Dynamic Causal Modelling DCM Theory Demis Hassabis Hanneke

Dynamic Causal Modelling (DCM): Theory Demis Hassabis & Hanneke den Ouden Thanks to Klaas Enno Stephan Functional Imaging Lab Wellcome Dept. of Imaging Neuroscience Institute of Neurology University College London

Overview • Classical approaches to functional & effective connectivity • Generic concepts of system analysis • DCM for f. MRI: – Neural dynamics and hemodynamics – Bayesian parameter estimation • Interpretation of parameters – Statistical inference – Bayesian model selection
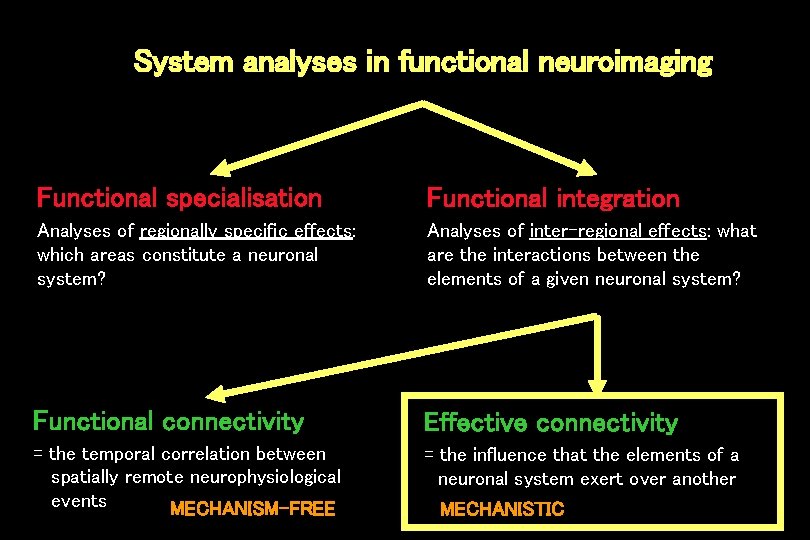
System analyses in functional neuroimaging Functional specialisation Functional integration Analyses of regionally specific effects: which areas constitute a neuronal system? Analyses of inter-regional effects: what are the interactions between the elements of a given neuronal system? Functional connectivity Effective connectivity = the temporal correlation between spatially remote neurophysiological events MECHANISM-FREE = the influence that the elements of a neuronal system exert over another MECHANISTIC

Models of effective connectivity • Structural Equation Modelling (SEM) • Psycho-physiological interactions (PPI) • Multivariate autoregressive models (MAR) & Granger causality techniques • Kalman filtering • Volterra series • Dynamic Causal Modelling (DCM) Friston et al. , Neuro. Image 2003

Overview • Classical approaches to functional & effective connectivity • Generic concepts of system analysis • DCM for f. MRI: – Neural dynamics and hemodynamics – Bayesian parameter estimation • Interpretation of parameters – Statistical inference – Bayesian model selection

