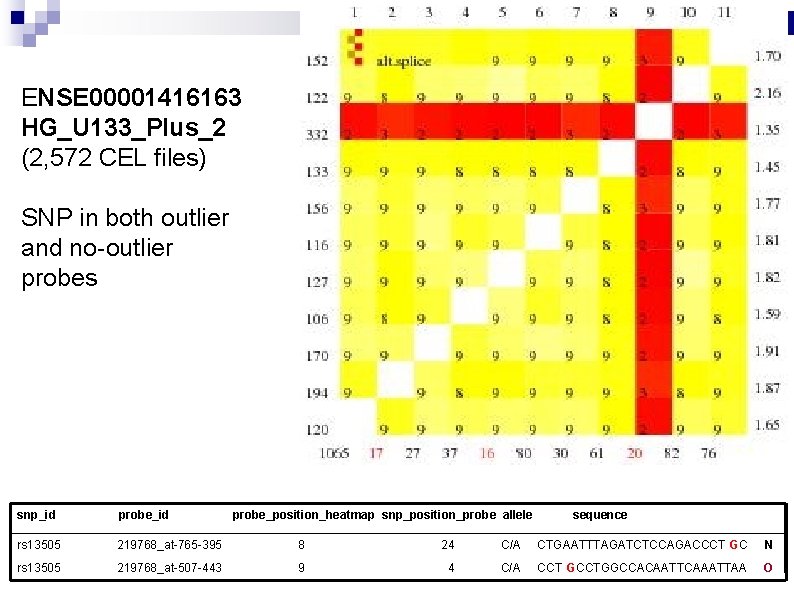
ENSE 00001416163 HG_U 133_Plus_2 (2, 572 CEL files) SNP in both outlier and no-outlier probes snp_id probe_position_heatmap snp_position_probe allele sequence rs 13505 219768_at-765 -395 8 24 C/A CTGAATTTAGATCTCCAGACCCT GC N rs 13505 219768_at-507 -443 9 4 C/A CCT GCCTGGCCACAATTCAAATTAA O

ENSE 00001416163 HG_U 133 A_2 (159 CEL files) SNP in only NO-outlier probes snp_id probe_position_heatmap snp_position_probe allele sequence rs 13505 219768_at-432 -225 8 24 C/A CTGAATTTAGATCTCCAGACCCT GC N rs 13505 219768_at-534 -259 4 C/A CCT GCCTGGCCACAATTCAAATTAA N

