Dynamic Causal Modelling DCM Theory Burkhard Pleger Thanks

Dynamic Causal Modelling (DCM): Theory Burkhard Pleger Thanks to Klaas Enno Stephan Functional Imaging Lab Wellcome Dept. of Imaging Neuroscience Institute of Neurology University College London

Overview • DCMs as generic models of dynamic systems • Neural and hemodynamic levels in DCM • Parameter estimation – Priors in DCM – Bayesian parameter estimation • Interpretation of parameters • Bayesian model selection

System analyses in functional neuroimaging Functional specialisation Functional integration Analyses of regionally specific effects: which areas constitute a neuronal system? Analyses of inter-regional effects: what are the interactions between the elements of a given neuronal system? Functional connectivity Effective connectivity = the temporal correlation between spatially remote neurophysiological events MECHANISM-FREE = the influence that the elements of a neuronal system exert over another MECHANISTIC MODEL

Models of effective connectivity • Structural Equation Modelling (SEM) • Psycho-physiological interactions (PPI) • Multivariate autoregressive models (MAR) & Granger causality techniques • Kalman filtering • Volterra series • Dynamic Causal Modelling (DCM) Friston et al. , Neuro. Image 2003

Models of effective connectivity = system models. But what precisely is a system? • System = set of elements which interact in a spatially and temporally specific fashion. • System dynamics = change of state vector in time overall system state represented by state variables • Causal effects in the system: – interactions between elements – external inputs u • System parameters : specify the nature of the interactions • general state equation for nonautonomous systems change of state vector in time
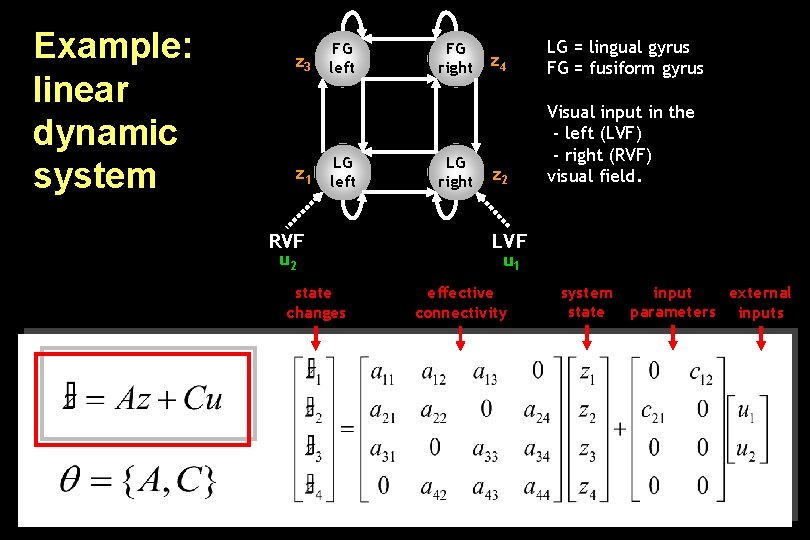
Example: linear dynamic system FG z 3 left z 1 LG left RVF u 2 state changes FG right LG right z 4 LG = lingual gyrus FG = fusiform gyrus z 2 Visual input in the - left (LVF) - right (RVF) visual field. LVF u 1 effective connectivity system state input external parameters inputs

Extension: bilinear dynamic system z 3 FG left FG right z 4 z 1 LG left LG right z 2 RVF u 2 CONTEXT u 3 LVF u 1

Bilinear state equation in DCM state changes intrinsic connectivity modulation of system connectivity state direct inputs m external inputs

Overview • DCMs as generic models of dynamic systems • Neural and hemodynamic levels in DCM • Parameter estimation – Priors in DCM – Bayesian parameter estimation • Interpretation of parameters • Bayesian model selection

DCM for f. MRI: the basic idea • Using a bilinear state equation, a cognitive system is modelled at its underlying neuronal level (which is not directly accessible for f. MRI). • The modelled neuronal dynamics (z) is transformed into area-specific BOLD signals (y) by a hemodynamic forward model (λ). The aim of DCM is to estimate parameters at the neuronal level such that the modelled BOLD signals are maximally similar to the experimentally measured BOLD signals. z λ y