Models of effective connectivity = system models. But what precisely is a system? • System = set of elements which interact in a spatially and temporally specific fashion. • System dynamics = change of state vector in time overall system state represented by state variables • Causal effects in the system: – interactions between elements – external inputs u • System parameters : specify the nature of the interactions • general state equation for nonautonomous systems change of state vector in time
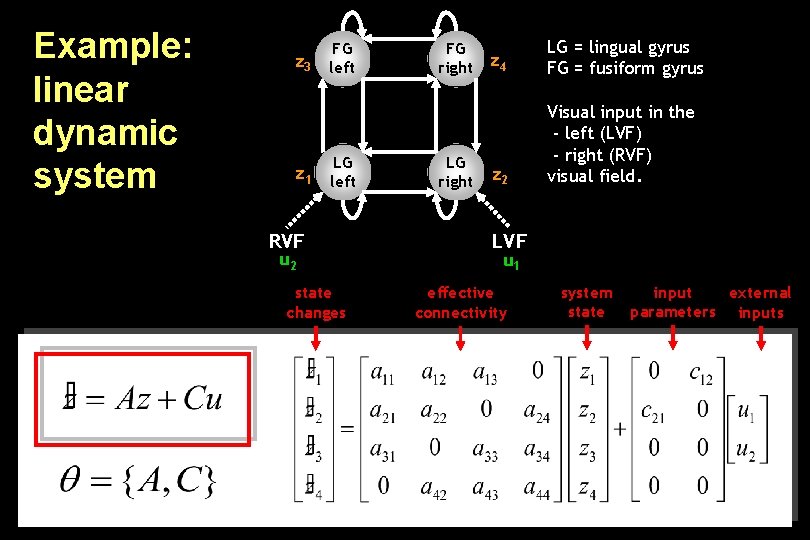
Example: linear dynamic system FG z 3 left z 1 LG left RVF u 2 state changes FG right LG right z 4 LG = lingual gyrus FG = fusiform gyrus z 2 Visual input in the - left (LVF) - right (RVF) visual field. LVF u 1 effective connectivity system state input external parameters inputs
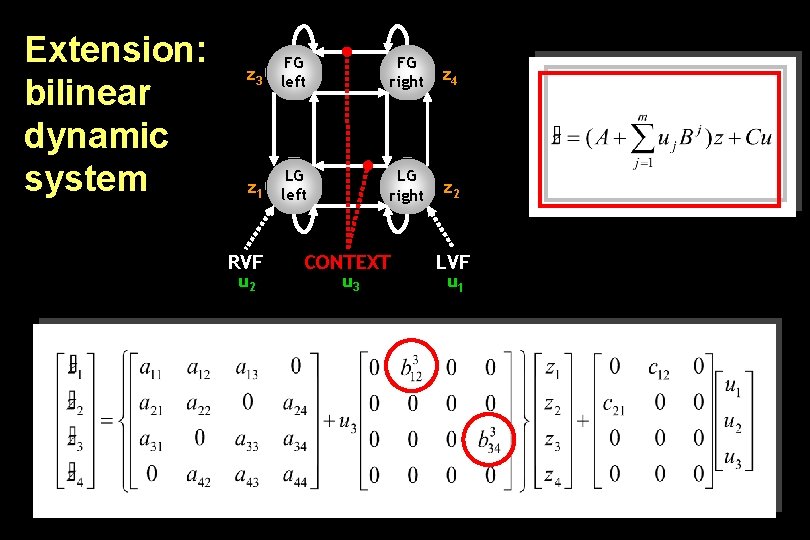
Extension: bilinear dynamic system z 3 FG left FG right z 4 z 1 LG left LG right z 2 RVF u 2 CONTEXT u 3 LVF u 1

Bilinear state equation in DCM state changes intrinsic connectivity modulation of system connectivity state direct inputs m external inputs

Overview • Classical approaches to functional & effective connectivity • Generic concepts of system analysis • DCM for f. MRI: – Neural dynamics and hemodynamics – Bayesian parameter estimation • Interpretation of parameters – Statistical inference – Bayesian model selection

