Effect of Plasmid Concentration on Bacterial Transformation Emmanuel

Effect of Plasmid Concentration on Bacterial Transformation Emmanuel J. Eppinger Campus School of Carlow University 8 th Grade
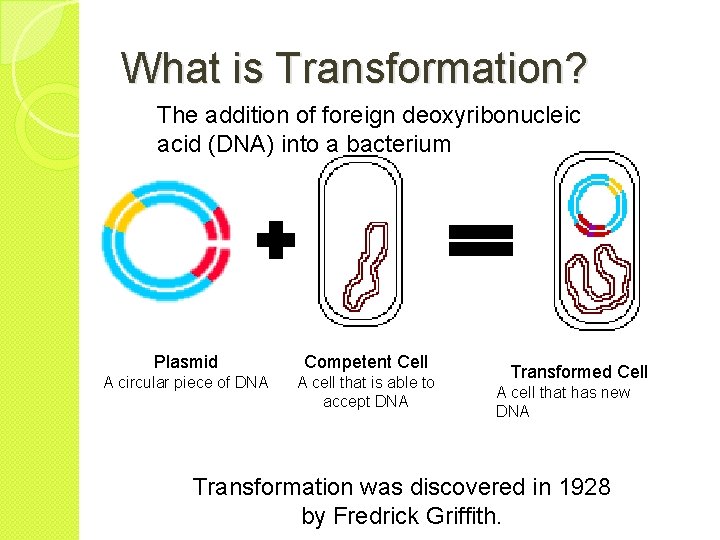
What is Transformation? The addition of foreign deoxyribonucleic acid (DNA) into a bacterium Plasmid Competent Cell A circular piece of DNA A cell that is able to accept DNA Transformed Cell A cell that has new DNA Transformation was discovered in 1928 by Fredrick Griffith.
Uses of Bacterial Transformation To mass produce proteins e. g. DRACO protein gives immunity to all viruses (e. g. , cold, flu, Ebola) To transfer traits among species e. g. Transfer of luminescence genes can be used to create secret codes To mass produce DNA is multiplied each time a cell reproduces
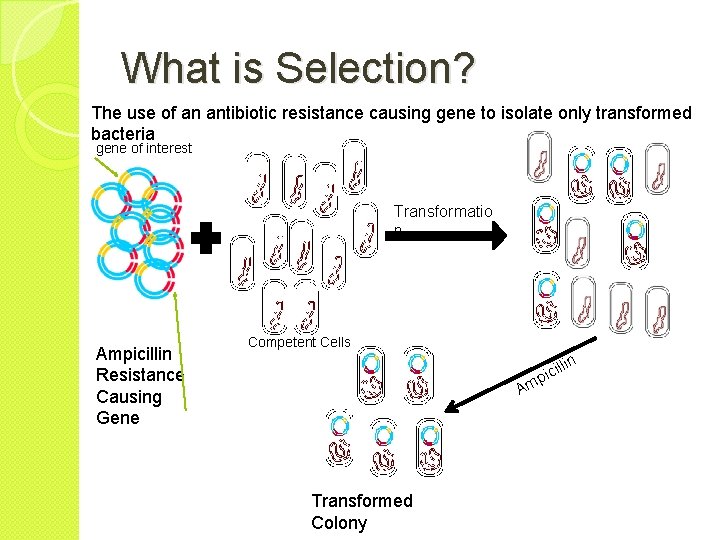
What is Selection? The use of an antibiotic resistance causing gene to isolate only transformed bacteria gene of interest Transformatio n Ampicillin Resistance Causing Gene Competent Cells in ill c i p Am Transformed Colony
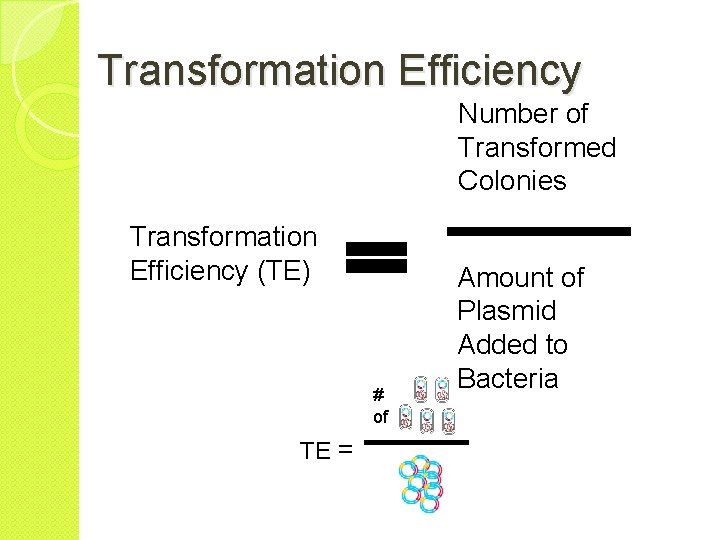
Transformation Efficiency Number of Transformed Colonies Transformation Efficiency (TE) # of TE = Amount of Plasmid Added to Bacteria

