Drug Metabolism Phase II Dr Arthur G Roberts

Drug Metabolism: Phase II Dr. Arthur G. Roberts

Outline • • • Introduction UDP-glucuronosyltransferases (UGTs) Sulfotransferases (SULTs) Glutathione-S-Transferases (GSTs) N-acetyltransferases (NATs) Methyltransferases – Catechol O-Methyltransferases (COMTs) – Histamine Methyltransferases (HMTs) – Thiopurine Methyltransferases (TPMTs)
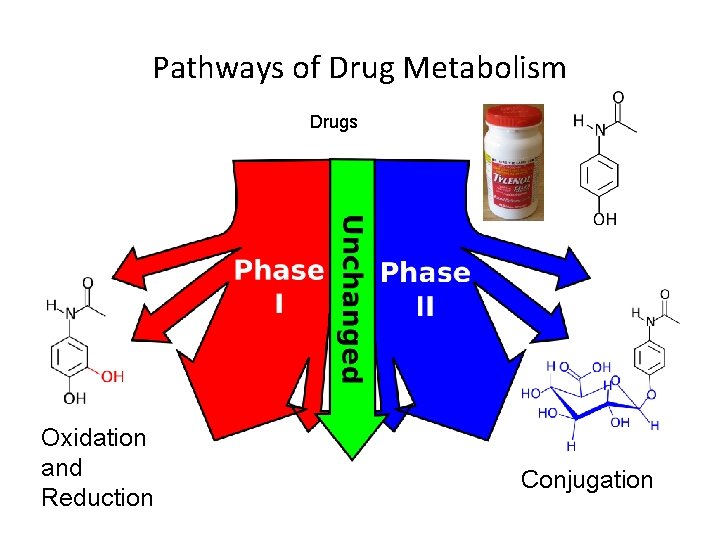
Pathways of Drug Metabolism Drugs Oxidation and Reduction Conjugation
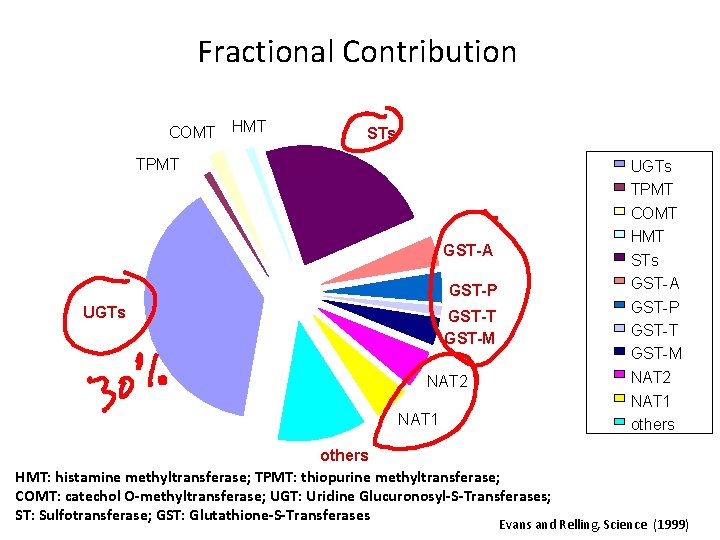
Fractional Contribution COMT HMT STs TPMT UGTs TPMT COMT HMT STs GST-A GST-P UGTs GST-P GST-T GST-M NAT 2 NAT 1 others HMT: histamine methyltransferase; TPMT: thiopurine methyltransferase; COMT: catechol O-methyltransferase; UGT: Uridine Glucuronosyl-S-Transferases; ST: Sulfotransferase; GST: Glutathione-S-Transferases Evans and Relling, Science (1999)
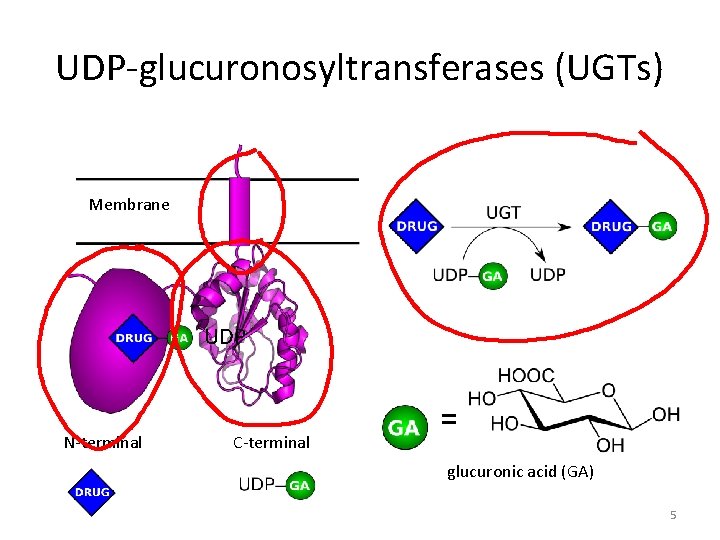
UDP-glucuronosyltransferases (UGTs) Membrane UDP N-terminal C-terminal = glucuronic acid (GA) 5
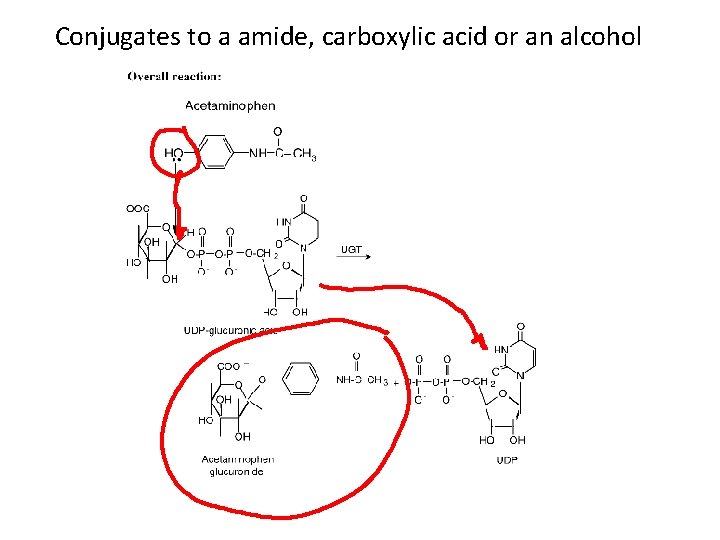
Conjugates to a amide, carboxylic acid or an alcohol
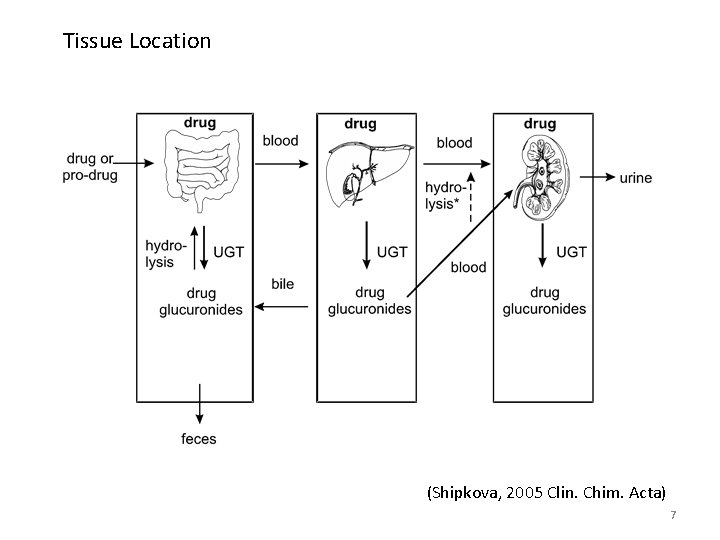
Tissue Location (Shipkova, 2005 Clin. Chim. Acta) 7
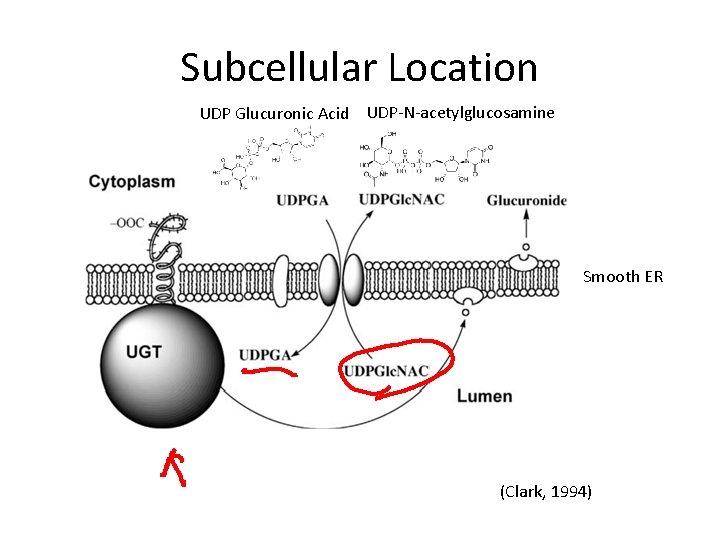
Subcellular Location UDP Glucuronic Acid UDP-N-acetylglucosamine Smooth ER (Clark, 1994)

