DNA Topoisomerases DNA Supercoiling in vivo In most




![Processive activity at higher [ATP] Magnet rotation applied Topo II activity • Trelax << Processive activity at higher [ATP] Magnet rotation applied Topo II activity • Trelax <<](https://slidetodoc.com/presentation_image_h/a8f9643655f690e96bf449fc56862d39/image-12.jpg)
![[ATP] and force-dependence of strand passage Km = 270 m. M ATP Vsat = [ATP] and force-dependence of strand passage Km = 270 m. M ATP Vsat =](https://slidetodoc.com/presentation_image_h/a8f9643655f690e96bf449fc56862d39/image-15.jpg)



- Slides: 34

DNA Topoisomerases

DNA Supercoiling in vivo • In most organisms, DNA is negatively supercoiled (s ~ -0. 06) • Supercoiling is involved in initiation of transcription, replication, repair & recombination • Actively regulated by topoisomerases, ubiquitous and essential family of proteins

Chromosomes: the ultimate Gordian knot? EM by U. Laemmli

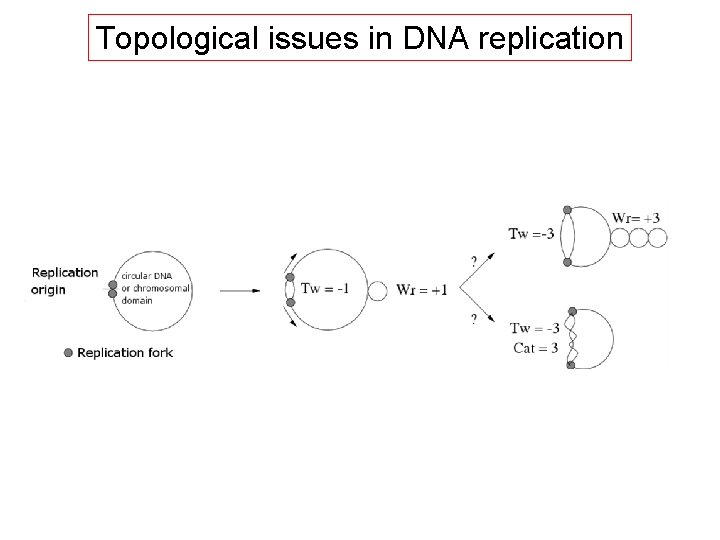
Topological issues in DNA replication

Supercoiling and transcription • In bacteria, gyrase helps maintain negative supercoiling. • This can help drive transcription in many genes (although gyrase is, itself, downregulated by negative supercoiling). • Mutations in gyrase are compensated by mutations in topo I to prevent it from removing negative supercoiling. • Positive supercoils ahead of RNAP, negative supercoils behind?

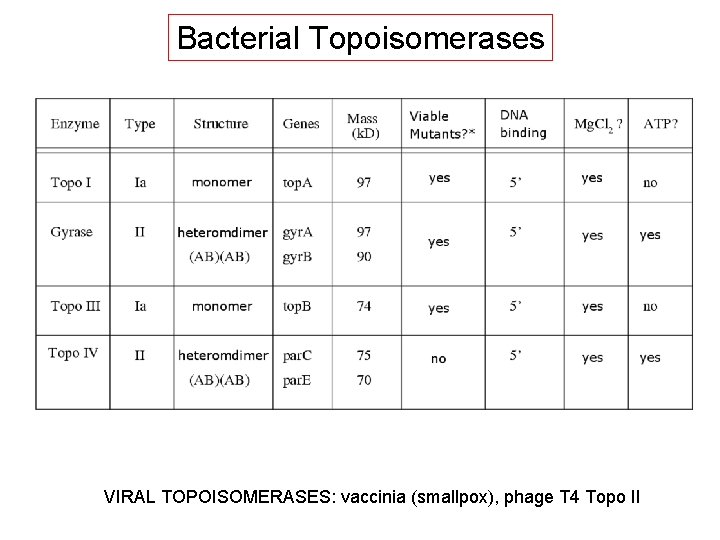
Bacterial Topoisomerases VIRAL TOPOISOMERASES: vaccinia (smallpox), phage T 4 Topo II