Purpose �During transformation, not all bacteria are transformed. ◦ Only some bacteria accept the plasmid. �The purpose of this experiment is to determine whether the concentration of plasmid will affect the efficiency with which bacteria transform

Hypothesis If the rate at which bacteria colonies transform is not proportional to the plasmid concentration, then the transformation efficiencies of different concentrations of plasmid will not be equal because transformation efficiency is a function of plasmid concentration.

Materials � Experiment-Specific Supplies � Disposable Lab Supplies ◦ 1 m. L of competent DH 5 alpha E. ◦ Goggles, gloves, ice, sterile Coli cells distilled water ◦ 15 µL of Puc 18 ampicillin ◦ Sterile micro pipette tips resistance causing plasmid at. 82 ◦ Sterile 1. 5 m. L micro tubes µg of plasmid per µL of solution ◦ 6 m. L of sterile LB nutrient agar � Lab Equipment ◦ 23 ampicillin positive agar plates ◦ Timer ◦ 1 ampicillin negative agar plates ◦ Closed test tube rack ◦ 37˚C Incubator ◦ Sterile spreader bars ◦ Turn table ◦ Labeling utensil ◦ Pipetter

Procedure Step 1: Prepare Plasmid o Wear goggles and gloves Keep plasmid on ice for entire dilution 1. Create various concentrations of plasmid by diluting with water according to table o 2. Concentration Plasmid (µL) Water (µL) 1 x 6 0 . 5 x 3 3 . 1 x 1 9 . 05 x 1 19 . 01 x 1 99 . 001 x 1 999 . 0001 x 1 9999 For each concentration, add 2µL of diluted plasmid to a 1. 5 m. L tube labeled with the concentration

Procedure Step 2: Transform Bacteria 3. 4. 5. 6. 7. 8. Add 100 µL of the competent cells to each of the labeled 1. 5 m. L tubes Re-suspend the cells and plasmid by drawing in 30 µL of the solution and adding it back repeatedly for 5 to 10 seconds Keep the tubes on ice for 40 minutes Add 450 µL of LB nutrient agar solution to each tube Place the tubes in pre-heated 37˚C water in a 37˚C incubator for 5 minutes to allow plasmids to be drawn into cells Place the tubes back on ice

Procedure Step 3: Plate Bacteria 9. Re-suspend all the solutions in each of the tubes For each concentration, add 100 µL of the solution to each of 3 ampicillin positive plates 11. Spread 100 µL of cells on both an ampicillin positive and negative plate (controls) 12. Spread the liquid using a spreader bar and a turn table 13. Invert the plates and incubate at 37˚C for 48 hours 10. Procedure Step 4: Analyze Results 14. Count and record the number of colonies on each plate 15. Compute the transformation efficiencies for each plate 16. Run an ANOVA test on the transformation