Substrate Specificity • Promiscuous • Drug-drug interaction • UGT-UGT Interaction
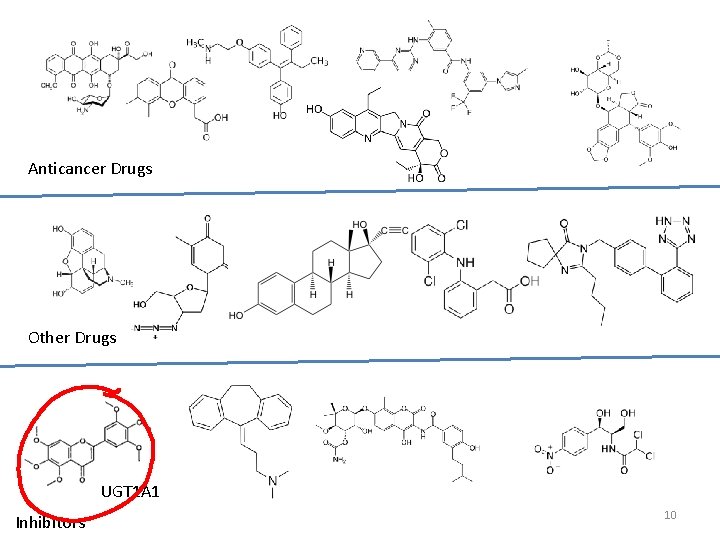
Anticancer Drugs Other Drugs UGT 1 A 1 Inhibitors 10
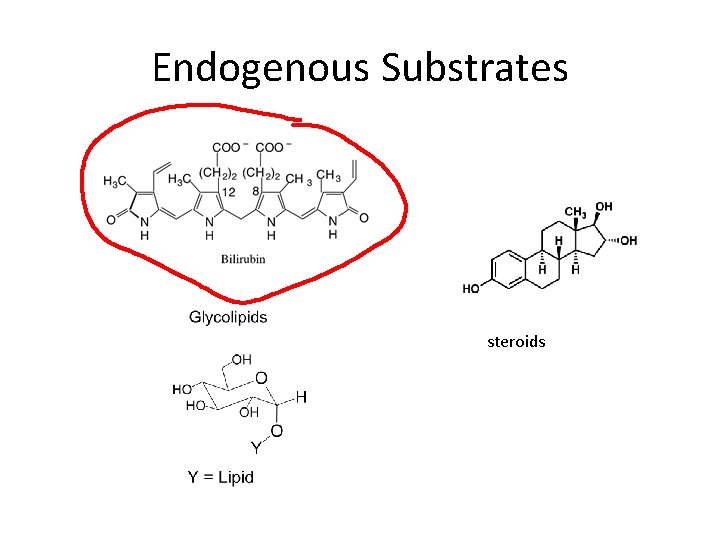
Endogenous Substrates steroids
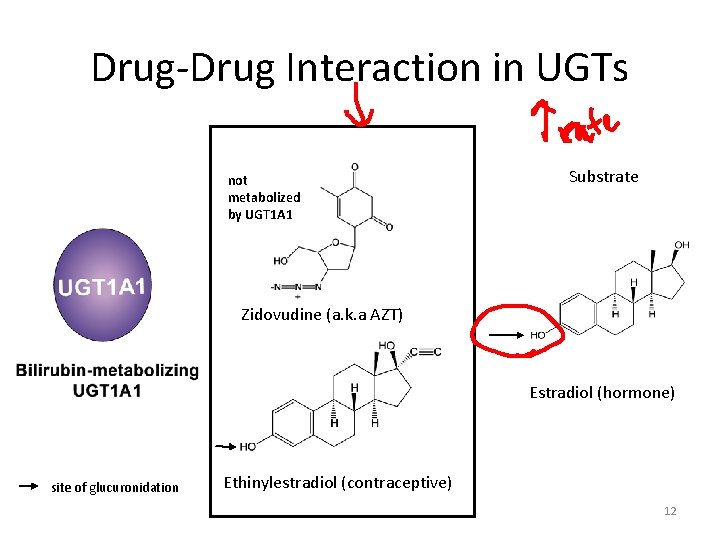
Drug-Drug Interaction in UGTs not metabolized by UGT 1 A 1 Substrate Zidovudine (a. k. a AZT) Estradiol (hormone) site of glucuronidation Ethinylestradiol (contraceptive) 12
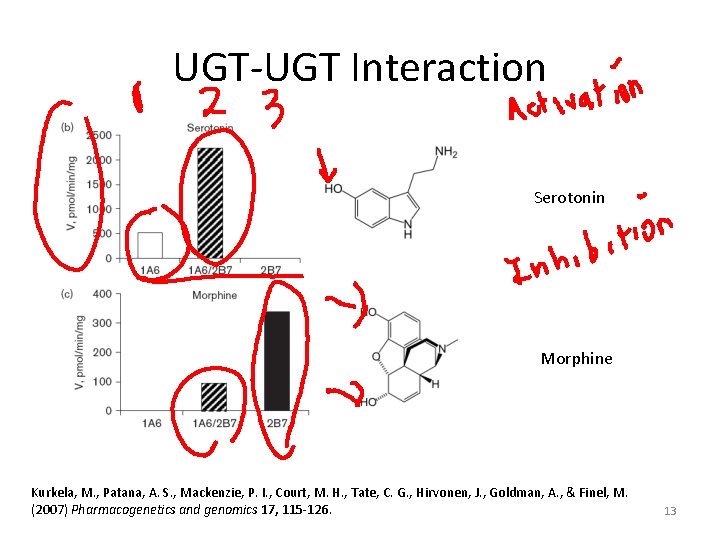
UGT-UGT Interaction Serotonin Morphine Kurkela, M. , Patana, A. S. , Mackenzie, P. I. , Court, M. H. , Tate, C. G. , Hirvonen, J. , Goldman, A. , & Finel, M. (2007) Pharmacogenetics and genomics 17, 115 -126. 13

Families • UGT 1 A – Planar and bulky phenols (e. g. estrogens), amines and unsaturated carboxylic acids – UGT 1 A 1: bilirubin and morphine • UGT 2 B – drugs, steroids, bile acids and unsaturated carboxylic acids
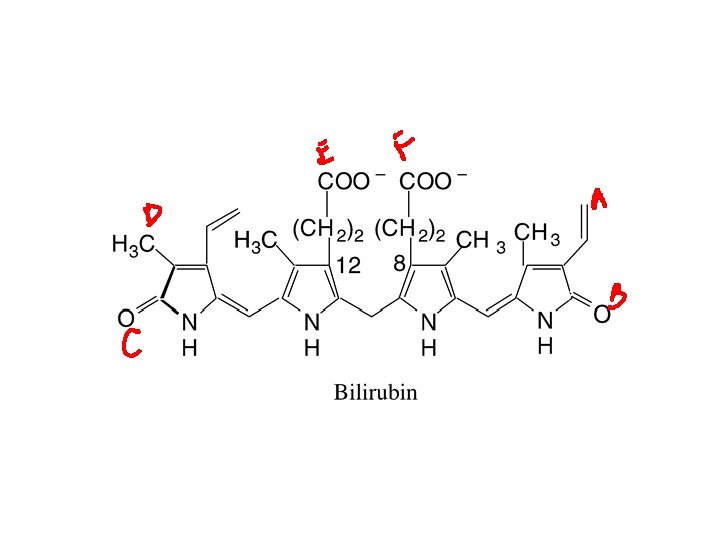
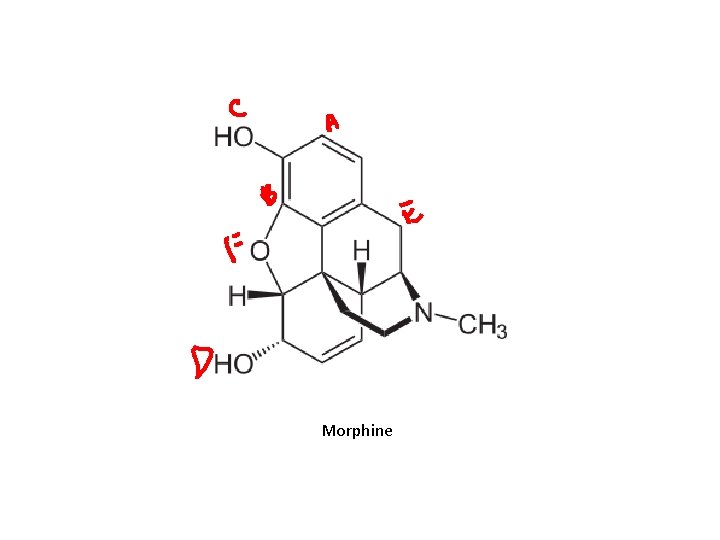
Morphine
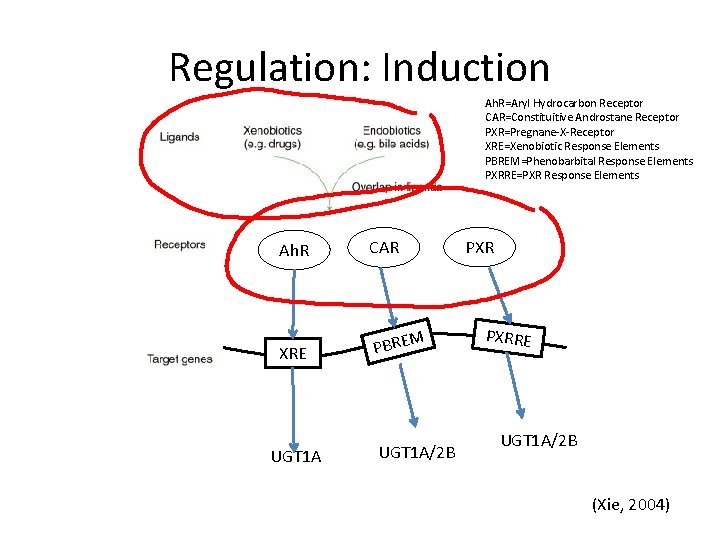
Regulation: Induction Ah. R=Aryl Hydrocarbon Receptor CAR=Constituitive Androstane Receptor PXR=Pregnane-X-Receptor XRE=Xenobiotic Response Elements PBREM=Phenobarbital Response Elements PXRRE=PXR Response Elements Ah. R CAR XRE PBRE UGT 1 A PXR M UGT 1 A/2 B PXRRE UGT 1 A/2 B (Xie, 2004)

