Drug Design drug discovery Jerome Baudry Assistant Professor

Drug Design / drug discovery Jerome Baudry Assistant Professor BCMB UT/ORNL Center for Molecular Biophysics 2 previous incarnations: Research faculty at UIUC Research scientist at Transtech Pharma, Inc.

Drug Design / drug discovery What’s a drug? A substance that treats/cure a disease. A small molecule that interacts with a target, (often protein involved in the disease process; activator/inhibitor) Drug discovery: The process of finding such a small molecule – combination of approaches Drug discovery or drug design? In principle: “Design” is more rational and targeted, and “discovery” is more serendipitous. But design and discovery share a lot and are ~ synonymous in a pharmaceutical context. 5% of human genome is “druggeable” Hoopkins, Groom, Nat Rev Drug Discov. 2002 1(9): 727 -30.

Drug discovery market Gigantic economic importance: 10 years & $200 to $1, 900 million to develop a drug 25 new molecules /year in millions US$ Revenue R&D income Johnson & Johnson 53, 324 7, 125 11, 053 Pfizer 48, 371 7, 599 19, 337 Glaxo. Smith. Kline 42, 813 6, 373 10, 135 Novartis 37, 020 5, 349 7, 202 Sanofi-Aventis 35, 645 5, 565 5, 033 Hoffmann–La Roche 33, 547 5, 258 7, 318 Astra. Zeneca 26, 475 3, 902 6, 063 Merck & Co. 22, 636 4, 783 4, 434 Abbott Laboratories 22, 476 2, 255 1, 717 Wyeth 20, 351 3, 109 4, 197 > $340 billion http: //en. wikipedia. org/wiki/List_of_pharmaceutical_companies Intense scientific activity: very interdisciplinary approach
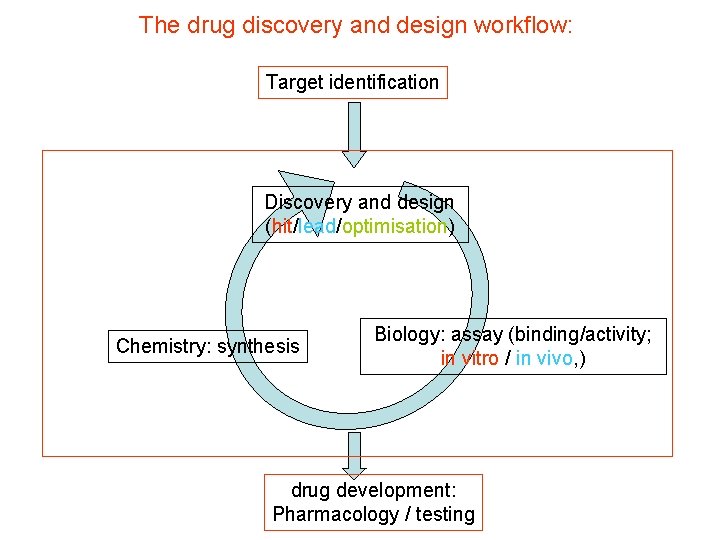