Variables Dependent Variable Bacterial Transformation Efficiency: Number of colonies per plate/amount of DNA plated Independent Variable Concentration of Plasmid: 1 x, . 5 x, . 1 x, . 05 x, . 01 x, . 001 x, and. 0001 x Controls Positive Control: Untransformed bacteria on ampicillin negative plate Negative Control: Untransformed bacteria on ampicillin positive plate
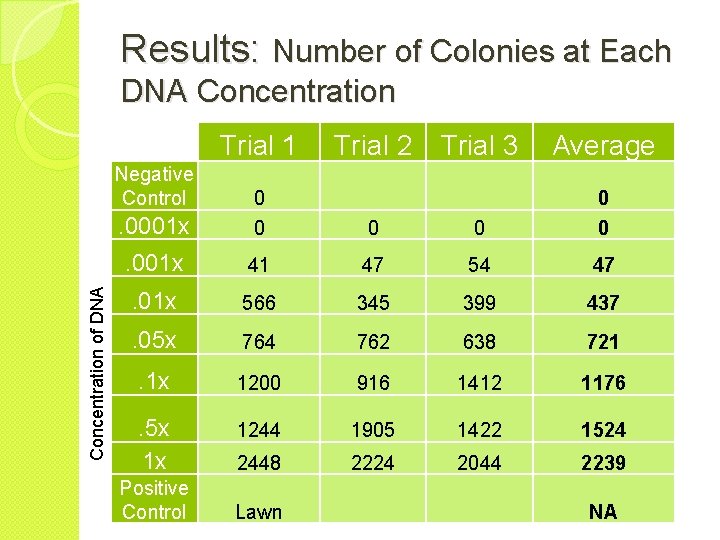
Results: Number of Colonies at Each DNA Concentration Trial 1 Concentration of DNA Negative Control Trial 2 Trial 3 Average . 0001 x 0 0 0 . 001 x 41 47 54 47 . 01 x 566 345 399 437 . 05 x 764 762 638 721 . 1 x 1200 916 1412 1176 . 5 x 1 x 1244 1905 1422 1524 2448 2224 2044 2239 Positive Control Lawn NA
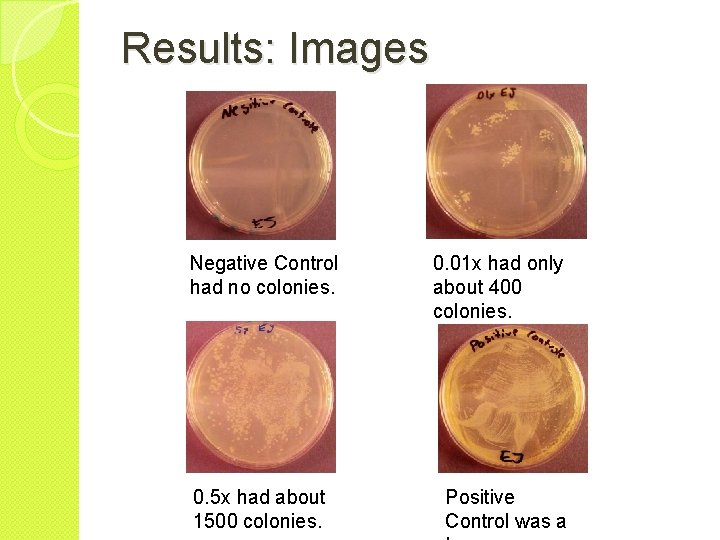
Results: Images Negative Control had no colonies. 0. 5 x had about 1500 colonies. 0. 01 x had only about 400 colonies. Positive Control was a
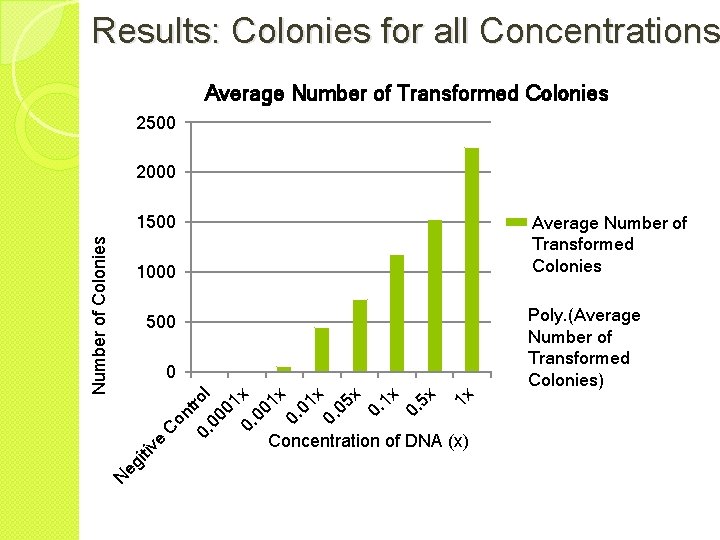
Results: Colonies for all Concentrations Average Number of Transformed Colonies 2500 2000 Average Number of Transformed Colonies 1000 Poly. (Average Number of Transformed Colonies) 500 1 x ro 00 l 01 0. x 00 1 x 0. 01 x 0. 05 x 0. 1 x 0. 5 x 0 N eg iti 0. ve C on t Number of Colonies 1500 Concentration of DNA (x)
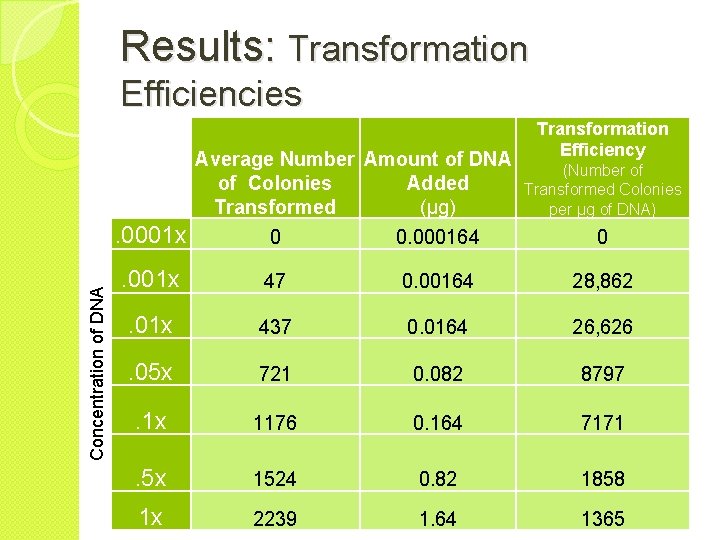
Results: Transformation Efficiencies Concentration of DNA Average Number Amount of DNA of Colonies Added Transformed (µg) Transformation Efficiency (Number of Transformed Colonies per µg of DNA) . 0001 x 0 0. 000164 0 . 001 x 47 0. 00164 28, 862 . 01 x 437 0. 0164 26, 626 . 05 x 721 0. 082 8797 . 1 x 1176 0. 164 7171 . 5 x 1524 0. 82 1858 1 x 2239 1. 64 1365