Diseases • Hyperbilirubinemia – Gilbert’s syndrome (20 -30% UGT 1 A 1 Activity) – Crigler-Najjar syndrome • Type I (0% UGT 1 A 1 Activity) • Type II (10% UGT 1 A 1 Activity) • Gray Baby Syndrome (Chloramphenicol) • Neonatal Jaundice – UGT 1 A 1*28/*28 – light therapy • Autoimmune Hepatitis
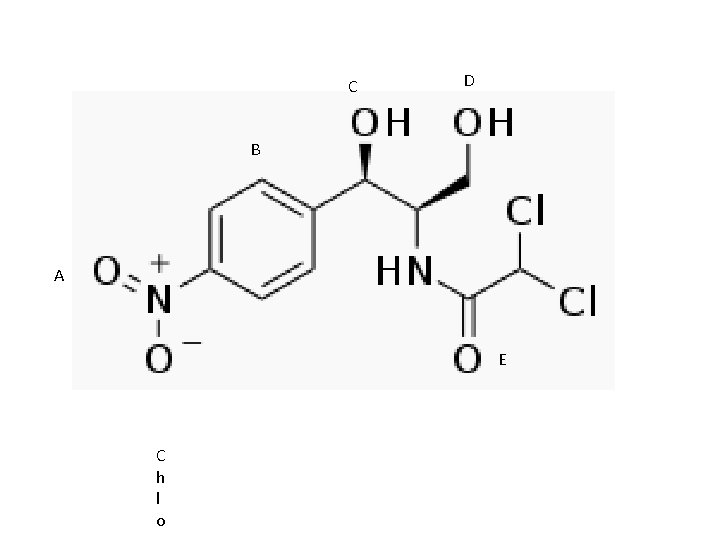
C D B A E C h l o
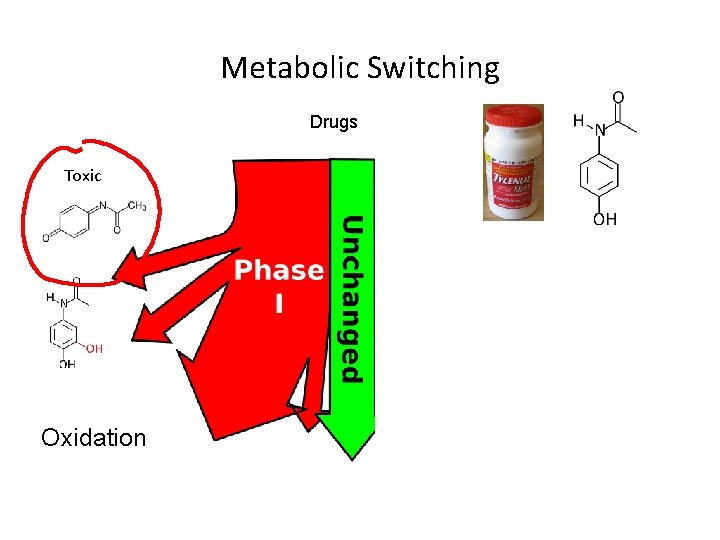
Metabolic Switching Drugs Toxic Oxidation Conjugation
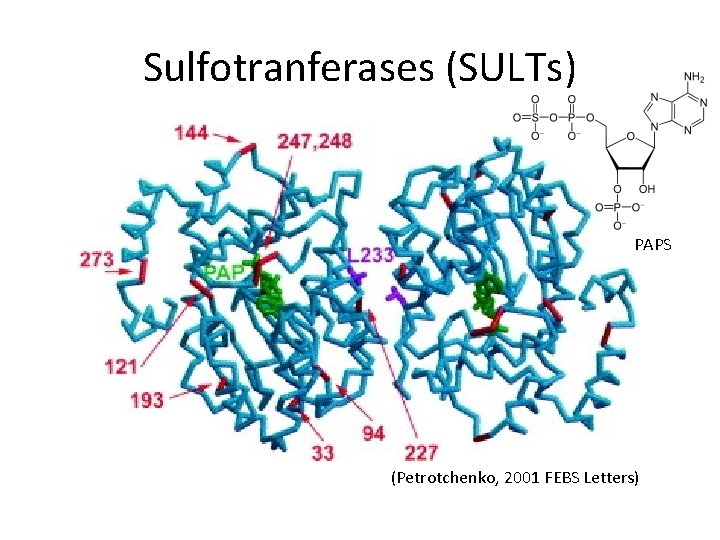
Sulfotranferases (SULTs) PAPS (Petrotchenko, 2001 FEBS Letters)
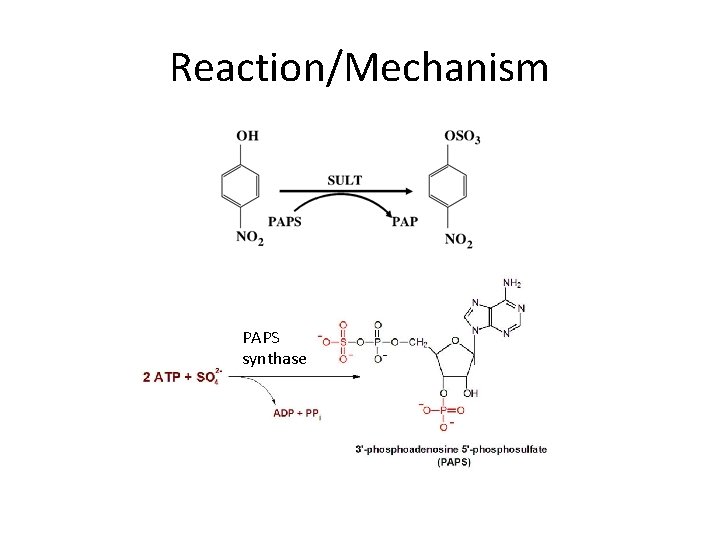
Reaction/Mechanism PAPS synthase
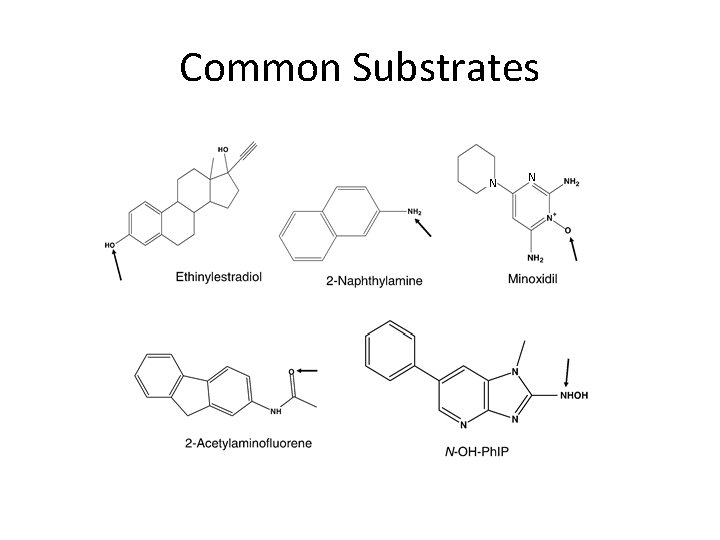
Common Substrates N N
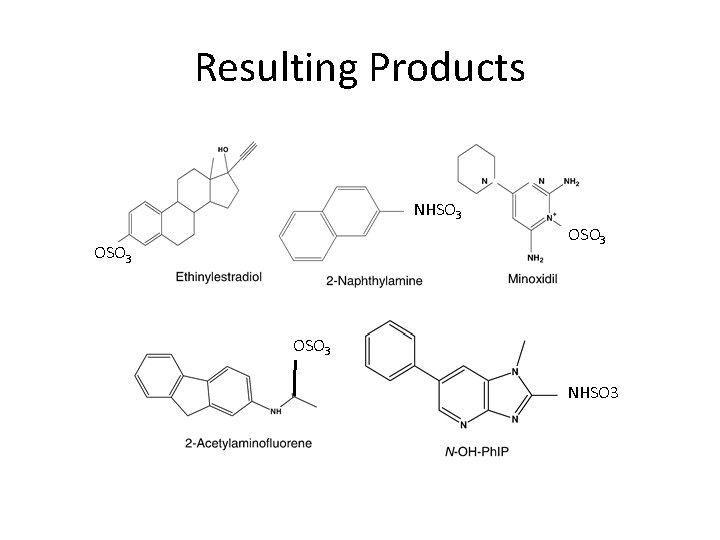
Resulting Products NHSO 3 OSO 3 NHSO 3