DCM for f. MRI: the basic idea • Using a bilinear state equation, a cognitive system is modelled at its underlying neuronal level (which is not directly accessible for f. MRI). • The modelled neuronal dynamics (z) is transformed into area-specific BOLD signals (y) by a hemodynamic forward model (λ). The aim of DCM is to estimate parameters at the neuronal level such that the modelled BOLD signals are maximally similar to the experimentally measured BOLD signals. z λ y
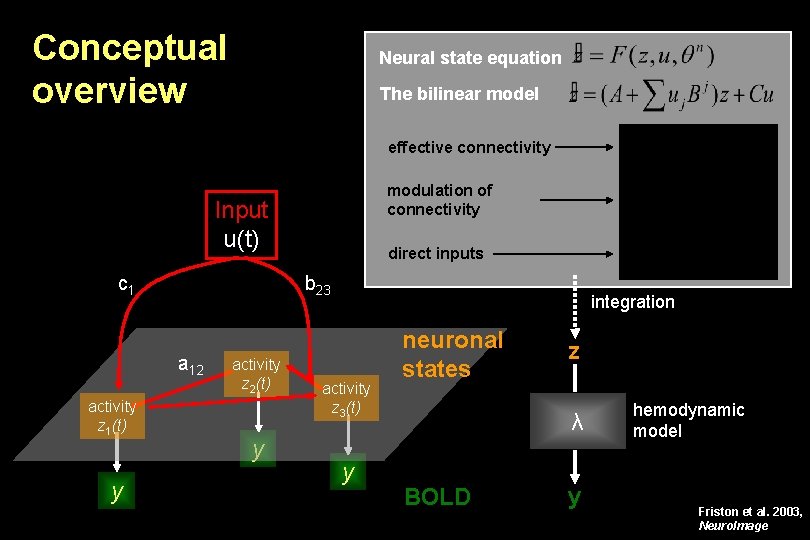
Conceptual overview Neural state equation The bilinear model effective connectivity modulation of connectivity Input u(t) c 1 b 23 a 12 activity z 1(t) y direct inputs activity z 2(t) y integration activity z 3(t) y neuronal states z λ BOLD y hemodynamic model Friston et al. 2003, Neuro. Image
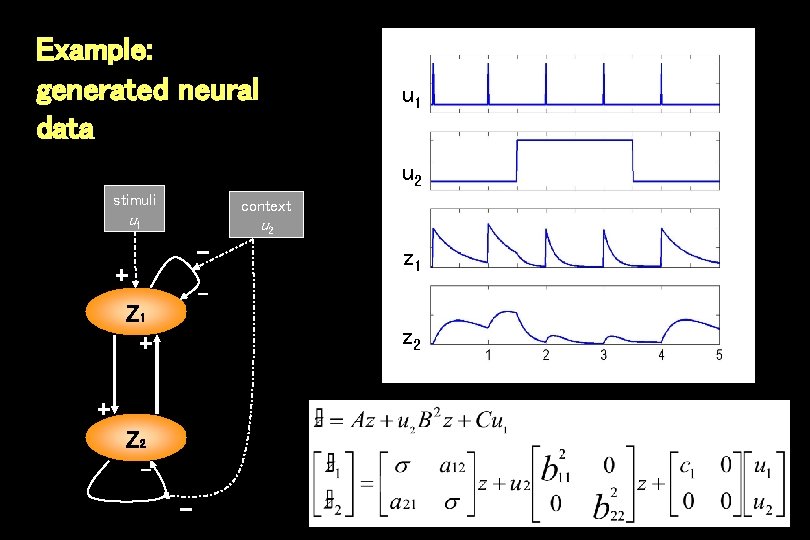
Example: generated neural data u 1 u 2 stimuli context u 1 Z 1 u 2 - + z Z 2 1 - Z 1 + z 2 + Z 2 -
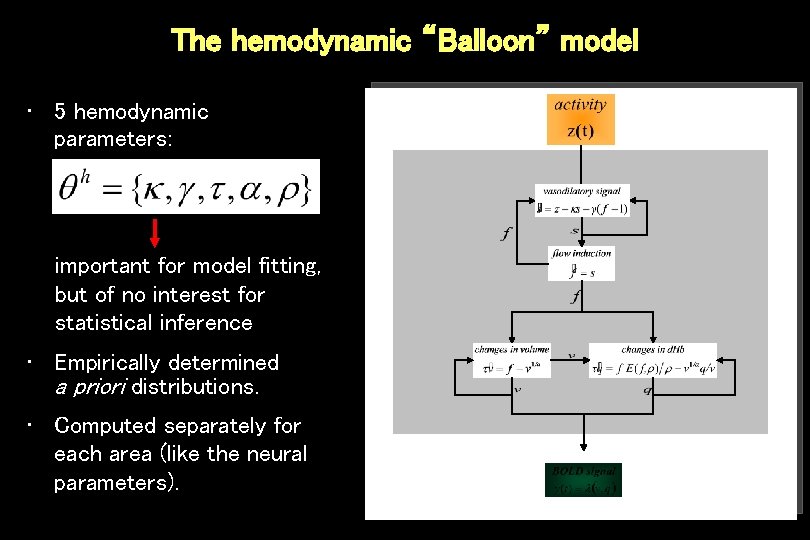
The hemodynamic “Balloon” model • 5 hemodynamic parameters: important for model fitting, but of no interest for statistical inference • Empirically determined a priori distributions. • Computed separately for each area (like the neural parameters).
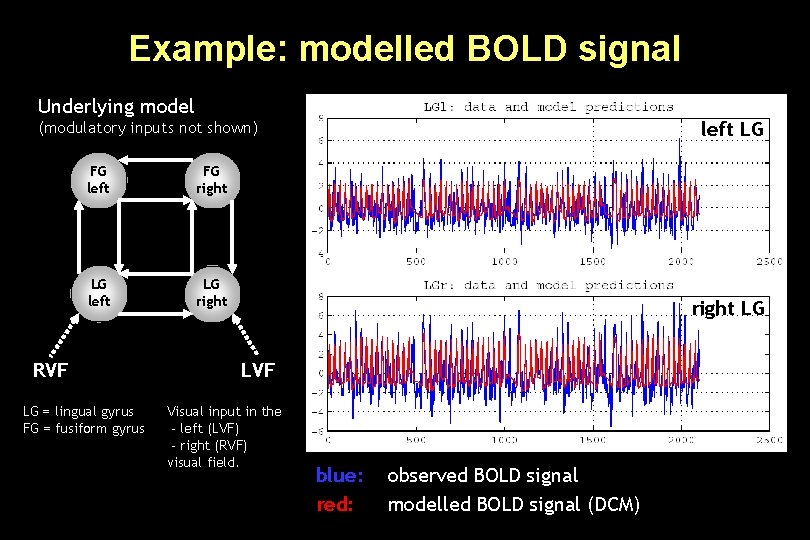
Example: modelled BOLD signal Underlying model left LG (modulatory inputs not shown) FG left FG right LG left LG right RVF LG = lingual gyrus FG = fusiform gyrus right LG LVF Visual input in the - left (LVF) - right (RVF) visual field. blue: red: observed BOLD signal modelled BOLD signal (DCM)