The drug discovery and design workflow: Target identification Discovery and design (hit/lead/optimisation) Chemistry: synthesis Biology: assay (binding/activity; in vitro / in vivo, ) drug development: Pharmacology / testing
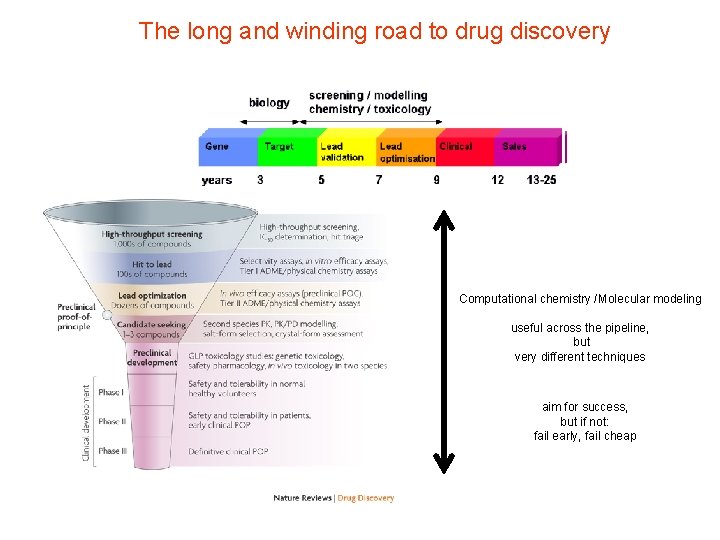
The long and winding road to drug discovery Computational chemistry /Molecular modeling useful across the pipeline, but very different techniques aim for success, but if not: fail early, fail cheap
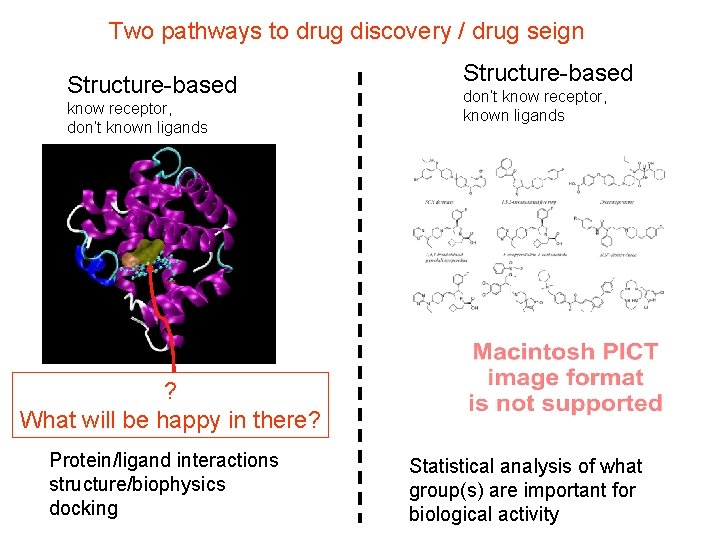
Two pathways to drug discovery / drug seign Structure-based know receptor, don’t known ligands Structure-based don’t know receptor, known ligands ? What will be happy in there? Protein/ligand interactions structure/biophysics docking Statistical analysis of what group(s) are important for biological activity
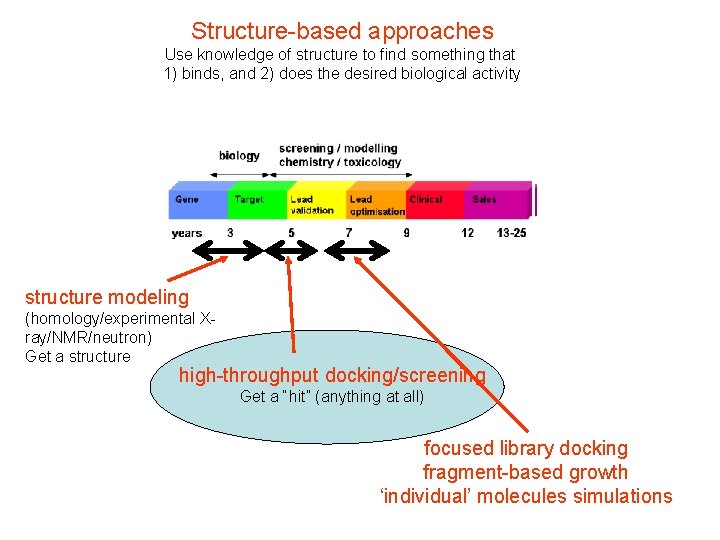
Structure-based approaches Use knowledge of structure to find something that 1) binds, and 2) does the desired biological activity structure modeling (homology/experimental Xray/NMR/neutron) Get a structure high-throughput docking/screening Get a “hit” (anything at all) focused library docking fragment-based growth ‘individual’ molecules simulations

Structure-based library screening What do we need: 1) Compounds libraries 2) Protein target 3) Binding site in the protein 4) Docking: generate different (many) possible conformations of the compounds in the binding site 5) Scoring: evaluate the strength of the protein/ligand interactions (score). 6) Select preferred ligands to propose a list of prioritized compounds for experimental screening.