Subcellular Location • Cytosolic – Homodimers/Heterodimers • Membrane-bound – Golgi Apparatus

Families and General Substrate Specificities • SULT 1 – Phenols, Catechols, Benzylic Alcohols, Amines, Estrogens • SULT 2 – Steroids
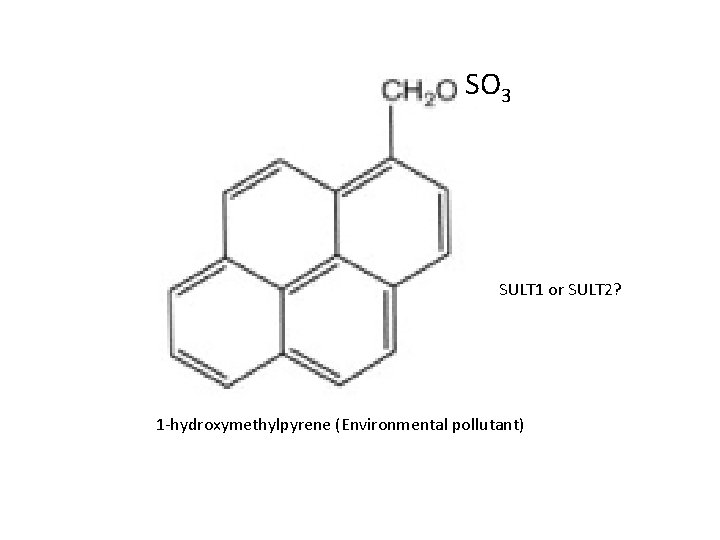
SO 3 SULT 1 or SULT 2? 1 -hydroxymethylpyrene (Environmental pollutant)
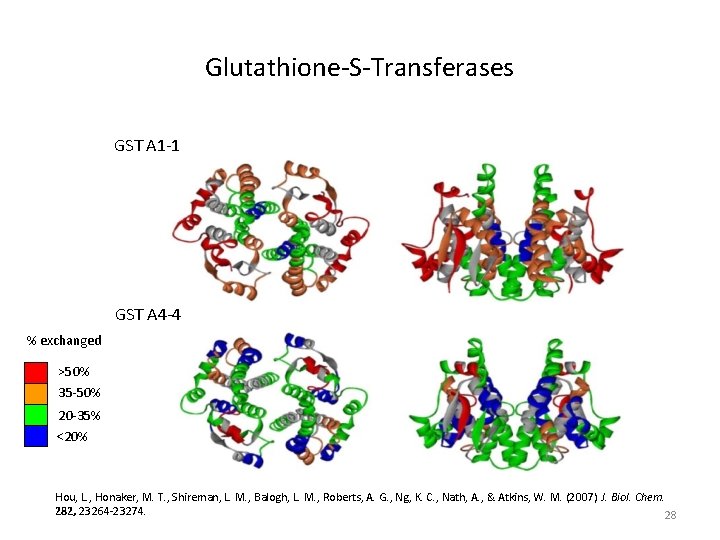
Glutathione-S-Transferases GST A 1 -1 GST A 4 -4 % exchanged >50% 35 -50% 20 -35% <20% Hou, L. , Honaker, M. T. , Shireman, L. M. , Balogh, L. M. , Roberts, A. G. , Ng, K. C. , Nath, A. , & Atkins, W. M. (2007) J. Biol. Chem. 282, 23264 -23274. 28
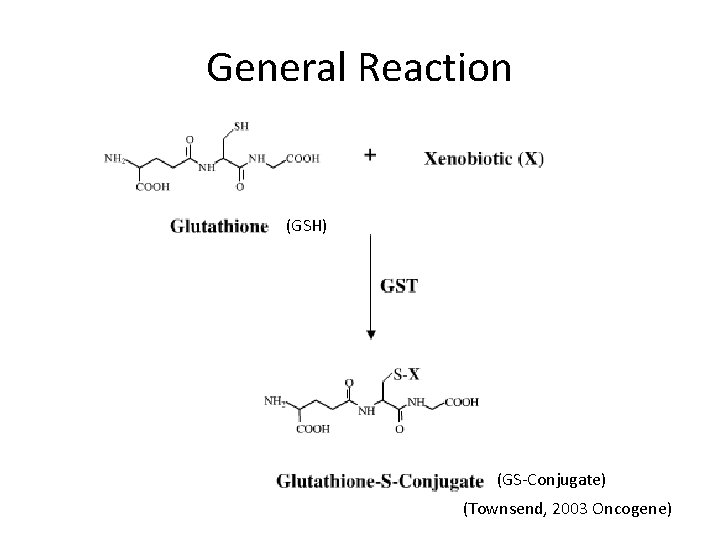
General Reaction (GSH) (GS-Conjugate) (Townsend, 2003 Oncogene)
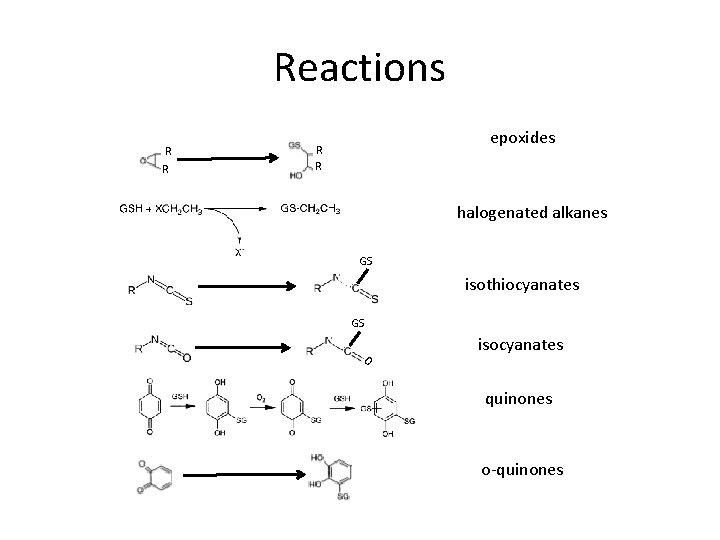
Reactions R R epoxides R R halogenated alkanes GS isothiocyanates GS O isocyanates quinones o-quinones
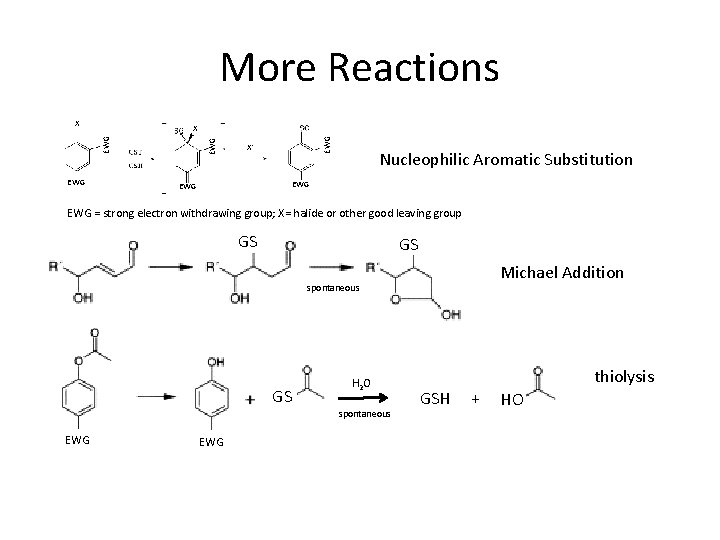
More Reactions X EWG EWG X X- Nucleophilic Aromatic Substitution EWG EWG = strong electron withdrawing group; X= halide or other good leaving group GS GS Michael Addition spontaneous GS H 2 O spontaneous EWG thiolysis GSH + HO
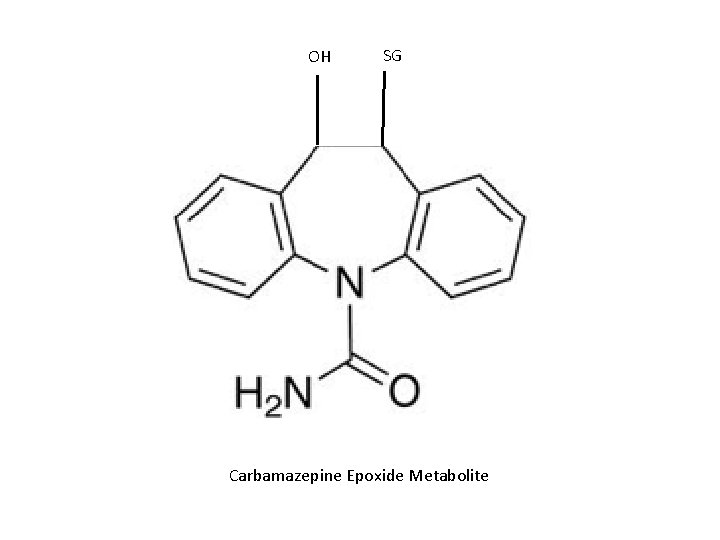
OH SG Carbamazepine Epoxide Metabolite
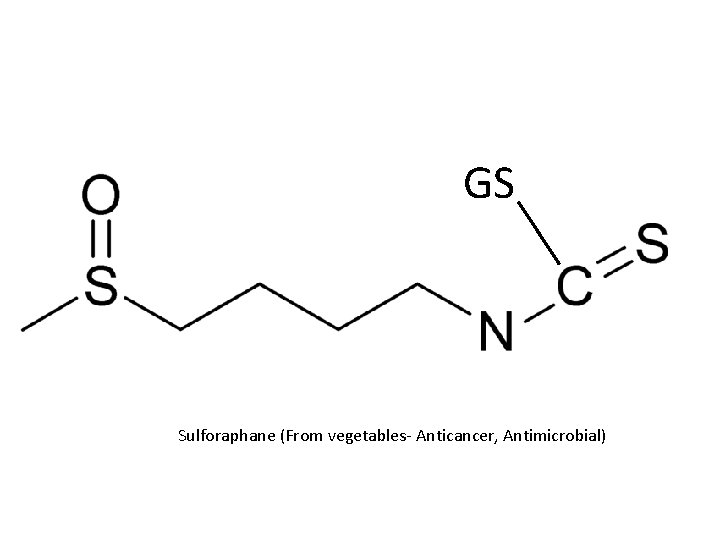
GS Sulforaphane (From vegetables- Anticancer, Antimicrobial)
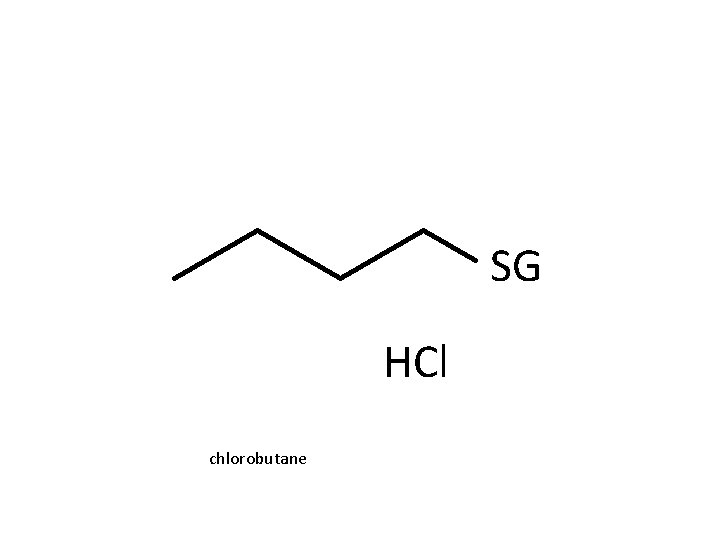
SG HCl chlorobutane
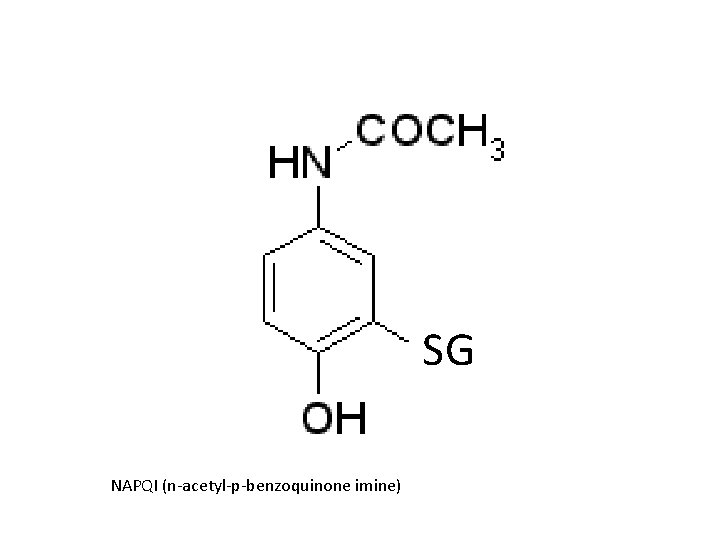
SG NAPQI (n-acetyl-p-benzoquinone imine)