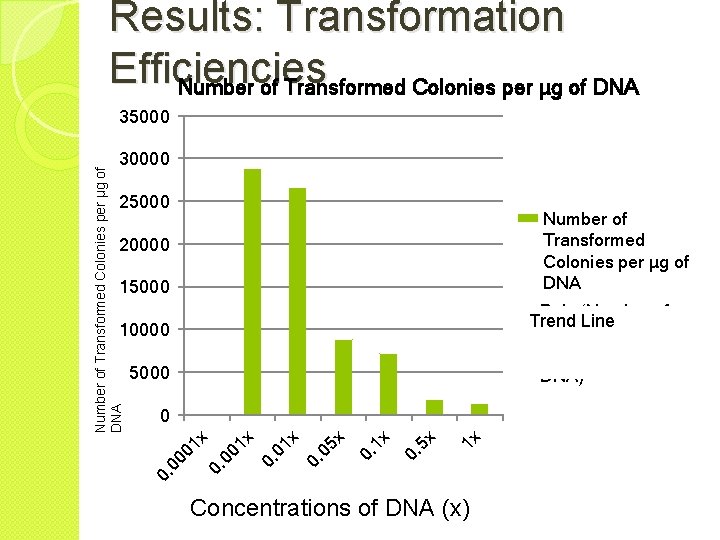
Results: Transformation Efficiencies Number of Transformed Colonies per µg of DNA 35000 Number of Transformed Colonies per µg of DNA 30000 25000 Number of Transformed Colonies per µg of DNA 20000 15000 Poly. (Number of Trend Line Transformed Colonies per µg of DNA) 10000 5000 1 x x 0. 5 x 0. 1 x 0. 05 x 0. 01 01 x 0. 0 0. 00 0 1 x 0 Concentrations of DNA (x)
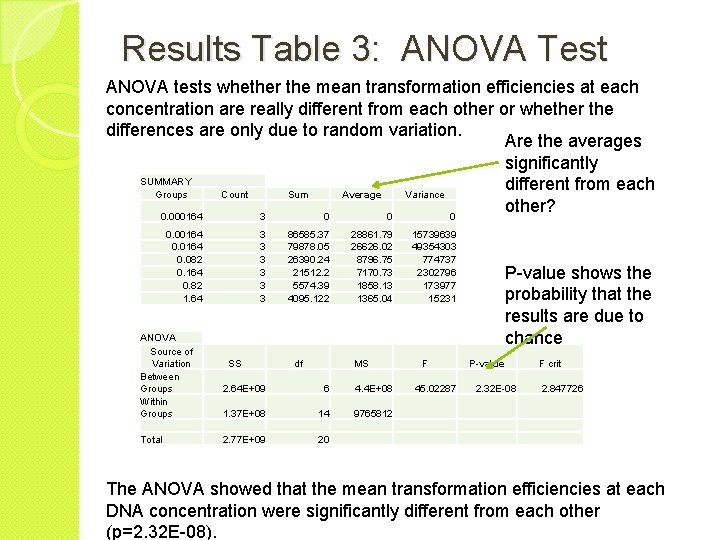
Results Table 3: ANOVA Test ANOVA tests whether the mean transformation efficiencies at each concentration are really different from each other or whether the differences are only due to random variation. Are the averages significantly SUMMARY different from each Groups Count Sum Average Variance other? 0. 000164 3 0 0. 00164 0. 082 0. 164 0. 82 1. 64 ANOVA Source of Variation Between Groups Within Groups Total 3 3 3 SS 86585. 37 79878. 05 26390. 24 21512. 2 5574. 39 4095. 122 df 28861. 79 26626. 02 8796. 75 7170. 73 1858. 13 1365. 04 15739639 49354303 774737 2302796 173977 15231 MS 2. 64 E+09 6 4. 4 E+08 1. 37 E+08 14 9765812 2. 77 E+09 20 P-value shows the probability that the results are due to chance F P-value 45. 02287 F crit 2. 32 E-08 2. 847726 The ANOVA showed that the mean transformation efficiencies at each DNA concentration were significantly different from each other (p=2. 32 E-08).