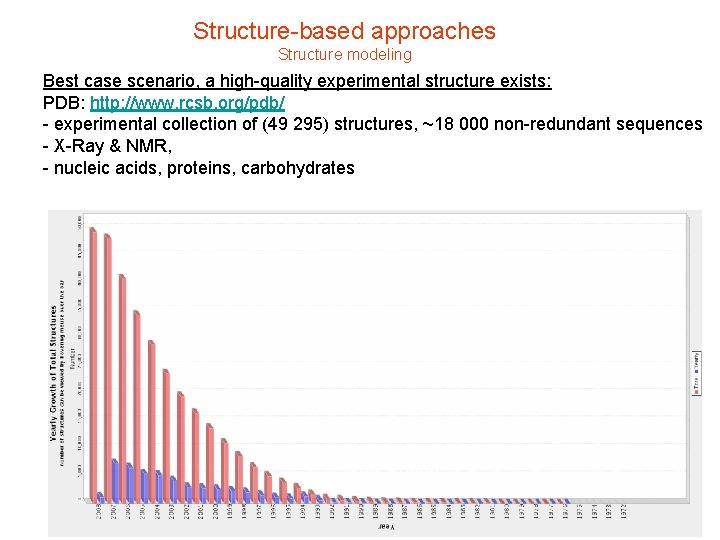
Structure-based approaches Structure modeling Best case scenario, a high-quality experimental structure exists: PDB: http: //www. rcsb. org/pdb/ - experimental collection of (49 295) structures, ~18 000 non-redundant sequences - X-Ray & NMR, - nucleic acids, proteins, carbohydrates

Structure-based approaches Structure modeling ~50, 000 non-redundant protein structures in the PDB: is that a lot? that’s ~1% of the 5. 5 million protein sequences in swissprot (http: //www. ebi. ac. uk/swissprot/sptr_stats/index. html) and < ~0. 00007% of earth’s proteins, (5 E 6 organisms, 5 K genes/genome, low-end estimate. ) …AQRTEVYTYRRS… protein sequence Must do for new pharmaceutical target (homology, ab-initio folding…) protein structure Structure-based drug discovery = “Post genomics challenge”: structural biology, functional genomics, chemical biology…
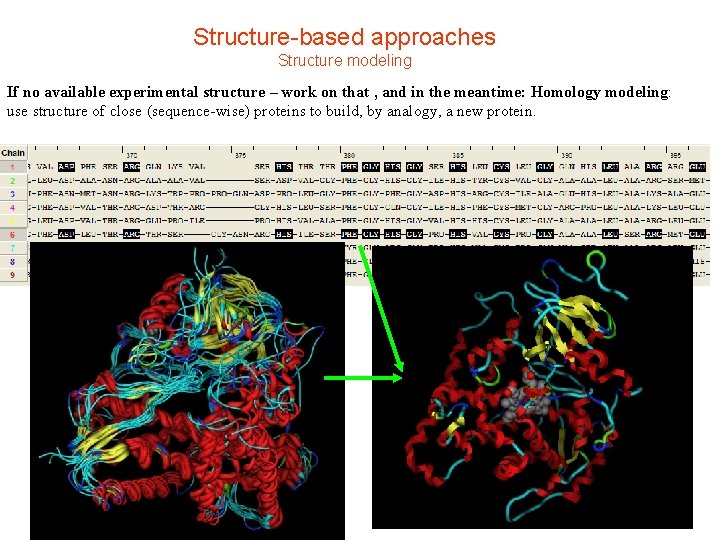
Structure-based approaches Structure modeling If no available experimental structure – work on that , and in the meantime: Homology modeling: use structure of close (sequence-wise) proteins to build, by analogy, a new protein.