Families and Tissue Locations • alpha (A) - Liver • kappa (K) • mu (M) – brain, lung , testes, lymphoblasts, skeletal muscle • omega (O) • pi (P) – all tissues except Liver • sigma (S) – macrophages, placenta and fat – prostaglandin metabolism • theta (T) • zeta (Z)

Location • Tissue – Everywhere • Subcellular – Cytosolic, Mitochondrial and Microsomal

Regulation • GST A, M and P – electrophiles that produce reactive oxygen – antioxidant response element

Genetics (Often Missing) • GST M 1 – <50% Caucasians and East Asians – >90% Polynesians and Micronesians – >50% Africans – absence increases risk for lung and bladder cancer • GST T 1 – 20% missing in Caucasians – 50% missing in Asians – 40% missing in Africans
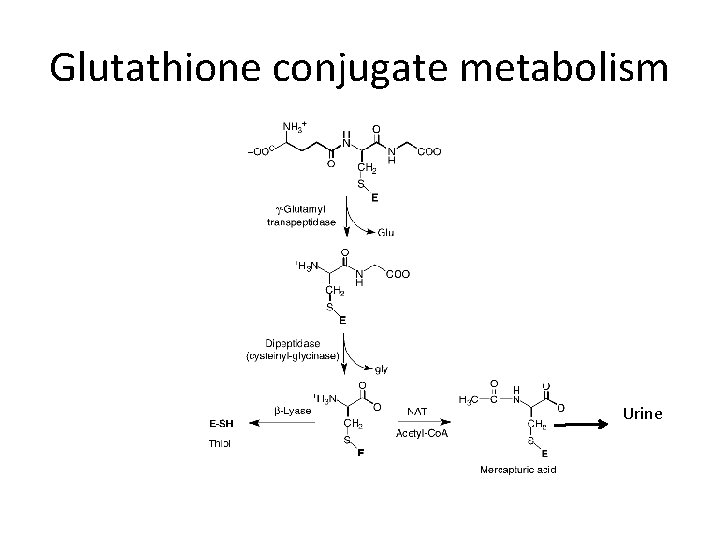
Glutathione conjugate metabolism Urine
N-acetyltransferases (NAT) Acetyl-Co. A
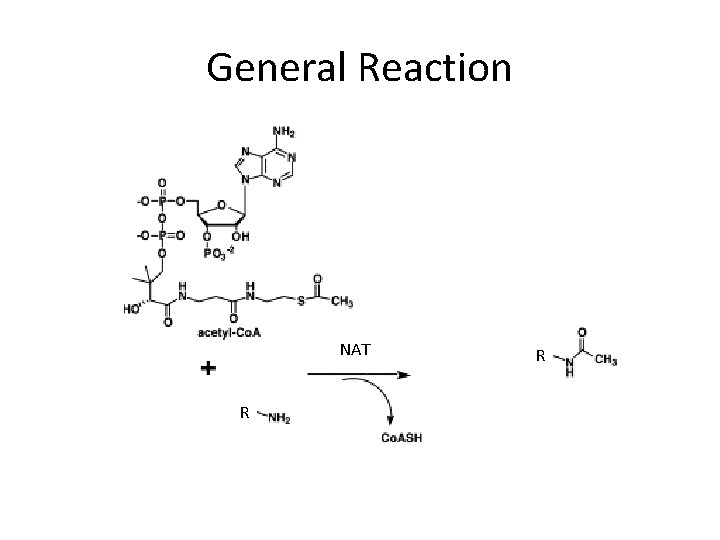
General Reaction NAT R R
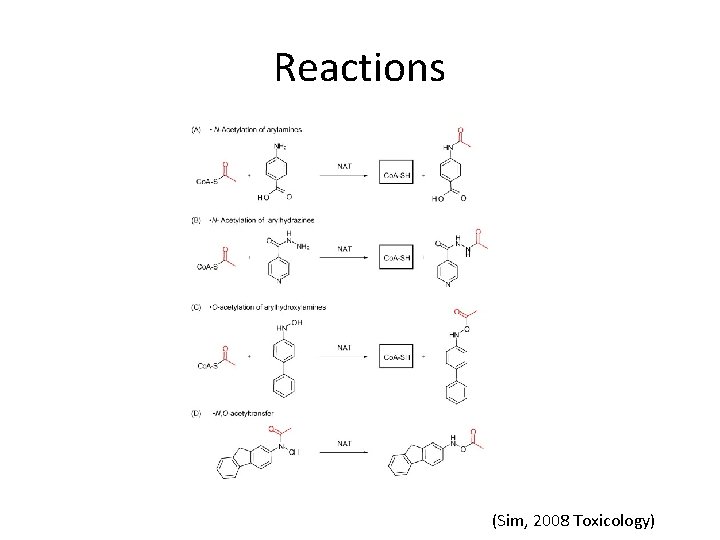
Reactions (Sim, 2008 Toxicology)
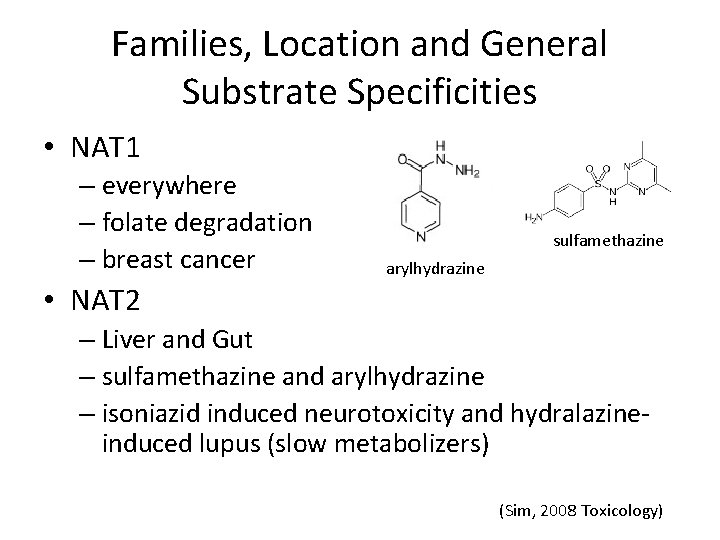