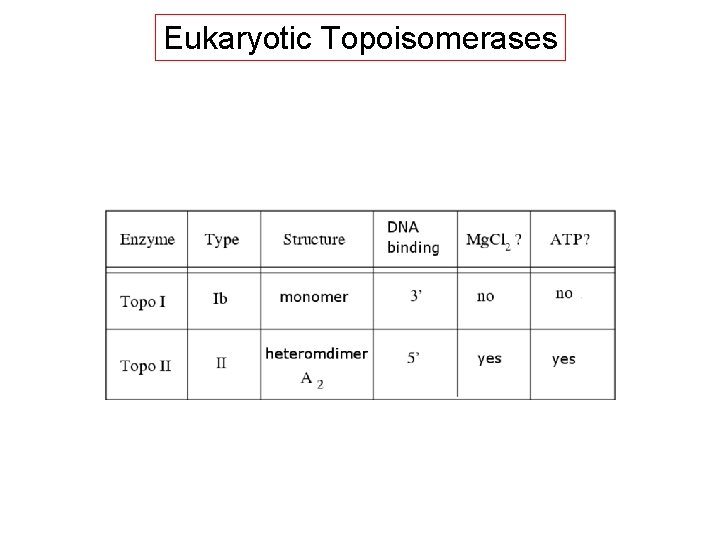
Eukaryotic Topoisomerases

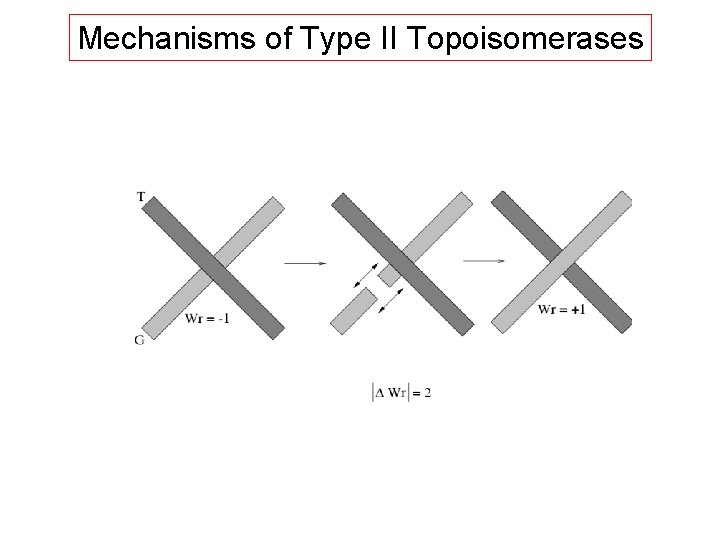
Mechanisms of Type II Topoisomerases

Therapeutic Implications Gyrase is a good target for antibacterial quinolones (ciproflaxin). DNA Breakages are toxic… Reversed by tyrosyl-DNA phosphodiesterases (3’ topo Ib breaks)… How are tdp proteins and other break-repairing proteins (involved in recombinational repair) involved in resistance to chimiotherapeutic agents? Topoisomerase II poisons are used in chemotherapy (daunorubicin, doxorubicin, etoposide) as well as Topo I poisons (topotecan)

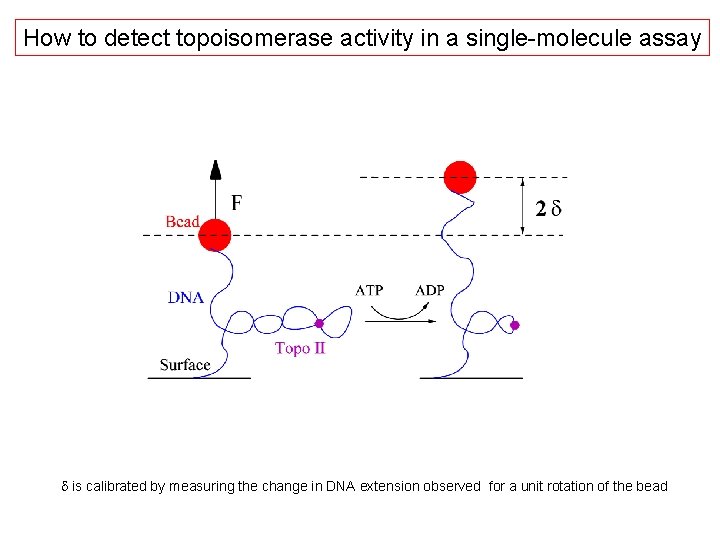
How to detect topoisomerase activity in a single-molecule assay d is calibrated by measuring the change in DNA extension observed for a unit rotation of the bead

