Do TS Genes From Transcribed Sequences to Genes

Do. TS Genes – From Transcribed Sequences to Genes via Genomic Alignment Y. Thomas Gan*#, Jonathan Crabtree*#, Joan Mazzarelli*#, Otto Valladares#, Maja Bucan#, and Christian Stoeckert*# * CBIL, Center for Bioinformatics, U. of Penn. , Phila. , PA # Department of Genetics, U. of Penn. , Phila. , PA

Background Although sequences for large eukaryotic genomes are being completed, it remains a challenge to identify all genes encoded by them. Approaches used to date include ab initio gene prediction, similarity-based annotation, and direct alignment of transcribed sequences. Here we demonstrate a process that uses a variant of the direct alignment approach. We align Do. TS Transcripts, instead of individual ESTs and m. RNAs, against genomes using BLAT (Kent WJ, 2002) and merge selected alignments to identify genes. Do. TS is a database of human/mouse transcribed sequences built by cleaning up, clustering and assembling the millions of ESTs (and RNAs) from Gen. Bank.

Result Summary n Do. TS Genes n n n ~ 47, 000 w/ high confidence for human ~ 42, 000 w/ high confidence for mouse Access of Do. TS Genes: http: //www. allgenes. org also accessible from n n n Human-mouse orthologs n n ~ 22, 000 human-mouse Reciprocal Best Hits Novel Gene Candidates 100+ w/ high confidence in ~75 Mb mouse chromosome 5 proximal region, not in the ENSEMBL or the Celera set. Candidates for experimental verification. Gene model enrichment of known genes n additional exons n extended UTRs n n UCSC custom tracks: http: //genome. ucsc. edu/golden. Path/custom. Tracks/cust. Tracks. html ENSEMBL genome browser: http: //www. ensembl. org (DAS Sources menu)

Method Overview n n n Cluster and Assemble EST/RNA into Do. TS Transcripts Align Do. TS Transcripts to Genome Load alignments into GUS Database Compute alignment “quality” Merge selected alignments into “genes” Assign confidence scores
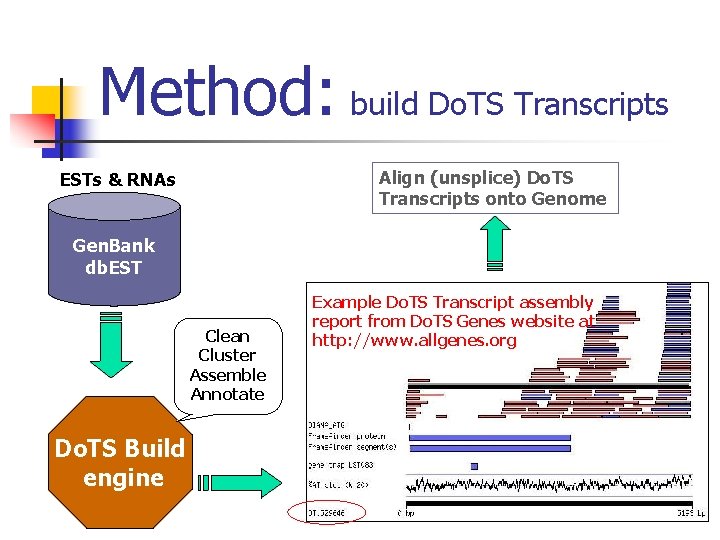
Method: build Do. TS Transcripts Align (unsplice) Do. TS Transcripts onto Genome ESTs & RNAs Gen. Bank db. EST Clean Cluster Assemble Annotate Do. TS Build engine Example Do. TS Transcript assembly report from Do. TS Genes website at http: //www. allgenes. org

Method: alignment quality n 1 = Very good n n n >= 95% identity (average) max_query_gap <= 5 both ends consistent n n n same as very good but internal and end mismatches allowed if there is a sufficiently large genomic sequence gap (within 10 X mismatch length for ends) 3 = Good n n not poly. A on both ends 2 = Very good with genomic gaps n n no more than 10 bp mismatch unless poly. A same as very good, but with max_query_gap <= 15, and inconsistent ends allowed if unaligned_bases <= 50 4 = Others

Method: “Gene” Creation n Select BLAT alignments n n Merge alignments w/ exon overlap, same strand Merge alignments w/ common EST clone info n n Distance: within 500 kb Merge nearby alignments n n quality: “good” & better Less than “good”, but best in the whole genome Parameter: within 75 bp (all merges are transitive)