Structure-based approaches Compound selection Databases of compounds Library designed against protein target, - vendors - based on hits from previous database - literature screening - corporate/laboratory - virtual compounds -A priori anything, but we can be smarter than that R 1 R 2 R 4 R 3 Millions of cmpds’ structures are available from public databases. Major NIH effort to fund & develop libraries: http: //nihroadmap. nih. gov/molecularlibraries/ http: //blaster. docking. org/zinc/ more exploratory more focused
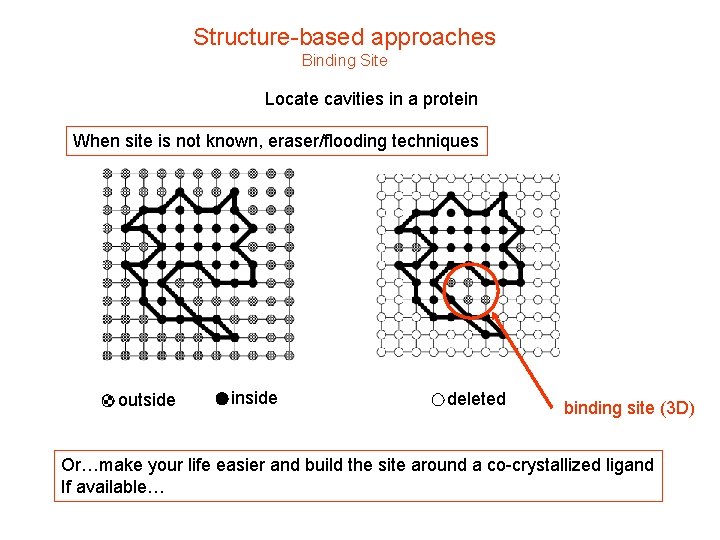
Structure-based approaches Binding Site Locate cavities in a protein When site is not known, eraser/flooding techniques outside inside deleted binding site (3 D) Or…make your life easier and build the site around a co-crystallized ligand If available…
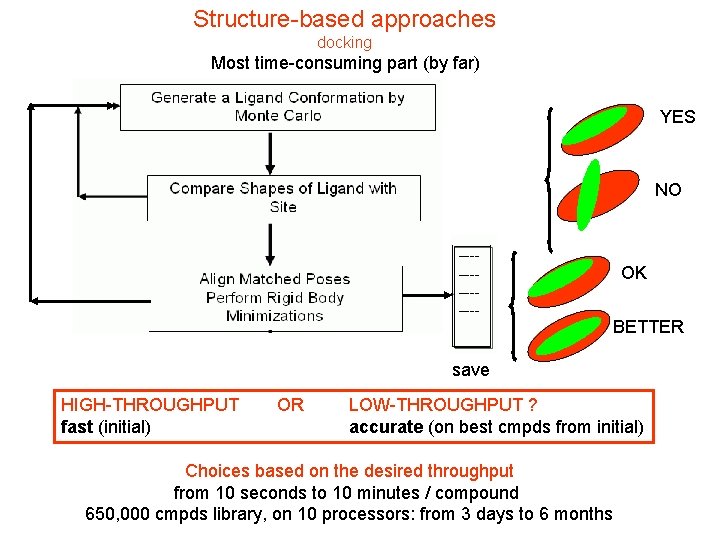
Structure-based approaches docking Most time-consuming part (by far) YES NO OK BETTER save HIGH-THROUGHPUT fast (initial) OR LOW-THROUGHPUT ? accurate (on best cmpds from initial) Choices based on the desired throughput from 10 seconds to 10 minutes / compound 650, 000 cmpds library, on 10 processors: from 3 days to 6 months
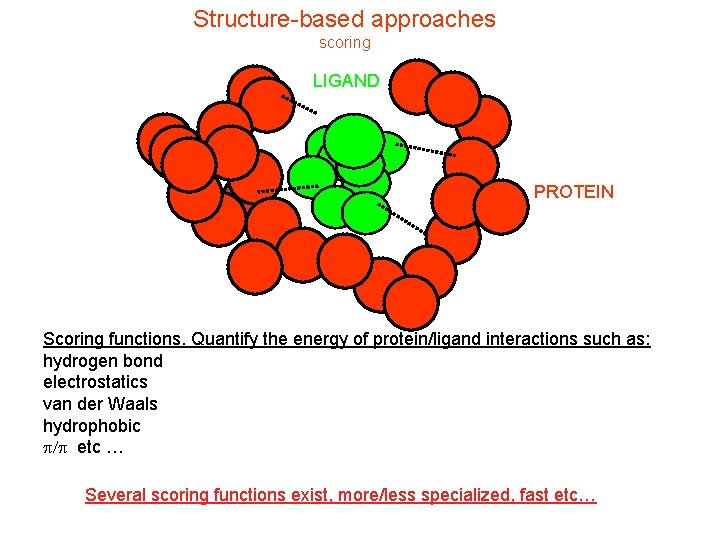