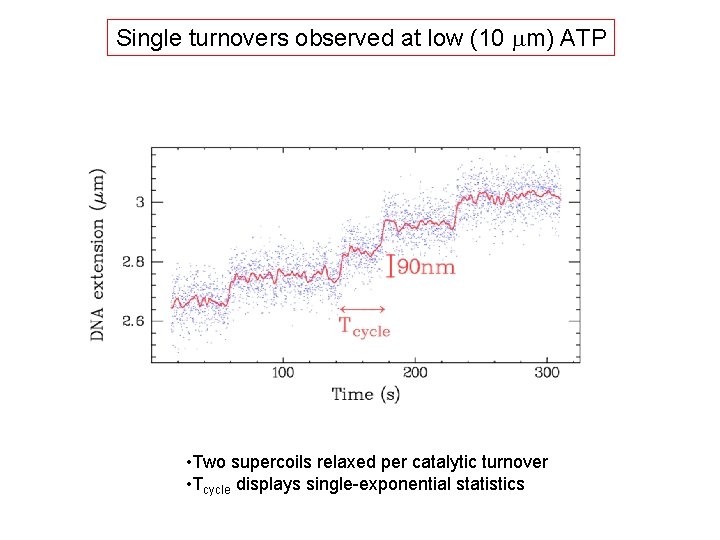
Single turnovers observed at low (10 mm) ATP • Two supercoils relaxed per catalytic turnover • Tcycle displays single-exponential statistics
![Processive activity at higher ATP Magnet rotation applied Topo II activity Trelax Processive activity at higher [ATP] Magnet rotation applied Topo II activity • Trelax <<](https://slidetodoc.com/presentation_image_h/a8f9643655f690e96bf449fc56862d39/image-12.jpg)
Processive activity at higher [ATP] Magnet rotation applied Topo II activity • Trelax << Twait single molecule bursts • Processivity on the order of ten cycles

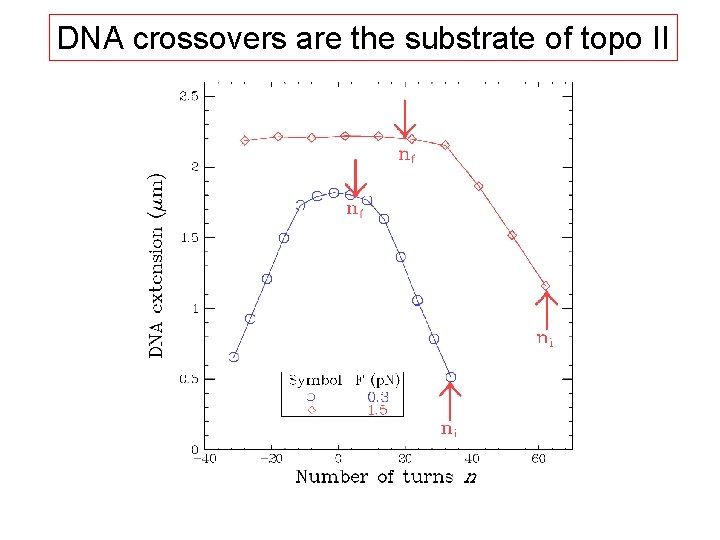
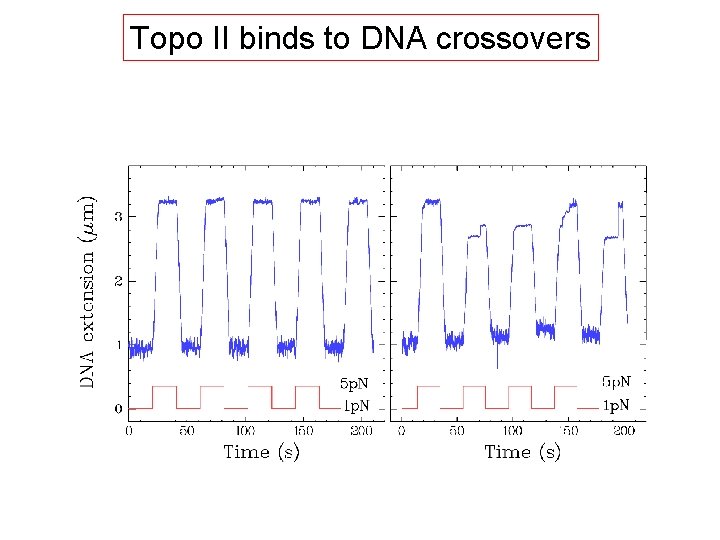
DNA crossovers are the substrate of topo II