Families, Location and General Substrate Specificities • NAT 1 – everywhere – folate degradation – breast cancer sulfamethazine arylhydrazine • NAT 2 – Liver and Gut – sulfamethazine and arylhydrazine – isoniazid induced neurotoxicity and hydralazineinduced lupus (slow metabolizers) (Sim, 2008 Toxicology)
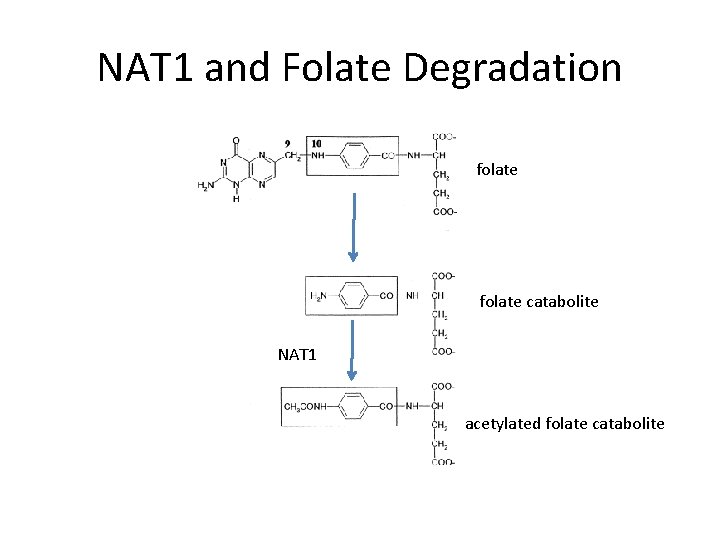
NAT 1 and Folate Degradation folate catabolite NAT 1 acetylated folate catabolite
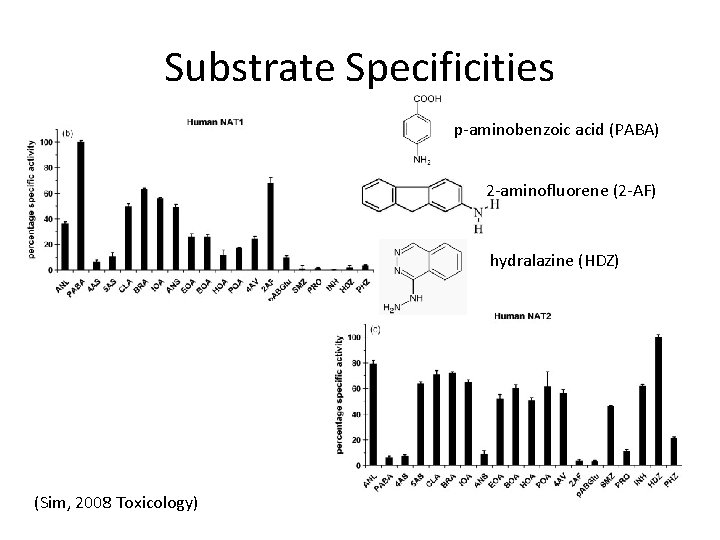
Substrate Specificities p-aminobenzoic acid (PABA) 2 -aminofluorene (2 -AF) hydralazine (HDZ) (Sim, 2008 Toxicology)
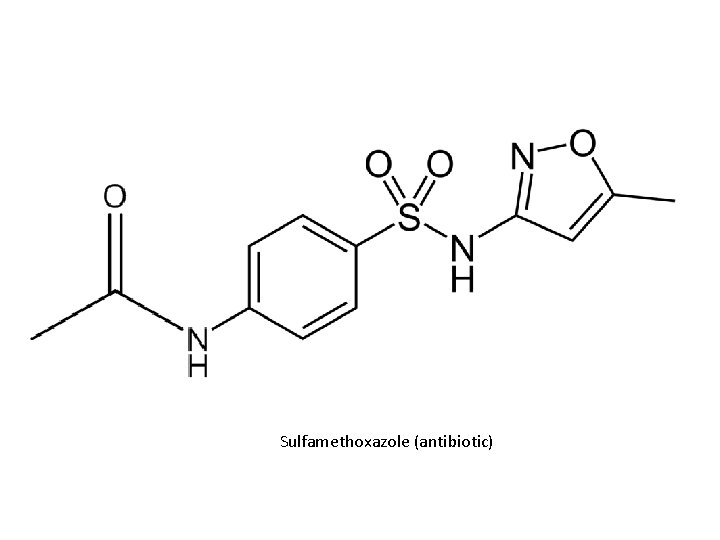
Sulfamethoxazole (antibiotic)
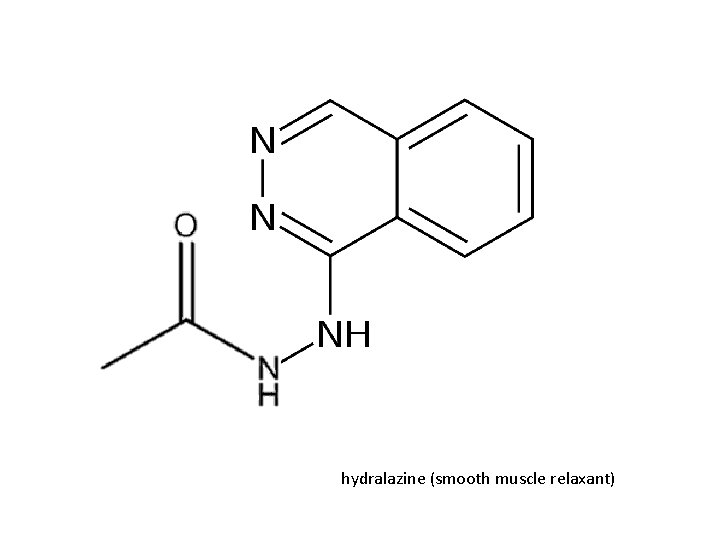
hydralazine (smooth muscle relaxant)
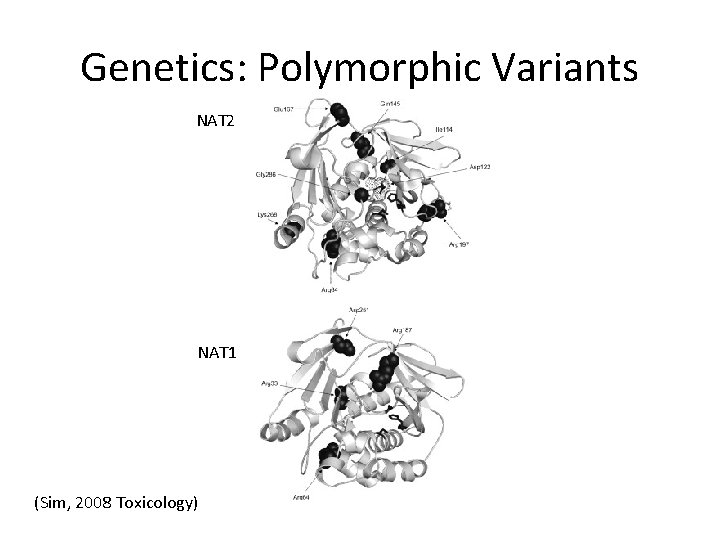
Genetics: Polymorphic Variants NAT 2 NAT 1 (Sim, 2008 Toxicology)
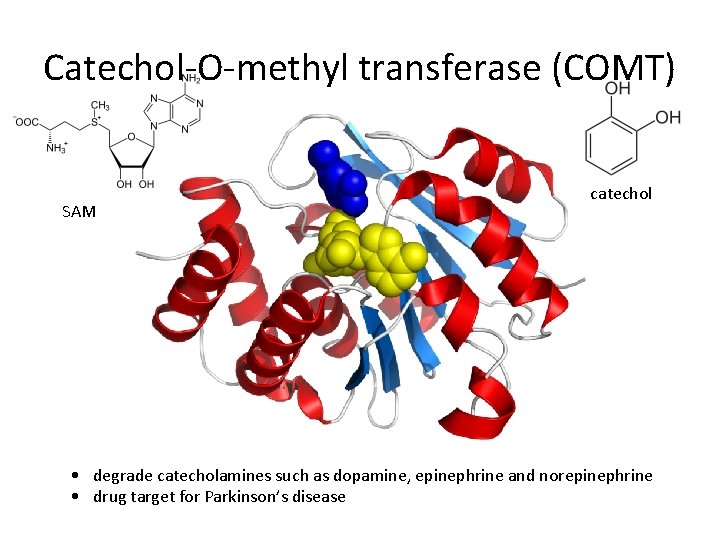
Catechol-O-methyl transferase (COMT) SAM catechol • degrade catecholamines such as dopamine, epinephrine and norepinephrine • drug target for Parkinson’s disease