Method: filter & score n High confidence Do. TS Genes (w/ stable identifiers) n n n contain m. RNA or spliced (w/ intron of at least 15 bp) cross-validated w/ our results of independent pre-genome approach Do. TS Genes are also scored based on n Gene structure and contents n n n n Genomic context n n splice: number of exons Alternative splice: number of Do. TS Transcripts m. RNA presence distinct EST libraries and/or clones 5’-3’ EST pairs from the same clone distribution of 5’/3’ EST end positions splice signal poly. A track Conservation: human/mouse reciprocal best hits
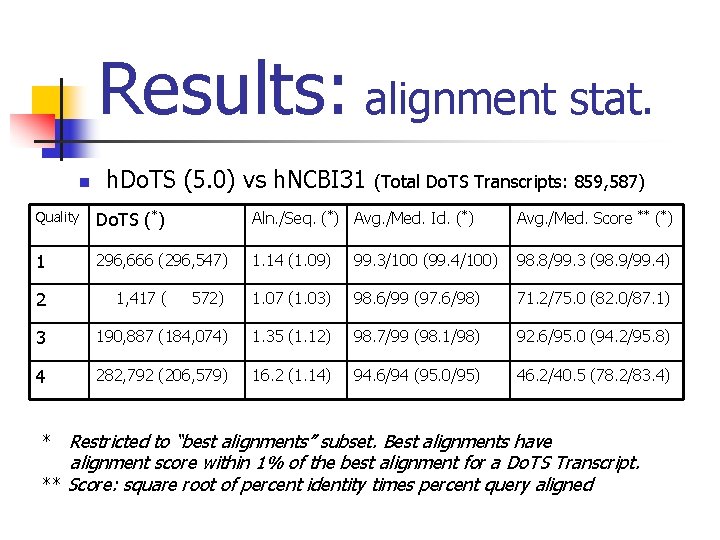
Results: alignment stat. n h. Do. TS (5. 0) vs h. NCBI 31 (Total Do. TS Transcripts: 859, 587) Quality Do. TS (*) Aln. /Seq. (*) Avg. /Med. Id. (*) Avg. /Med. Score 1 296, 666 (296, 547) 1. 14 (1. 09) 99. 3/100 (99. 4/100) 98. 8/99. 3 (98. 9/99. 4) 572) 1. 07 (1. 03) 98. 6/99 (97. 6/98) 71. 2/75. 0 (82. 0/87. 1) 3 190, 887 (184, 074) 1. 35 (1. 12) 98. 7/99 (98. 1/98) 92. 6/95. 0 (94. 2/95. 8) 4 282, 792 (206, 579) 16. 2 (1. 14) 94. 6/94 (95. 0/95) 46. 2/40. 5 (78. 2/83. 4) 2 1, 417 ( * Restricted to “best alignments” subset. Best alignments have ** alignment score within 1% of the best alignment for a Do. TS Transcript. ** Score: square root of percent identity times percent query aligned (*)

Results: human Do. TS Genes n n n h. Do. TS (5. 0) vs m. NCBI 31 48, 066 w/ high confidence 5. 8 transcripts / gene on average 6. 1 exons / gene on average 16, 950 contain m. RNA
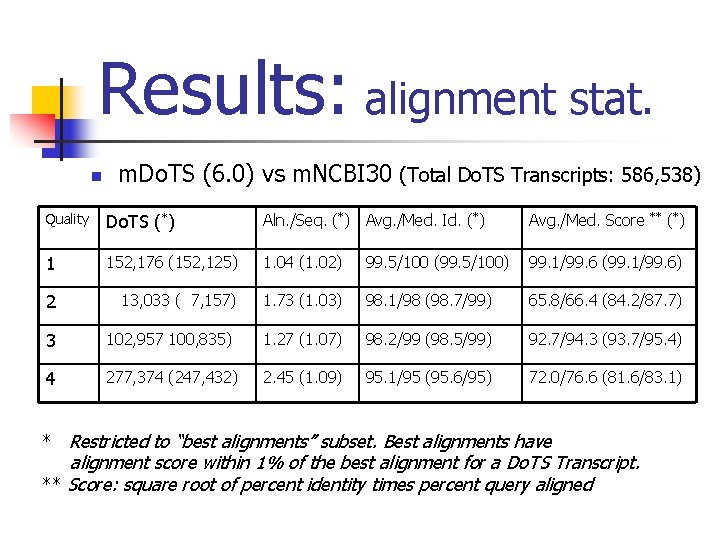
Results: alignment stat. n m. Do. TS (6. 0) vs m. NCBI 30 (Total Do. TS Transcripts: 586, 538) Quality Do. TS (*) Aln. /Seq. (*) Avg. /Med. Id. (*) Avg. /Med. Score 1 152, 176 (152, 125) 1. 04 (1. 02) 99. 5/100 (99. 5/100) 99. 1/99. 6 (99. 1/99. 6) 2 13, 033 ( 7, 157) 1. 73 (1. 03) 98. 1/98 (98. 7/99) 65. 8/66. 4 (84. 2/87. 7) ** (*) 3 102, 957 100, 835) 1. 27 (1. 07) 98. 2/99 (98. 5/99) 92. 7/94. 3 (93. 7/95. 4) 4 277, 374 (247, 432) 2. 45 (1. 09) 95. 1/95 (95. 6/95) 72. 0/76. 6 (81. 6/83. 1) * Restricted to “best alignments” subset. Best alignments have alignment score within 1% of the best alignment for a Do. TS Transcript. ** Score: square root of percent identity times percent query aligned