Overview • Classical approaches to functional & effective connectivity • Generic concepts of system analysis • DCM for f. MRI: – Neural dynamics and hemodynamics – Bayesian parameter estimation • Interpretation of parameters – Statistical inference – Bayesian model selection
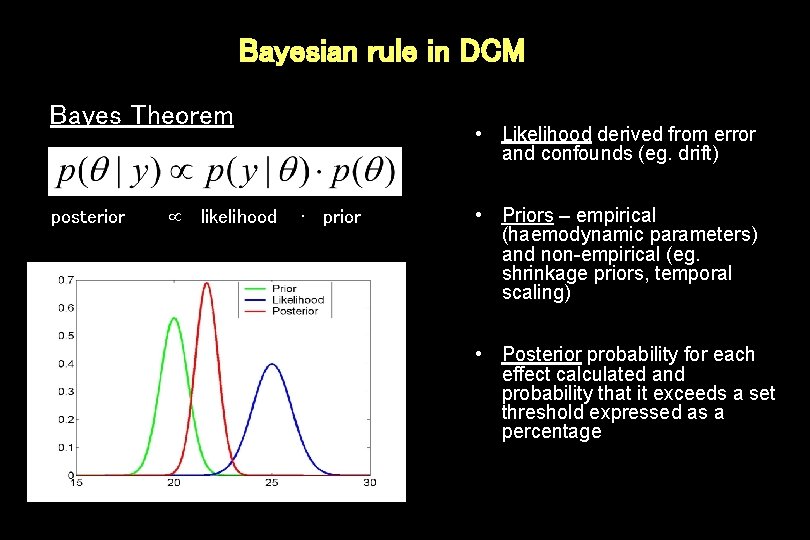
Bayesian rule in DCM Bayes Theorem posterior likelihood • Likelihood derived from error and confounds (eg. drift) ∙ prior • Priors – empirical (haemodynamic parameters) and non-empirical (eg. shrinkage priors, temporal scaling) • Posterior probability for each effect calculated and probability that it exceeds a set threshold expressed as a percentage
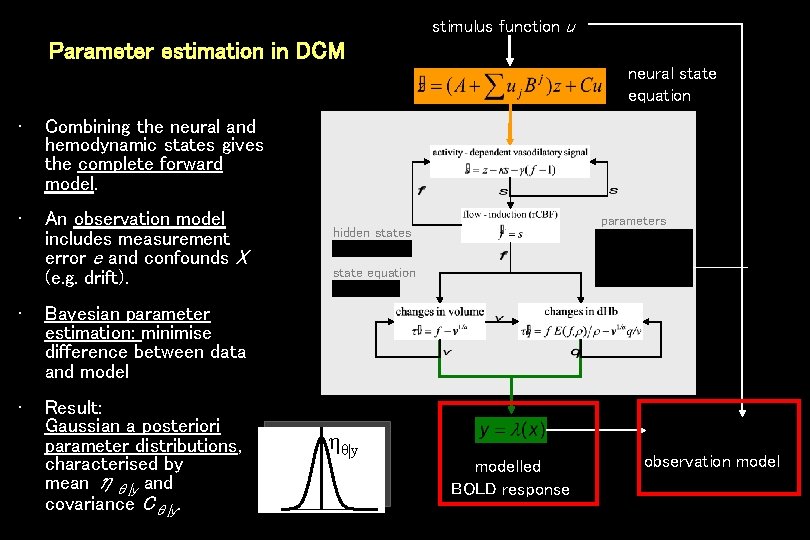
stimulus function u Parameter estimation in DCM neural state equation • Combining the neural and hemodynamic states gives the complete forward model. • An observation model includes measurement error e and confounds X (e. g. drift). • Bayesian parameter estimation: minimise difference between data and model • Result: Gaussian a posteriori parameter distributions, characterised by mean ηθ|y and covariance Cθ|y. parameters hidden states state equation ηθ|y modelled BOLD response observation model