Structure-based approaches scoring LIGAND PROTEIN Scoring functions. Quantify the energy of protein/ligand interactions such as: hydrogen bond electrostatics van der Waals hydrophobic p/p etc … Several scoring functions exist, more/less specialized, fast etc…

scoring functions: Force-field based: (CHARMM, AMBER etc). MMFF: very popular one because of “modular parametrisation”: easy to derive parameters from functional groups, well adapted to organic molecules. Physically ‘accurate’ but slow, parametrisation issues. Empirical – count the number of interactions and assign a score based on the # of occurrences. E. g. : H-bonds, ionic interactions (easy because very directional and well quantified) Hydrophobic interactions (more difficult to assess and quantify) Number of rotatable bonds frozen (link to entropic cost of binding, quite difficult to estimate) Knowledge-based – observe known protein/ligand structures, and favor interactions and geometries that are seen often. Idea: directly link to free energy because “real life” distribution (potential of mean force). But: based on small # of entries. Intense competition “my scoring function is better than yours” Future: force-field based / even QM-based Different approaches depending on size
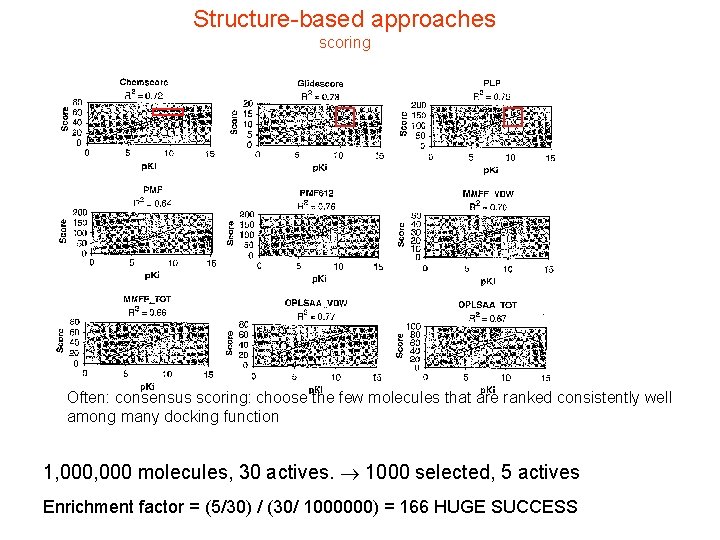