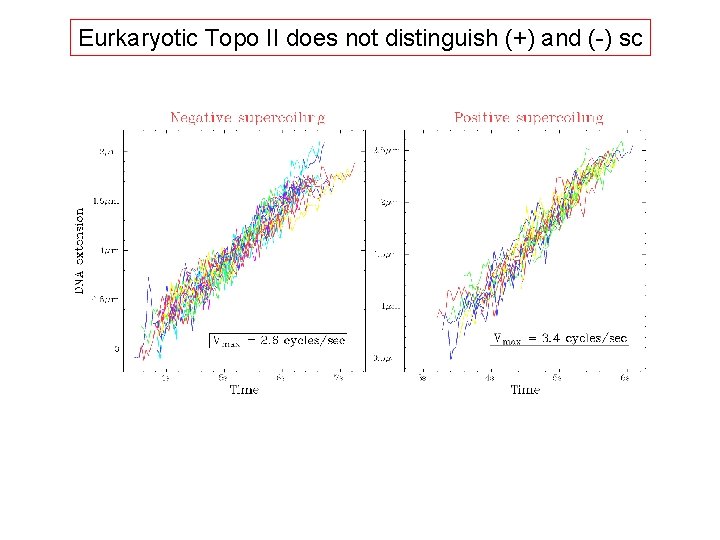
Eurkaryotic Topo II does not distinguish (+) and (-) sc
![ATP and forcedependence of strand passage Km 270 m M ATP Vsat [ATP] and force-dependence of strand passage Km = 270 m. M ATP Vsat =](https://slidetodoc.com/presentation_image_h/a8f9643655f690e96bf449fc56862d39/image-15.jpg)
[ATP] and force-dependence of strand passage Km = 270 m. M ATP Vsat = 3 cycles/sec • Rate-limiting step coupled to ~1 nm motion against the applied force

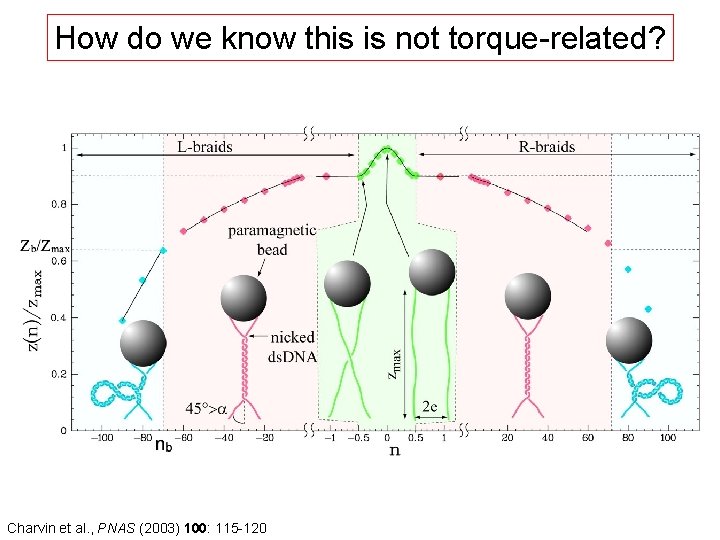
How do we know this is not torque-related? Charvin et al. , PNAS (2003) 100: 115 -120

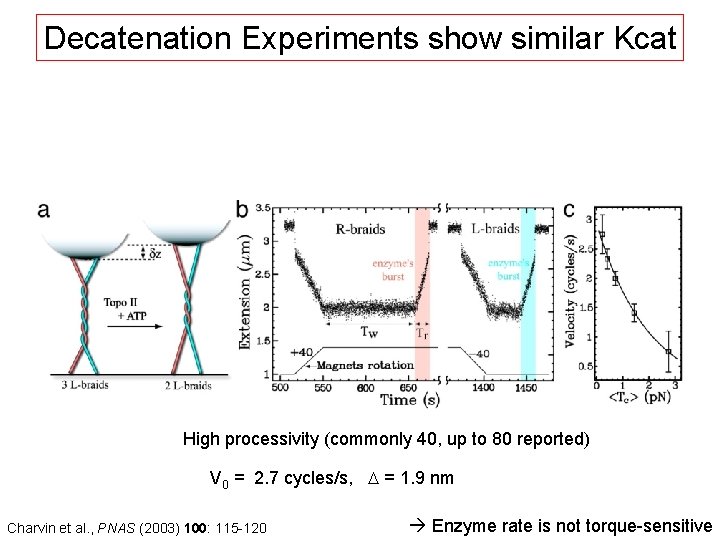
Decatenation Experiments show similar Kcat High processivity (commonly 40, up to 80 reported) V 0 = 2. 7 cycles/s, D = 1. 9 nm Charvin et al. , PNAS (2003) 100: 115 -120 Enzyme rate is not torque-sensitive