Conceptual overview Neural state equation The bilinear model effective connectivity modulation of connectivity Input u(t) c 1 b 23 a 12 activity z 1(t) y direct inputs activity z 2(t) y integration activity z 3(t) y neuronal states z λ BOLD y hemodynamic model Friston et al. 2003, Neuro. Image
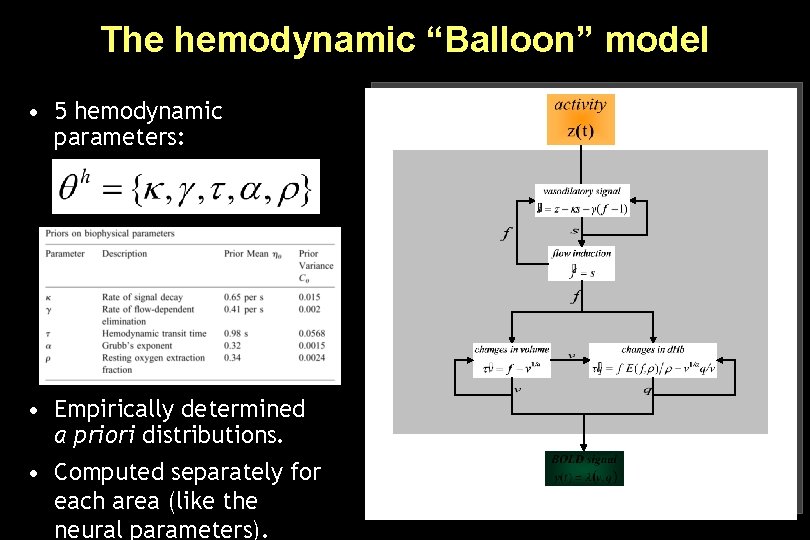
The hemodynamic “Balloon” model • 5 hemodynamic parameters: • Empirically determined a priori distributions. • Computed separately for each area (like the neural parameters).
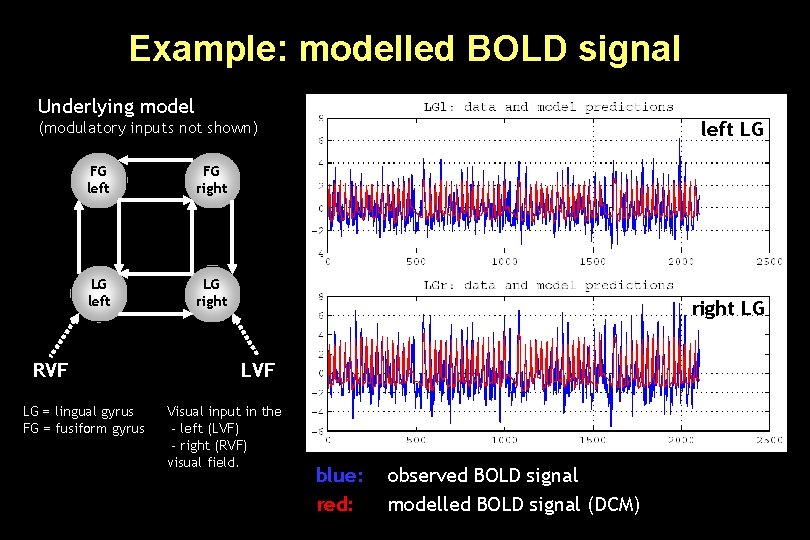
Example: modelled BOLD signal Underlying model left LG (modulatory inputs not shown) FG left FG right LG left LG right RVF LG = lingual gyrus FG = fusiform gyrus right LG LVF Visual input in the - left (LVF) - right (RVF) visual field. blue: red: observed BOLD signal modelled BOLD signal (DCM)

Overview • DCMs as generic models of dynamic systems • Neural and hemodynamic levels in DCM • Parameter estimation – Priors in DCM – Bayesian parameter estimation • Interpretation of parameters • Bayesian model selection
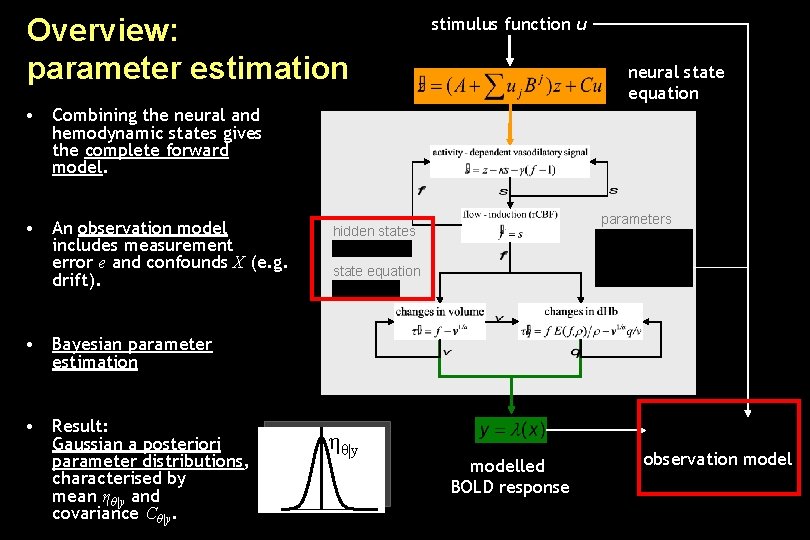
Overview: parameter estimation stimulus function u neural state equation • Combining the neural and hemodynamic states gives the complete forward model. • An observation model includes measurement error e and confounds X (e. g. drift). parameters hidden states state equation • Bayesian parameter estimation • Result: Gaussian a posteriori parameter distributions, characterised by mean ηθ|y and covariance Cθ|y. ηθ|y modelled BOLD response observation model

Priors in DCM • needed for Bayesian estimation, embody constraints on parameter estimation • express our prior knowledge or “belief” about parameters of the model • hemodynamic parameters: empirical priors • temporal scaling: principled prior, self-inhibition • coupling parameters: shrinkage priors Bayes Theorem posterior likelihood ∙ prior

Priors in DCM, self-inhibition • self-inhibition: ensured by priors on the decay rate constant σ (ησ=1, Cσ=0. 105) → these allow for neural transients with a half life in the range of 300 ms to 2 seconds NB: a single rate constant for all regions! • Identical temporal scaling in all areas by factorising A and B with σ: all connection strengths are relative to the self-connections.

Priors in DCM, coupling • system stability: in the absence of input, the neuronal states must return to a stable mode → constraints on prior variance of intrinsic connections (A) • shrinkage priors for coupling parameters (η=0) → conservative estimates!

Shrinkage Priors Small & variable effect Large & variable effect Small but clear effect Large & clear effect
- Slides: 19