Structure-based approaches scoring Often: consensus scoring: choose the few molecules that are ranked consistently well among many docking function 1, 000 molecules, 30 actives. 1000 selected, 5 actives Enrichment factor = (5/30) / (30/ 1000000) = 166 HUGE SUCCESS
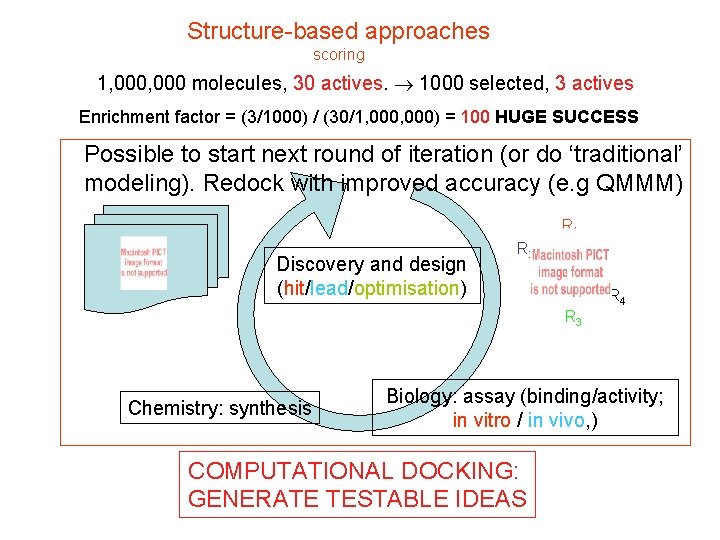
Structure-based approaches scoring 1, 000 molecules, 30 actives. 1000 selected, 3 actives Enrichment factor = (3/1000) / (30/1, 000) = 100 HUGE SUCCESS Possible to start next round of iteration (or do ‘traditional’ modeling). Redock with improved accuracy (e. g QMMM) R 1 Discovery and design (hit/lead/optimisation) R 2 R 4 R 3 Chemistry: synthesis Biology: assay (binding/activity; in vitro / in vivo, ) COMPUTATIONAL DOCKING: GENERATE TESTABLE IDEAS
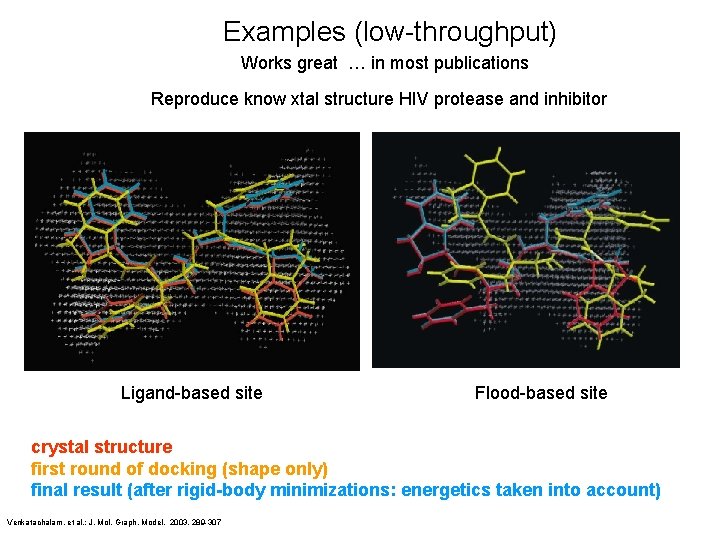
Examples (low-throughput) Works great … in most publications Reproduce know xtal structure HIV protease and inhibitor Ligand-based site Flood-based site crystal structure first round of docking (shape only) final result (after rigid-body minimizations: energetics taken into account) Venkatachalam, et al. ; J. Mol. Graph. Model. 2003, 289 -307
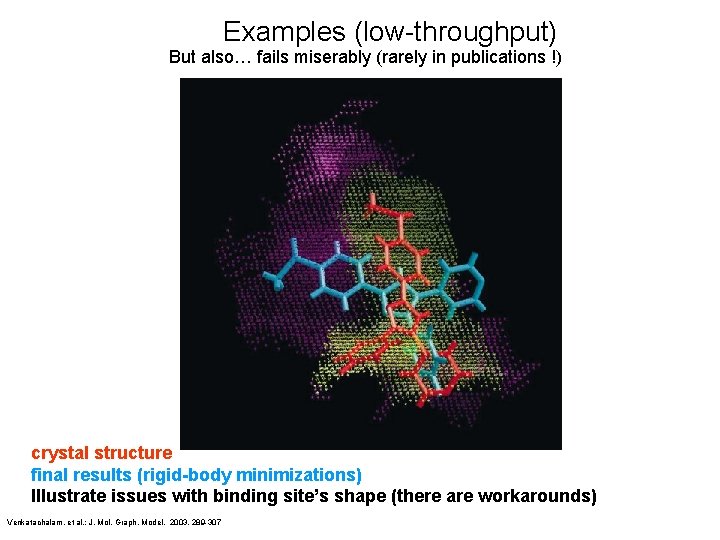