Model: closure of the DNA gap is rate-limiting

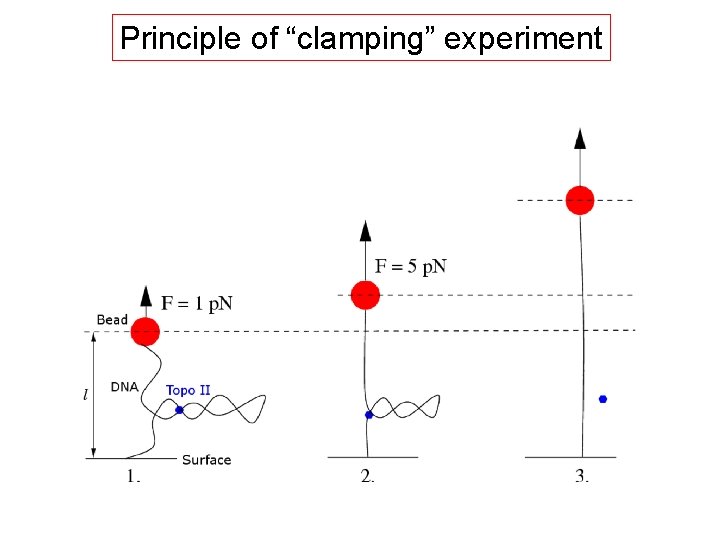
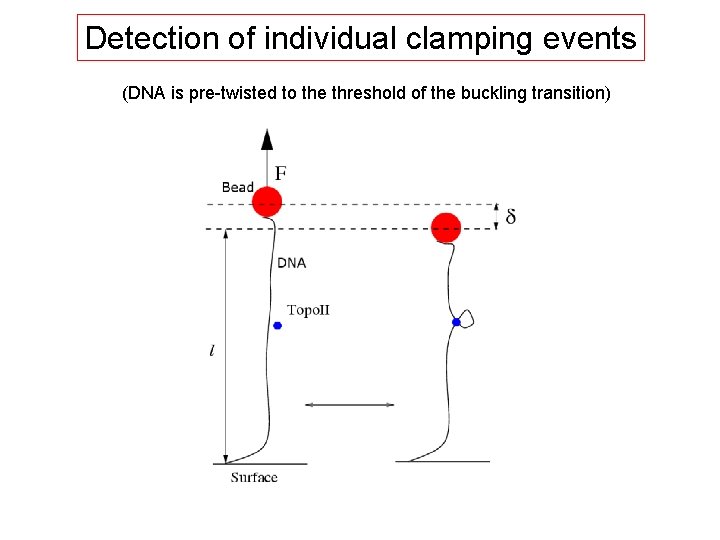
Principle of “clamping” experiment

Topo II binds to DNA crossovers

Detection of individual clamping events (DNA is pre-twisted to the threshold of the buckling transition)

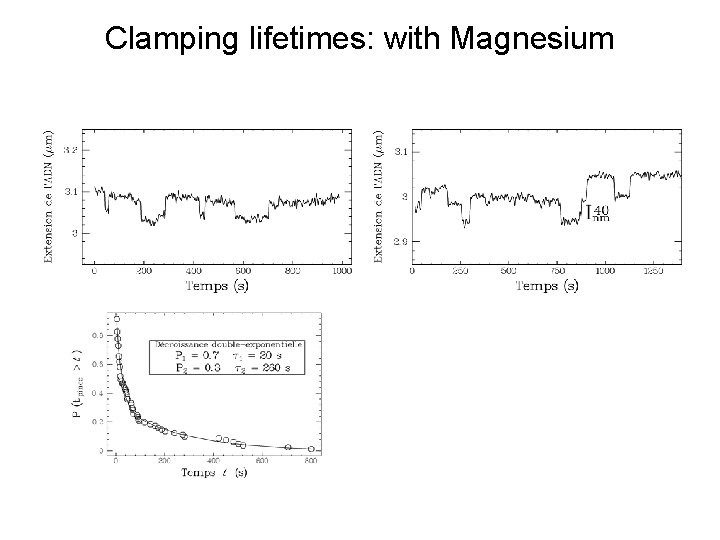
Clamping lifetimes: with Magnesium

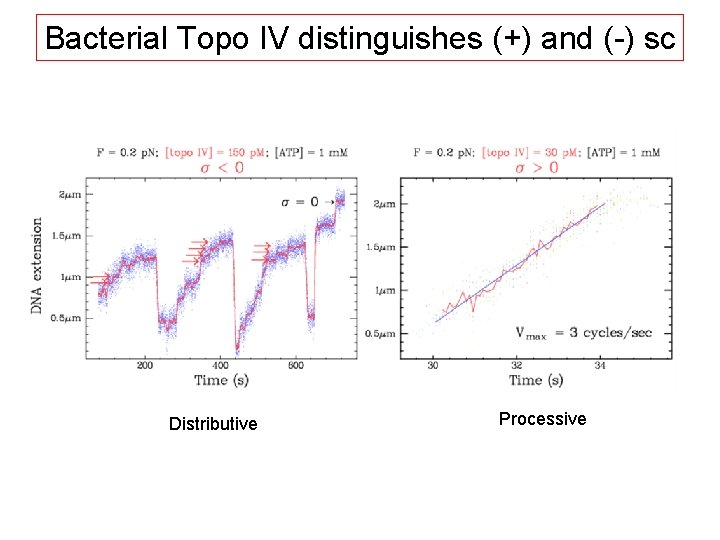
Bacterial Topo IV distinguishes (+) and (-) sc Distributive Processive