~60, 000 SNPs distributed in unique exons of ten array designs. 11% in unique exons in which all probes that contain the same SNP are outliers. 5% in which not all the probes containing the same SNP are outliers. 84% in which all probes are not outliers. These numbers may vary according to the Ensembl version used and the threshold for outliers chosen.
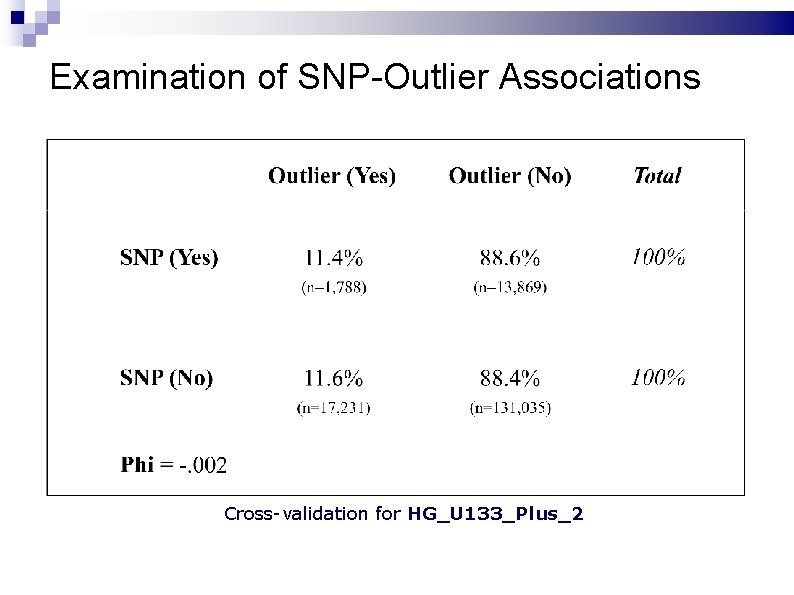
Examination of SNP-Outlier Associations Cross-validation for HG_U 133_Plus_2
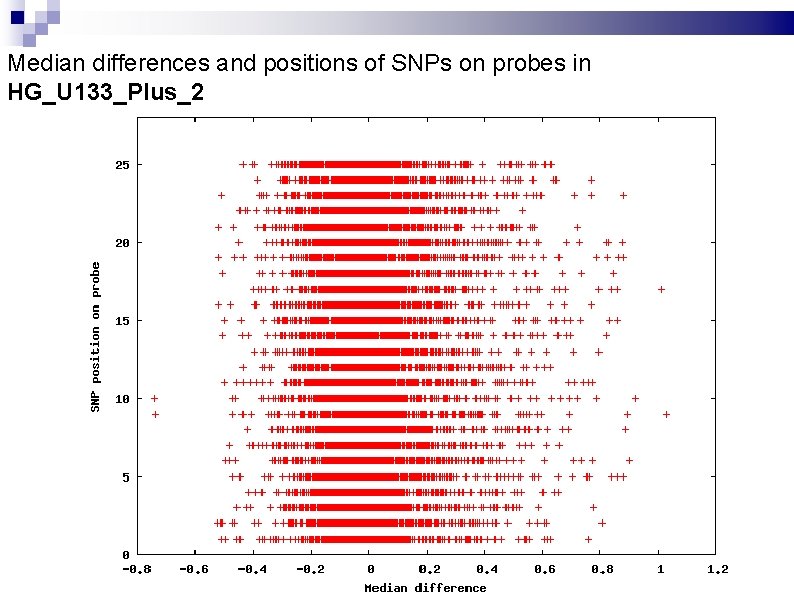
Median differences and positions of SNPs on probes in HG_U 133_Plus_2
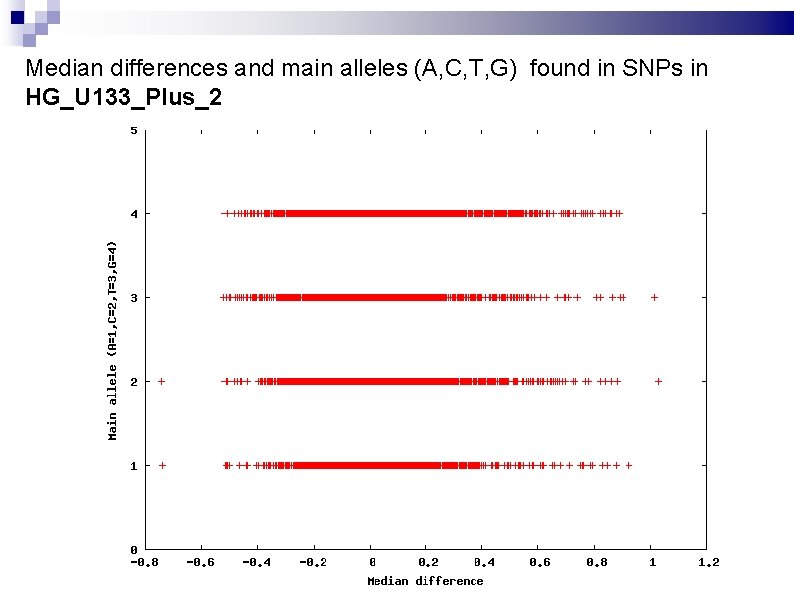
Median differences and main alleles (A, C, T, G) found in SNPs in HG_U 133_Plus_2