Examples (low-throughput) But also… fails miserably (rarely in publications !) crystal structure final results (rigid-body minimizations) Illustrate issues with binding site’s shape (there are workarounds) Venkatachalam, et al. ; J. Mol. Graph. Model. 2003, 289 -307

Example II): discovery of ligand/function for a new P 450 Ke et al, Archives of Biochemistry and Biophysics 436 (2005) 110– 120
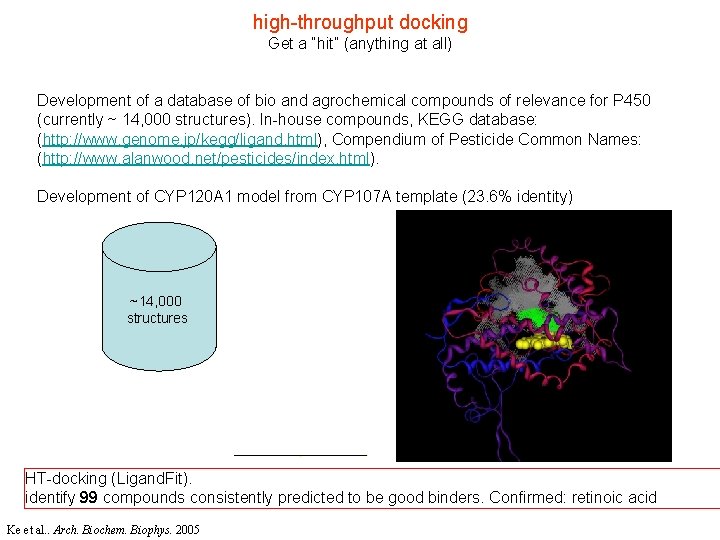
high-throughput docking Get a “hit” (anything at all) Development of a database of bio and agrochemical compounds of relevance for P 450 (currently ~ 14, 000 structures). In-house compounds, KEGG database: (http: //www. genome. jp/kegg/ligand. html), Compendium of Pesticide Common Names: (http: //www. alanwood. net/pesticides/index. html). Development of CYP 120 A 1 model from CYP 107 A template (23. 6% identity) ~14, 000 structures HT-docking (Ligand. Fit). identify 99 compounds consistently predicted to be good binders. Confirmed: retinoic acid Ke et al. . Arch. Biochem. Biophys. 2005

CONCLUSIONS In-silico combinatorial library design & structure-based screening: fast, efficient and inexpensive tool to : discover new possible ligands against a macromolecular target test library design ideas identify most promising scaffolds and R groups prior to synthesis HT-DOCKING SUCCESS IF: i) FIND A FEW MOLECULES OF INTEREST