Results: mouse Do. TS Genes n n n n m. Do. TS (6. 0) vs m. NCBI 30 42, 090 w/ high confidence 5. 4 transcripts / gene on average 6. 1 exons / gene on average 11, 715 contain m. RNA 17, 146 contain at least 2 splice signals 16, 654 have poly. A signals

Do. TS Gene result in good agreement w/ that of an independent approach n Compared with our pre-genome method: n n similarity-based graph algorithm genomic sequence info not used 86% (79%) human (mouse) Do. TS Genes have unique mappings Given a high confidence Do. TS Gene uniquely mapped to similarity-based cluster, on average 65% transcripts of the former are in common with 90% transcripts of the latter (for median, it is 62% vs 100%)
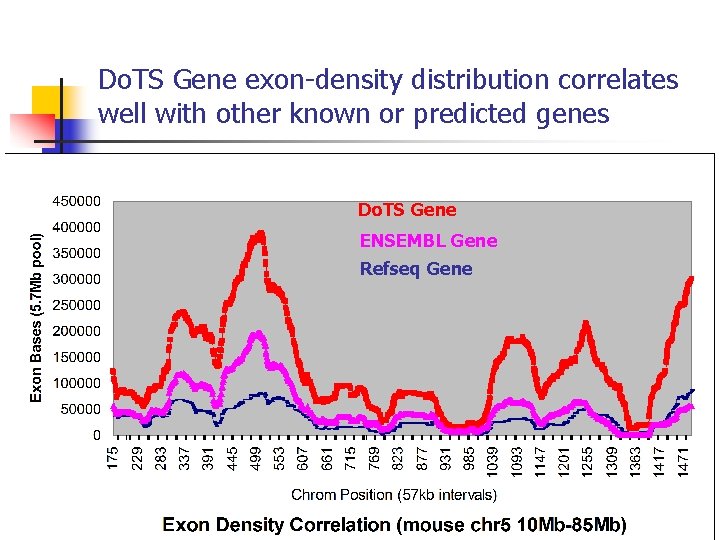
Do. TS Gene exon-density distribution correlates well with other known or predicted genes Do. TS Gene ENSEMBL Gene Refseq Gene
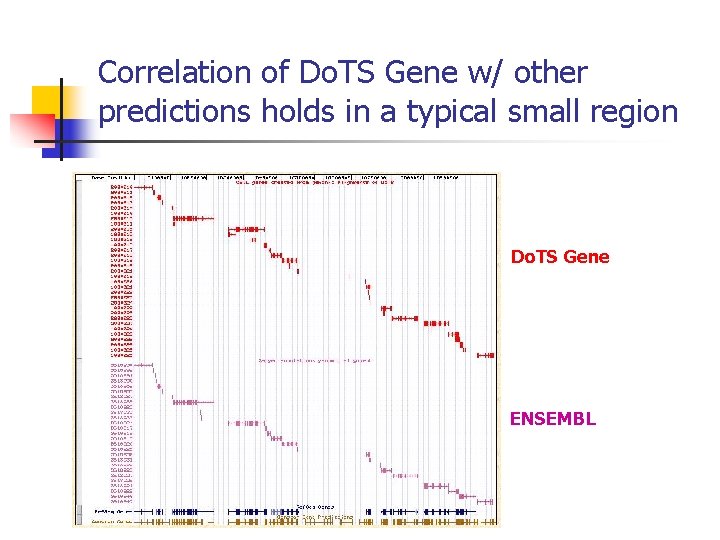
Correlation of Do. TS Gene w/ other predictions holds in a typical small region Do. TS Gene ENSEMBL

Do. TS Genes predict novel gene candidates n n n Examined in detail mouse chr 5 proximal ~75 Mb region Do. TS Gene predicted significantly more genes than ENSEMBL/Celera combined ~150 with highest scores are being experimentally verified

Do. TS Genes extend UTRs of known genes

Do. TS Genes add/extend internal exons of known genes

Do. TS Genes suggest alternative transcription starts of known genes n n Example w/ experimental evidence (Zhu Y. et. al. , Genome Biology 2003, 4: R 16) Example shown below

Further Directions n n n Continue efforts in experimental verification of predicted novel genes in mouse chromosome 5 proximal region Incorporate new sequence data (transcribed and genomic) and experiment feedback to fine-tune Do. TS Gene identification / scoring processes Enhance gene models and annotations of Do. TS Genes
- Slides: 20