Location • Tissue – Liver, some in brain. • Subcellular – cytosolic (soluble), rough endoplasmic reticulum (membrane-bound)
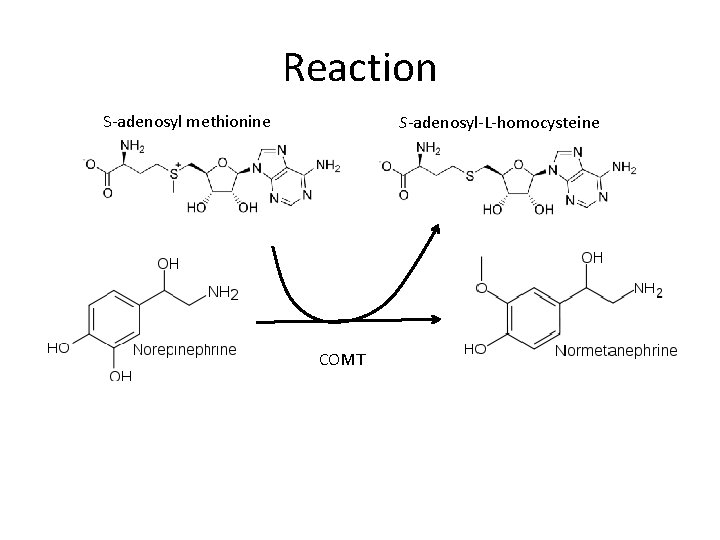
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine COMT
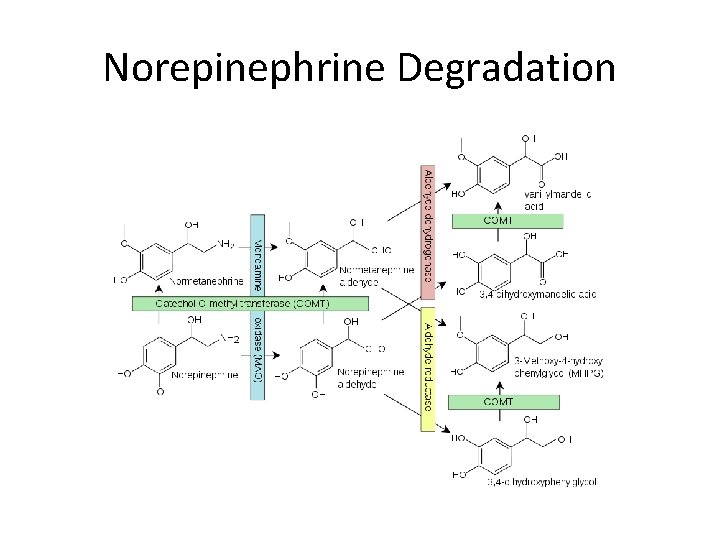
Norepinephrine Degradation
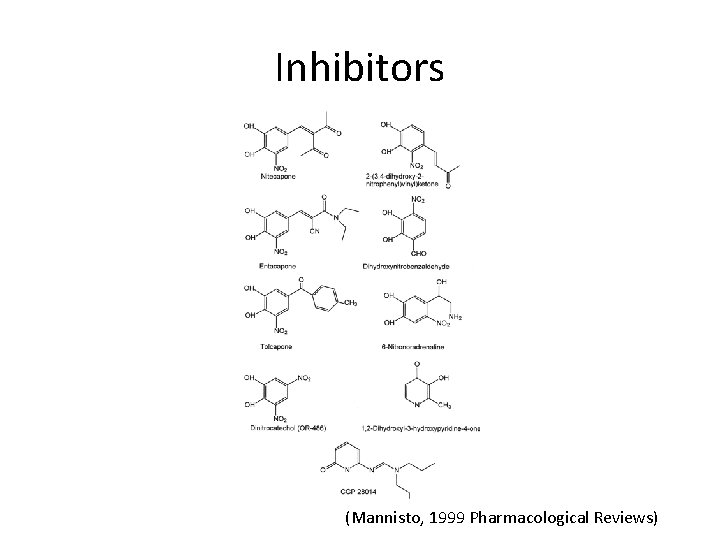
Inhibitors (Mannisto, 1999 Pharmacological Reviews)
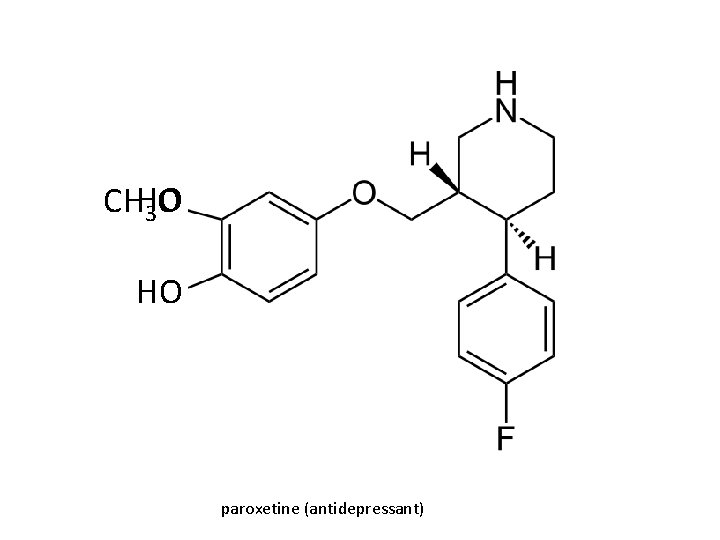
CHHO 3 O HO paroxetine (antidepressant)