Overview • Classical approaches to functional & effective connectivity • Generic concepts of system analysis • DCM for f. MRI: – Neural dynamics and hemodynamics – Bayesian parameter estimation • Interpretation of parameters – Statistical inference – Bayesian model selection
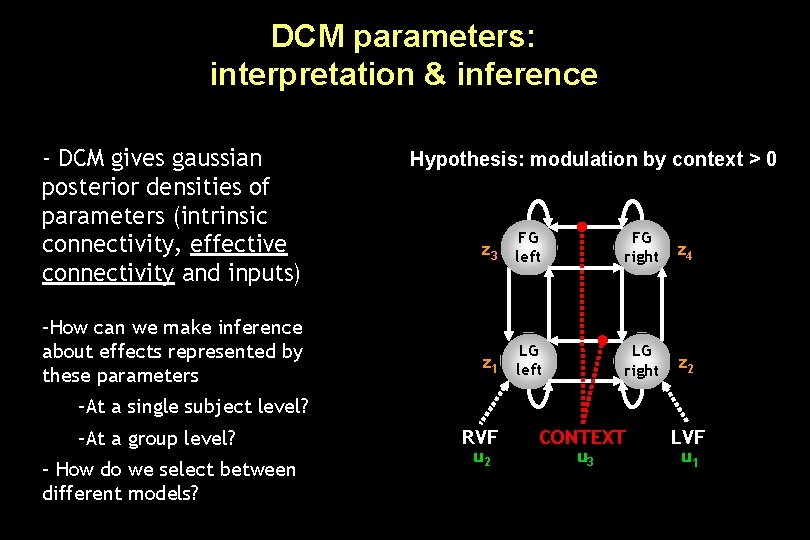
DCM parameters: interpretation & inference - DCM gives gaussian posterior densities of parameters (intrinsic connectivity, effective connectivity and inputs) –How can we make inference about effects represented by these parameters Hypothesis: modulation by context > 0 z 3 left FG FG right z 4 LG left LG right z 2 z 1 –At a single subject level? –At a group level? – How do we select between different models? RVF u 2 CONTEXT u 3 LVF u 1
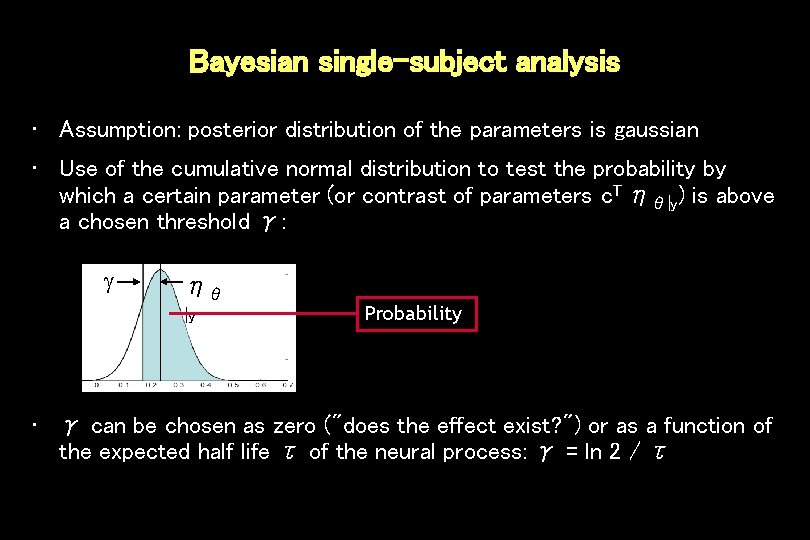
Bayesian single-subject analysis • Assumption: posterior distribution of the parameters is gaussian • Use of the cumulative normal distribution to test the probability by which