Conclusion � The greater the DNA concentration, the greater the average number of transformed colonies. � The transformation efficiencies were the opposite of the number of colonies: the lower the concentration of DNA, the higher the transformation efficiency. ◦ The exception was 0. 0001 x which had 0 colonies per µg of DNA added. � This suggests that ◦ Using lower concentrations of DNA is more efficient but that a point exists where so little plasmid is added that there is not enough to support a transformed colony. ◦ This point is between 0. 000164 µg of DNA (0. 0001 x)

Therefore… � The original hypothesis, “If the rate at which bacteria colonies transform is not proportional to the plasmid concentration, then the transformation efficiencies of different concentrations of plasmid will not be equal because transformation efficiency is a function of plasmid concentration, ” was supported. ◦ The number of colonies for each concentration was not proportional to the concentration of the plasmid. ◦ Since they were not proportional, the transformation efficiencies were different and not equal. ◦ Statistical testing from the ANOVA supported

Changes to Improve Experiment �Incubate for less time to prevent the formation of satellite colonies or multiple colonies merging �Test with larger samples of bacteria and DNA �Use an automated colony counter �Measure the surface area covered by colonies rather than the number of colonies

Future Research �Improving Transformation Efficiency ◦ Test lower and higher concentrations of plasmid ◦ Test different types and sizes of plasmids ◦ Test different bacteria species �e. g. , HB 101 ◦ Vary conditions of experiment �e. g. specifications of heat shock �Applications to other species ◦ Test transformation in plants, yeast, and animal cells ◦ Investigate potential uses in gene therapy in

Works Cited Maczulak, Anne. Allies and Enemies: How the World Depends on Bacteria. Upper Saddle River: Pearson Education, Inc. , 2011. Plattsburg. edu. 10 November. < http: //faculty. plattsburgh. edu/donald. slish/tr ansformation. html. > Science. Buddies. org. 10 November. < http: //www. sciencebuddies. org/science -fair-projects/project_ideas/Bio. Chem_p 013. shtml. > Wikipedia. org. 12 October. <http: //en. wikipedia. org/wiki/Transformato n_(genetics). > Wikipedia. org. 5 November 2012. <http: //en. wikipedia. org/wiki/DNA. >

Acknowledgements Mrs. Wojociechowski Middle School Science Teacher, Campus School of Carlow University Mr. Krotec High School Biology Teacher, Central Catholic High School
- Slides: 24