Dock Crunch and Beyond The future of receptorbased

Dock. Crunch and Beyond. . . The future of receptor-based virtual screening Bohdan Waszkowycz, Tim Perkins & Jin Li Protherics Molecular Design Ltd Macclesfield, UK

Outline Structure-based virtual screening – an achievable (and possibly useful) tool for drug discovery – the Dock. Crunch validation study Protherics’ experience since Dock. Crunch – methods: making VS a routine task – analysis: getting the most from your data – the future (and beyond)
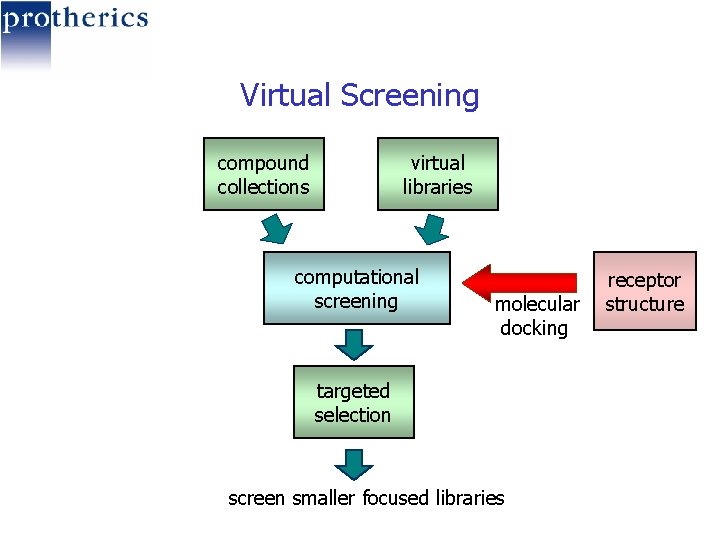
Virtual Screening compound collections virtual libraries computational screening molecular docking targeted selection screen smaller focused libraries receptor structure

Why Use Molecular Docking? Most detailed representation of binding site – overcomes simplifications of pharmacophores – identify both conservative and novel solutions – impetus for de novo design/optimisation Broad range of analyses applicable – diverse scoring/selection criteria Quality/throughput of available methods – good enough, despite technical limitations

Dock. Crunch Validation study for large-scale virtual screening – flexible ligand/rigid receptor docking – PRO_LEADS docking code using Chem. Score scoring function – 1. 1 M druglike ACD-SC compounds – dock versus oestrogen receptor (agonist and antagonist structures) – collaboration with SGI

Oestradiol: Oestrogen Receptor Complex

Docked. Energy Profiles Agonist Receptor Antagonist receptor • Achieve good separation in terms of predicted binding affinity

Dock. Crunch Results Demonstrated technical feasibility – 1. 1 M cpds docked in 6 days/64 processor Origin – implemented automated pre- and post-processing Demonstrated potential for lead identification – – successful discrimination of seeded known hits activity for 21 out of 37 assayed compounds ER binding affinities to 7 n. M Ki novel non-steroidal chemistries

Since Dock. Crunch. . . VS established as a routine CAMD task: – 2. 2 M structures docked in Dock. Crunch – 1. 5 M docked versus in-house target – 2. 5 M docked to date in external contracts – – – project 1: 0. 25 M Dec 2000 project 2: 0. 25 M Jan 2001 project 3: 1 M Feb 2001 project 4: 1 M March-April 2001 project 5: 0. 5 M to do in May. . . diverse targets/databases/project objectives

Virtual Screening within Prometheus Commercial databases Database preparation e. g. salt removal, protonation Database pre-filtering select drug-like profile Receptor structure Receptor-ligand docking predict binding mode/affinity Analysis graphical browsing, subset selection Virtual databases

PRO_LEADS Docking Tabu search + extended Chem. Score function – robust prediction of binding free energy – 85% success rate achieved across diverse test set Pre-calculated grids for energies/neighbour lists – defines extent of binding site – automatically/graphically defined Selection of PRO_LEADS docking protocol – use standard protocol across all receptors – specific constraints or modified energy terms available if desired

Example of Grid Definition c. AMP-dependent kinase (1 YDS) contact surface coloured by lipophilicity

Docking Throughput Standard protocols take 1– 5 mins/ligand – e. g. typical VS run at ~4 min for 3 M tabu steps – 250 k cpds/week on 100 processor Linux cluster (VA Linux 750 MHz PIII) PLUNDER script for parallelization – automatic processing of ligand batches – balances processor workload – works across heterogeneous architectures – supplies running time statistics – handles hardware failures

Data Analysis and Subset Selection Intrinsic problems of scoring functions: – – cannot parameterize all critical interactions try to take account of induced fit effects calibrated only versus good binders ignore co-operativity in binding When applied to random datasets: – predicted affinity typically normal distributed – overestimates binding affinity of random set energy alone not ideal for subset selection

Achieving Better Selection Need to supplement scoring function – consensus scoring schemes Explore more fundamental descriptors of receptor: ligand complementarity – capture characteristics of diverse receptor types – assess deficiencies of existing scoring functions – use as simple filters or as pseudo energy terms

Enrichment Rates Effect of different selection criteria for ER set for recovery of seeded compounds

Requirements for Analysis Package VS generates huge data output – want to be able to browse through entire dataset Real-time navigation of large datasets – – – graphing property distributions selections based on property filters browsing of 3 D models within selections initiating additional property calculations data transformations writing subset/reports

Property. Viewer

Approach to Analysis 1. Preliminary exploration – browse property distributions – comparisons with known ligands 2. Initial elimination of poor structures – – Docked. Energy, component energies DE corrected for size/functionality receptor: ligand steric complementarity polar/lipophilic surface complementarity

Approach to Analysis 3. Further filtering define focused subsets – tighter 2 D property filters – clustering by 2 D chemistry – presence of key 3 D binding interactions – specific H-bonds, specific lipo contacts, occupancy, volume overlap with reference ligand/fragment, etc – similarity/diversity of 3 D binding mode – 3 D similarity descriptors – final ranking by Docked. Energy or hybrid energy/complementarity scoring function pocket

Docked. Energy vs Size

Complementarity Space ER and FXa datasets

Addressing More Difficult Cases - COX 2 Knowns show clustering in property space despite modest Docked. Energy

Improvements in Docking Function original docking function some misdocked knowns new docking function more consistent docking +ve shift in random energies

Comparison of filters in subset selection 87% pass 2 D filters 37% pass energy filters 43% 22% 1% 22% pass complementarity filters 0% filtering to ~10% energy filters complementarity 2 D properties 2% 12% 9% Initial – – – Selection of final ~1% subset – 3 D structural features – preferred binding motifs – 2 D/3 D diversity

Conclusions Established VS as a routine CAMD task – focused software development – achieved success in drug discovery projects VS is more than a black box – data mining is worthwhile – explore receptor-ligand complementarity to achieve good subset selection and point towards better scoring functions

Future Directions for VS Exploit expanding computing resource – improved docking/scoring functions – improved receptor representations Broader application of VS – – evaluation of drugability of early targets screening of very large virtual libraries routine screening across protein families DMPK issues

Acknowledgements Tim Perkins Richard Sykes Richard Hall David Frenkel David Sheppard Martin Harrison Carol Baxter Chris Murray Jin Li Thanks to: SGI, MSI, MDL, VA Linux http: //www. protherics. com/crunch/
- Slides: 28