a certain parameter (or contrast of parameters c. T ηθ|y) is above a chosen threshold γ: ηθ |y Probability ηθ |y • γ can be chosen as zero ("does the effect exist? ") or as a function of the expected half life τ of the neural process: γ = ln 2 / τ
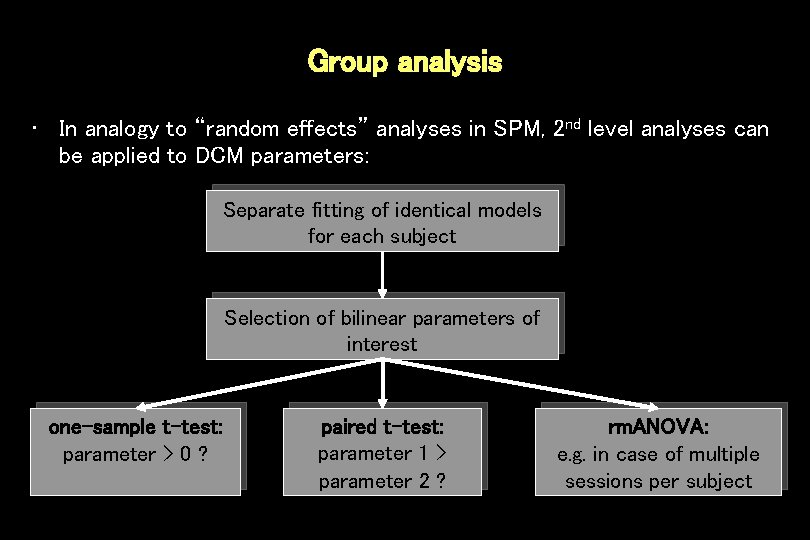
Group analysis • In analogy to “random effects” analyses in SPM, 2 nd level analyses can be applied to DCM parameters: Separate fitting of identical models for each subject Selection of bilinear parameters of interest one-sample t-test: parameter > 0 ? paired t-test: parameter 1 > parameter 2 ? rm. ANOVA: e. g. in case of multiple sessions per subject
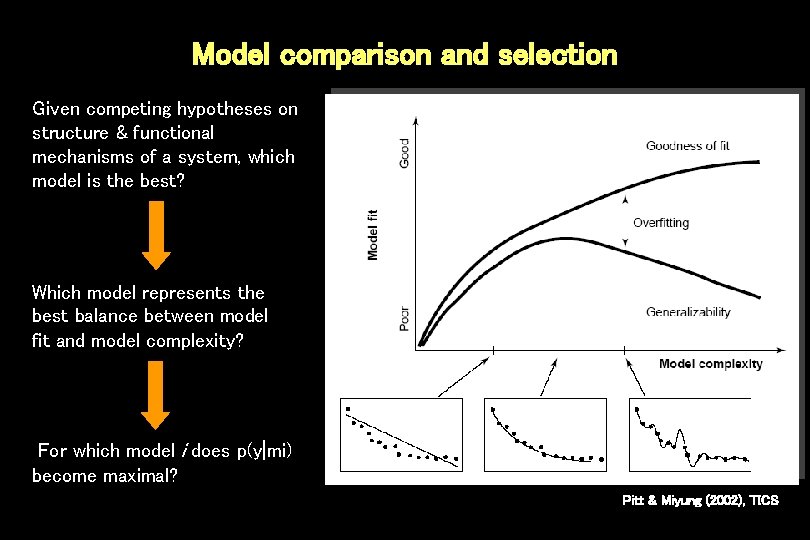
Model comparison and selection Given competing hypotheses on structure & functional mechanisms of a system, which model is the best? Which model represents the best balance between model fit and model complexity? For which model i does p(y|mi) become maximal? Pitt & Miyung (2002), TICS