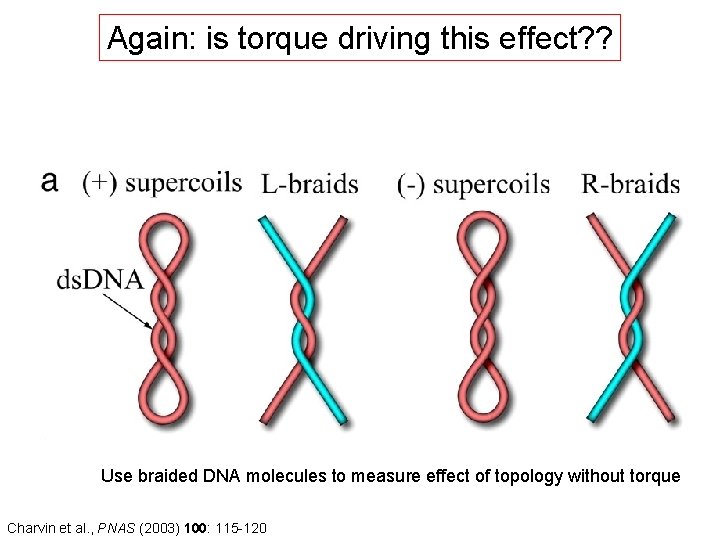
Again: is torque driving this effect? ? Use braided DNA molecules to measure effect of topology without torque Charvin et al. , PNAS (2003) 100: 115 -120

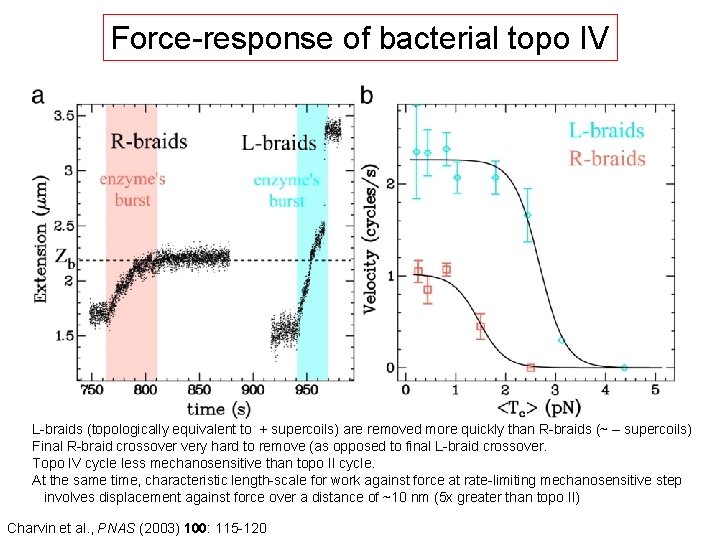
Force-response of bacterial topo IV L-braids (topologically equivalent to + supercoils) are removed more quickly than R-braids (~ – supercoils) Final R-braid crossover very hard to remove (as opposed to final L-braid crossover. Topo IV cycle less mechanosensitive than topo II cycle. At the same time, characteristic length-scale for work against force at rate-limiting mechanosensitive step involves displacement against force over a distance of ~10 nm (5 x greater than topo II) Charvin et al. , PNAS (2003) 100: 115 -120

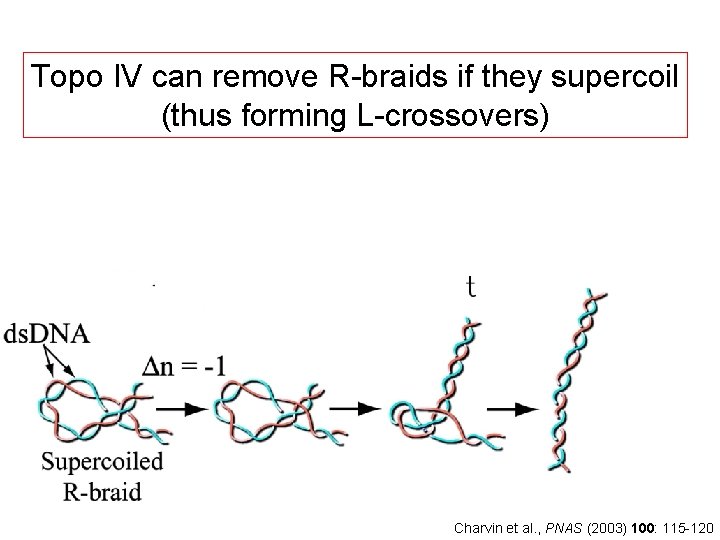
Topo IV can remove R-braids if they supercoil (thus forming L-crossovers) Charvin et al. , PNAS (2003) 100: 115 -120