Genetics: Polymorphisms • V 158 M – effects on cognitive function – schizophrenia
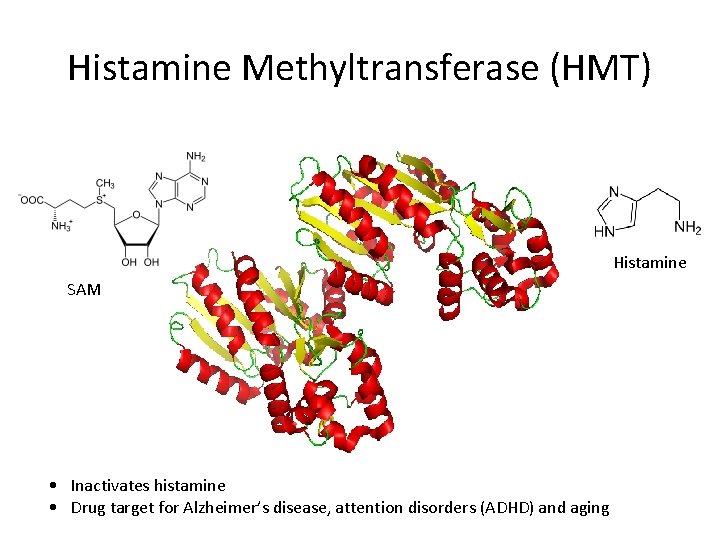
Histamine Methyltransferase (HMT) Histamine SAM • Inactivates histamine • Drug target for Alzheimer’s disease, attention disorders (ADHD) and aging
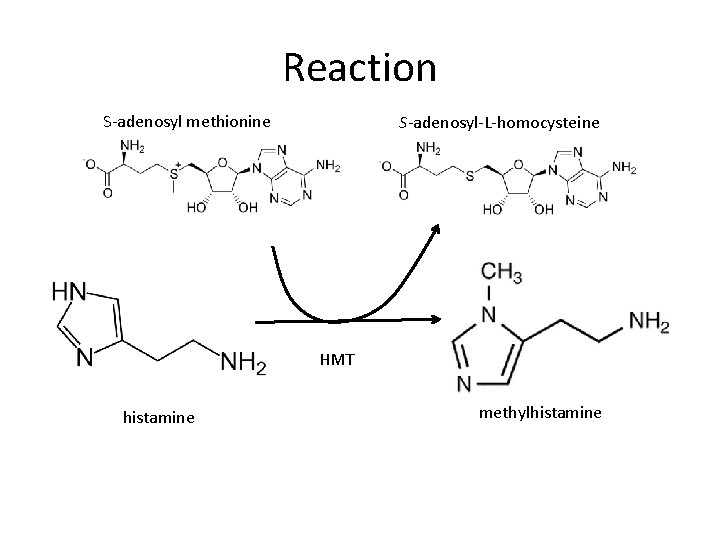
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine HMT histamine methylhistamine

Genetics: Polymorphisms • Residue 105 – T 105 (high activity phenotype, 90%) – I 105 (low activity phenotype, 10%)
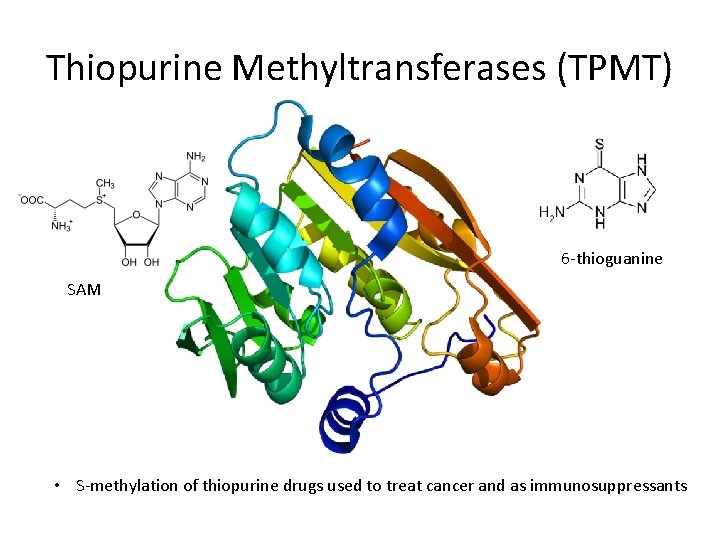
Thiopurine Methyltransferases (TPMT) 6 -thioguanine SAM • S-methylation of thiopurine drugs used to treat cancer and as immunosuppressants
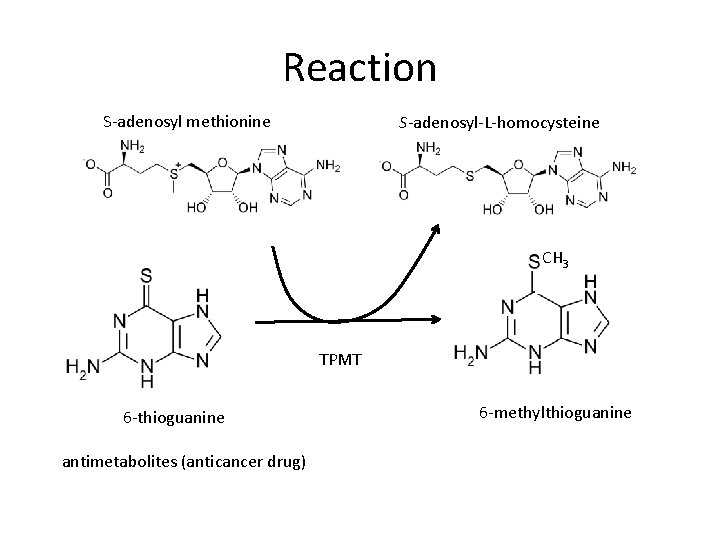
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine CH 3 TPMT 6 -thioguanine antimetabolites (anticancer drug) 6 -methylthioguanine
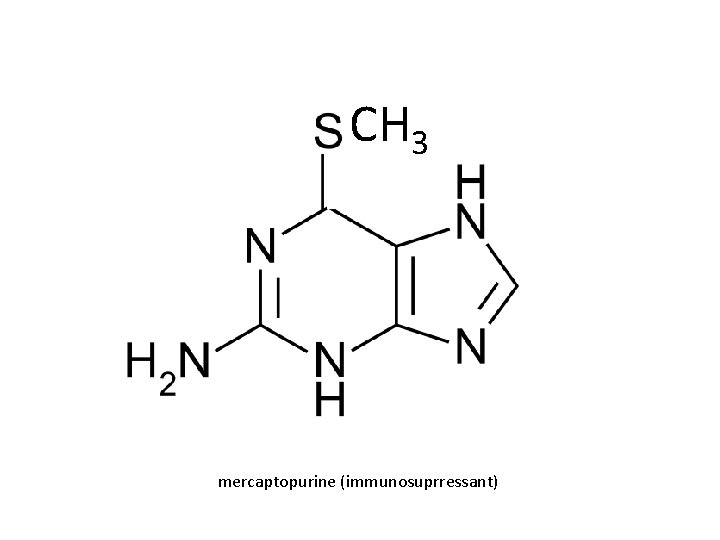
CH 3 mercaptopurine (immunosuprressant)
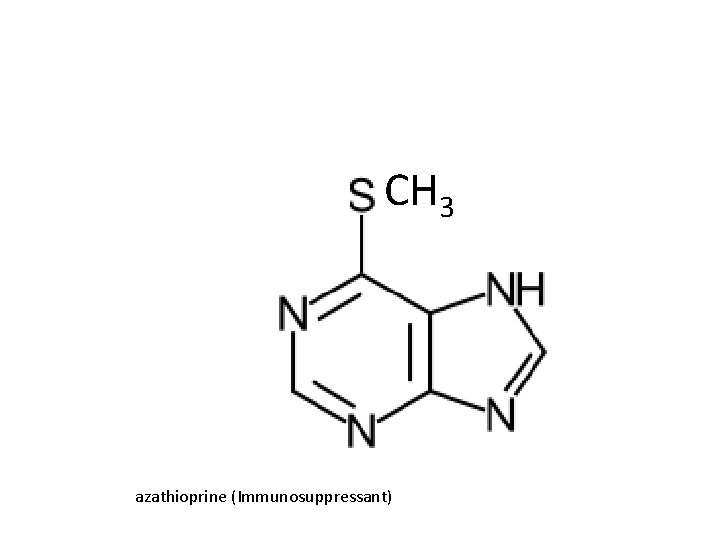
CH 3 azathioprine (Immunosuppressant)

Genetics: Polymorphisms • Low or No Activity Variants – TPMT*2 (2. 2%, G 238 C), TPMT*3 A (1. 5%, G 460 A and A 719 G), TPMT*3 B (0. 2%, G 460 A) and TPMT*3 C (1. 0%, A 719 G) – bone marrow toxicity (unmetabolized drug) – significant differences between ethnic groups

End of Phase II
- Slides: 65