Bayesian Model Selection Bayes theorem: Model evidence: The log model evidence can be represented as: Bayes factor: Penny et al. 2004, Neuro. Image
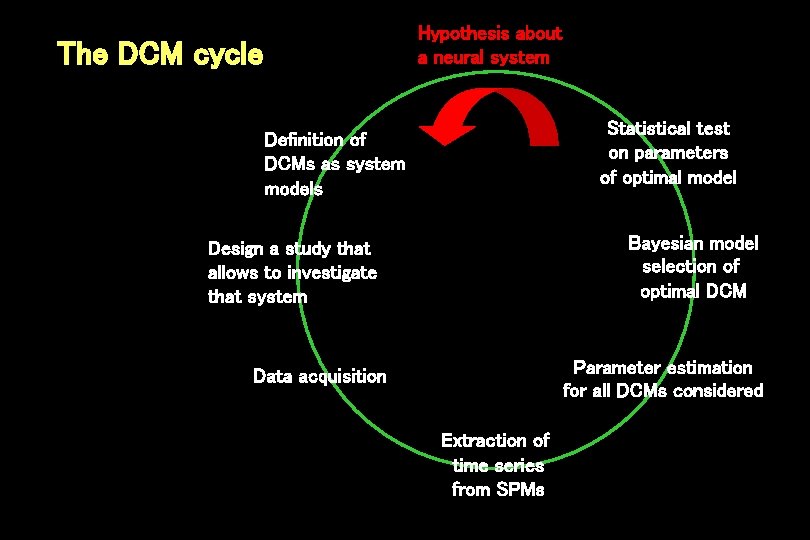
Hypothesis about a neural system The DCM cycle Statistical test on parameters of optimal model Definition of DCMs as system models Bayesian model selection of optimal DCM Design a study that allows to investigate that system Parameter estimation for all DCMs considered Data acquisition Extraction of time series from SPMs

Inference about DCM parameters: Bayesian fixed-effects group analysis Because the likelihood distributions from different subjects are independent, one can combine their posterior densities by using the posterior of one subject as the prior for the next: See: spm_dcm_average. m Neumann & Lohmann, Neuro. Image 2003 Under Gaussian assumptions this is easy to compute: group posterior covariance group posterior mean individual posterior covariances and means

Approximations to model evidence Laplace approximation: Unfortunately, the complexity term depends on the prior density, which is determined individually for each model to ensure stability. Therefore, we need other approximations to the model evidence. Akaike information criterion (AIC): Bayesian information criterion (BIC): Penny et al. 2004, Neuro. Image

DCM parameters = rate constants Integration of a first order linear differential equation gives an exponential function: The coupling parameter a is inversely proportional to the half life of z(t): The coupling parameter a thus describes the speed of the exponential growth/decay:
- Slides: 28