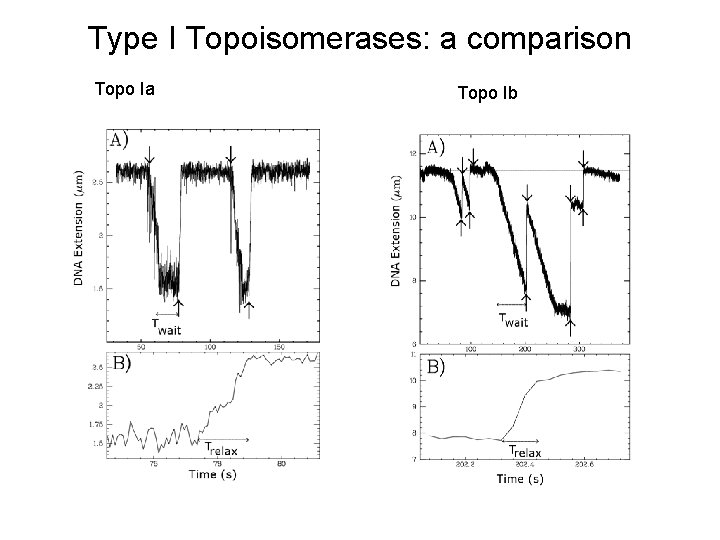
Type I Topoisomerases: a comparison Topo Ia Topo Ib

Measuring step-size by variance analysis Random (lt) ___n exp(-lt) P(n) = n! 1. X(t) is the recorded position of the system 2. Record many (long) traces and average them together mean = <X> = NPD variance = < (X - < X >)2 > = NP(1 -P)D 2

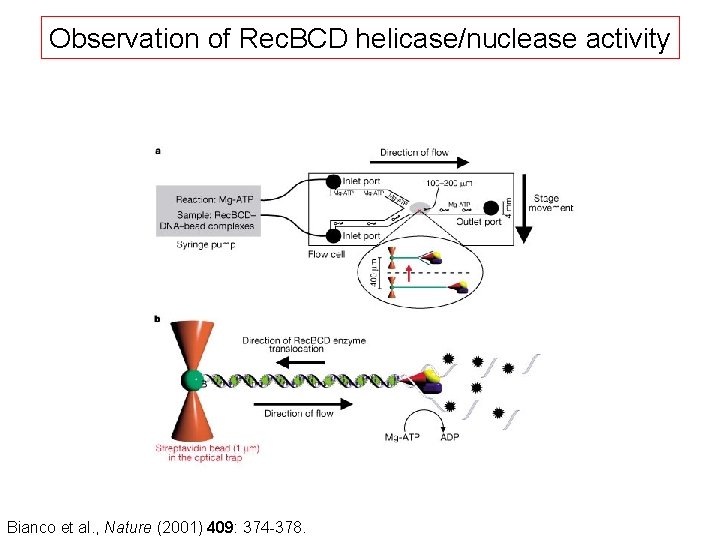
Observation of Rec. BCD helicase/nuclease activity Bianco et al. , Nature (2001) 409: 374 -378.

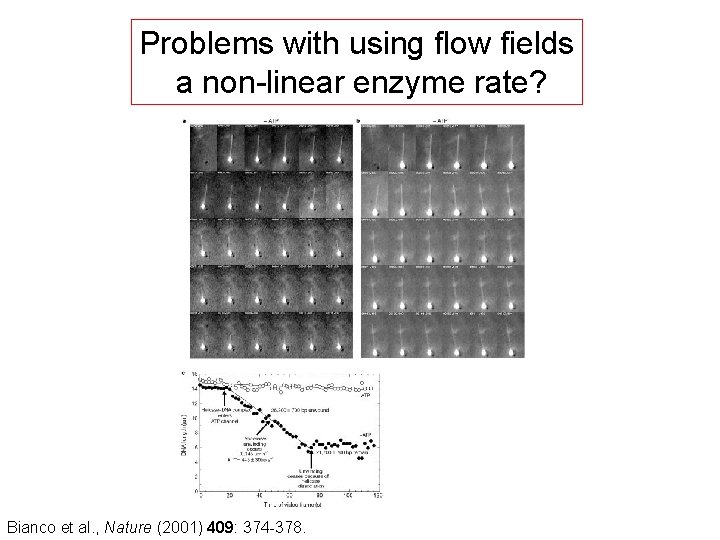
Problems with using flow fields a non-linear enzyme rate? Bianco et al. , Nature (2001) 409: 374 -378.