ii) MUCH QUICKER AND CHEAPER THAN “real” screening Baudry, J. ; Hergenrother, P. J. "Structure-based Design and In-Silico Virtual Screening of Combinatorial Libraries. A Combined Chemical/Computational Laboratory Assignment" J. Chem. Ed. 2005, 82, 890 -894. http: //www. scs. uiuc. edu/~phgroup/pdfs/2005 PJHchemed. pdf
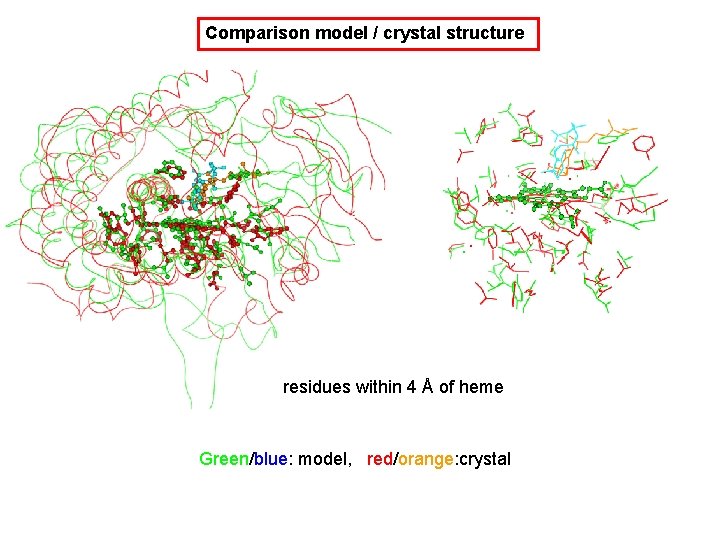
Comparison model / crystal structure residues within 4 Å of heme Green/blue: model, red/orange: crystal
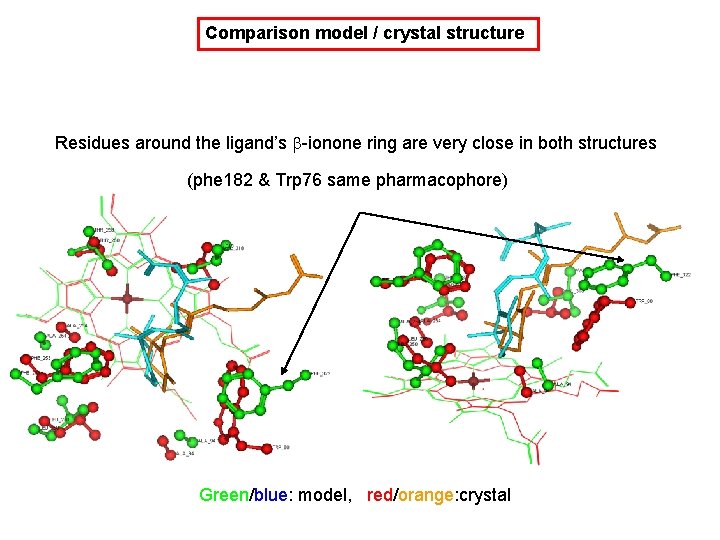
Comparison model / crystal structure Residues around the ligand’s b-ionone ring are very close in both structures (phe 182 & Trp 76 same pharmacophore) Green/blue: model, red/orange: crystal
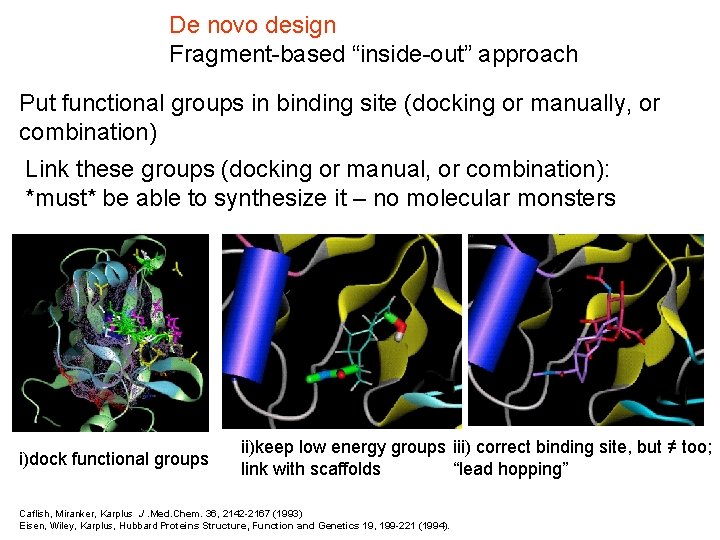
De novo design Fragment-based “inside-out” approach Put functional groups in binding site (docking or manually, or combination) Link these groups (docking or manual, or combination): *must* be able to synthesize it – no molecular monsters i)dock functional groups ii)keep low energy groups iii) correct binding site, but ≠ too; link with scaffolds “lead hopping” Caflish, Miranker, Karplus J. Med. Chem. 36, 2142 -2167 (1993) Eisen, Wiley, Karplus, Hubbard Proteins Structure, Function and Genetics 19, 199 -221 (1994).
- Slides: 26