We have identified other causes of outlier probes: Probes containing a contiguous run of 4 or more guanines: formation of G-quadruplexes occurring on the surface of a Gene. Chip. (Upton et al. , BMC Genomics (in press)). Probes located next to bright probes, such as at the edge of the Genechip, are affected by blur. Motifs or any other “problematic” subsequences.
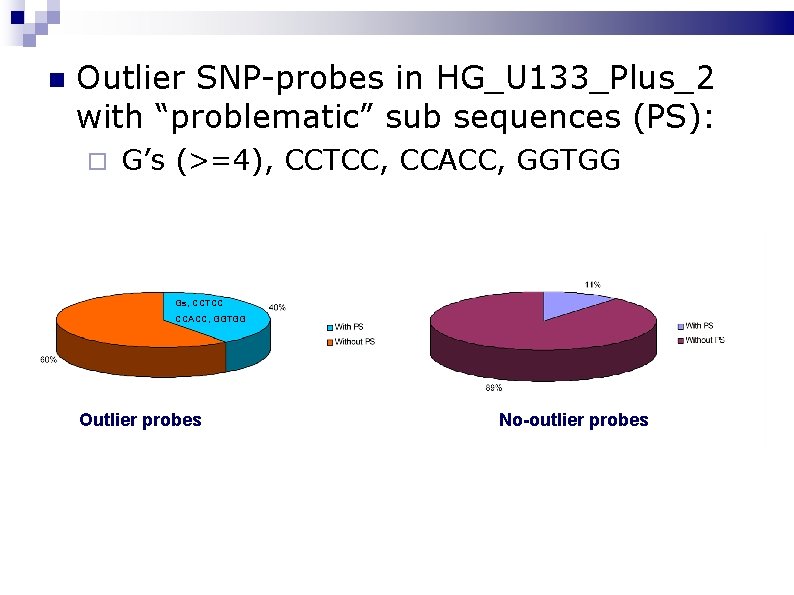
Outlier SNP-probes in HG_U 133_Plus_2 with “problematic” sub sequences (PS): G’s (>=4), CCTCC, CCACC, GGTGG Gs, CCTCC CCACC, GGTGG Outlier probes No-outlier probes

Conclusions

We have not found a common behaviour when SNPs are present in a probe. SNPs do not seem to cause outliers in groups of probes representing individual exons. SNPs may influence other biological events like alternative poly(A). The genomic region where SNPs are found, the position of the SNP in a probe, the main allele, and the number of SNPs in a probe does not make a probe an outlier in the correlation heatmap.

Bioinformatics Group Dr Andrew Harrison Dr Berthold Lausen Dr Abdel Salhi Professor Graham Upton Physics Statistics Mathematics Statistics Dr William Langdon Dr Olivia Sanchez Dr Maria Stalteri Physics and Computer Sc. Inorganic Chemistry & Bioinformatics Jose Arteaga-Salas Rohmatul Fajriyah Abdelhak Kheniche Rahim Bux Khokhar Zain-Ul-Abdin Khurho Farhat Memon Joanna Rowsell Statistics Pharmacology & Mathematics Computer Sc. Mathematics

Thank you!
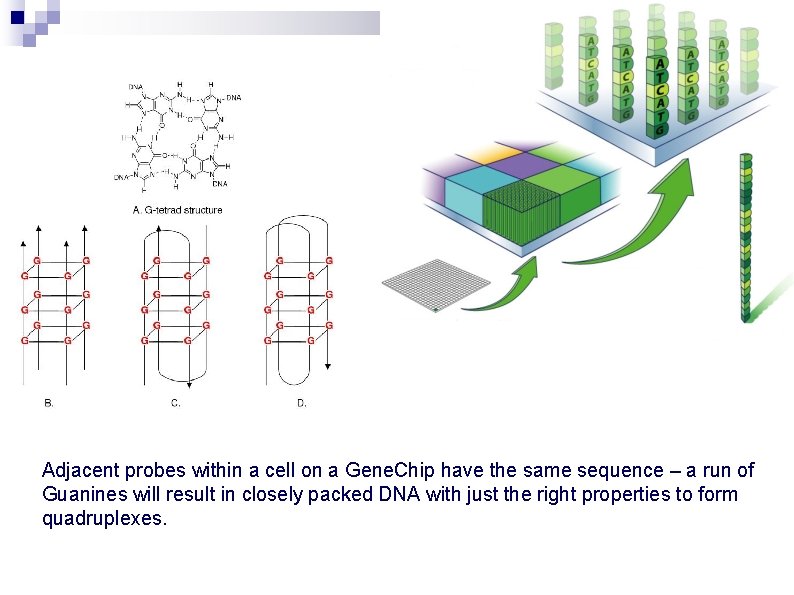
Adjacent probes within a cell on a Gene. Chip have the same sequence – a run of Guanines will result in closely packed DNA with just the right properties to form quadruplexes.
- Slides: 37