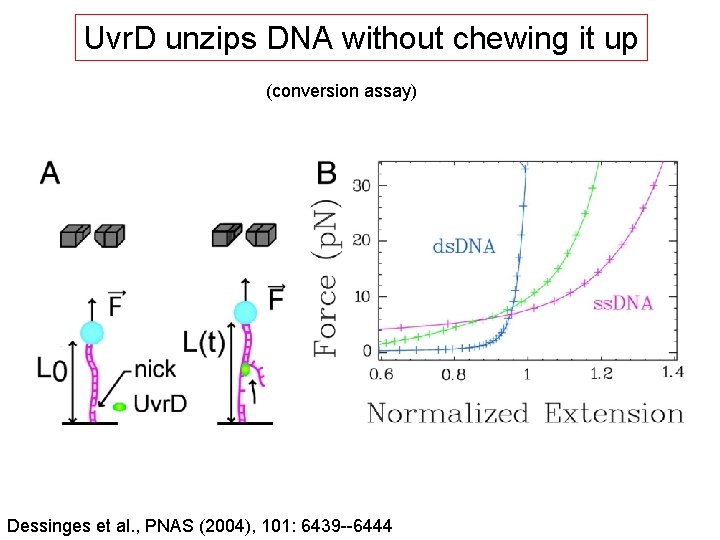
Uvr. D unzips DNA without chewing it up (conversion assay) Dessinges et al. , PNAS (2004), 101: 6439 --6444

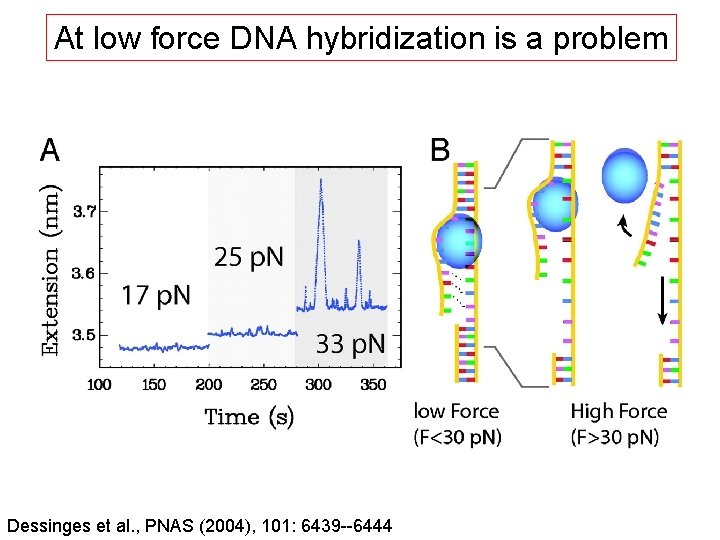
At low force DNA hybridization is a problem Dessinges et al. , PNAS (2004), 101: 6439 --6444

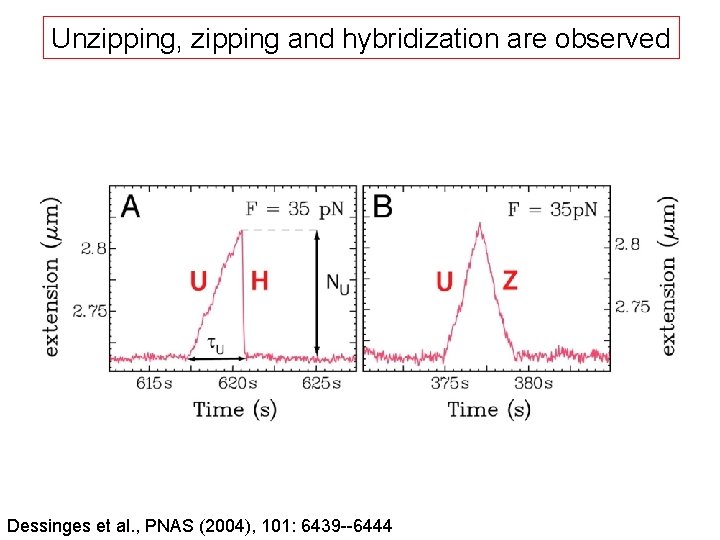
Unzipping, zipping and hybridization are observed Dessinges et al. , PNAS (2004), 101: 6439 --6444

Measuring step-size by variance analysis Like a random walk: N steps with a probability P (small) of moving forward a distance D Repeat the walk a large number of times and average the results together mean distance travelled = NPD variance of distance travelled = NP(1 -P)D 2 variance mean =D