DNA Sequencing DNA sequencing How we obtain the
DNA Sequencing
DNA sequencing How we obtain the sequence of nucleotides of a species …ACGTGACTGAGGACCGTG CGACTGACTGGGT CTAGACTACGTTTTA TATATACGTCGTCGT ACTGATGACTAGATTACAG ACTGATTTAGATACCTGAC TGATTTTAAAAAAATATT… CS 273 a Lecture 3, Spring 07, Batzoglou

Which representative of the species? Which human? Answer one: Answer two: it doesn’t matter Polymorphism rate: number of letter changes between two different members of a species Humans: ~1/1, 000 Other organisms have much higher polymorphism rates § Population size! CS 273 a Lecture 3, Spring 07, Batzoglou
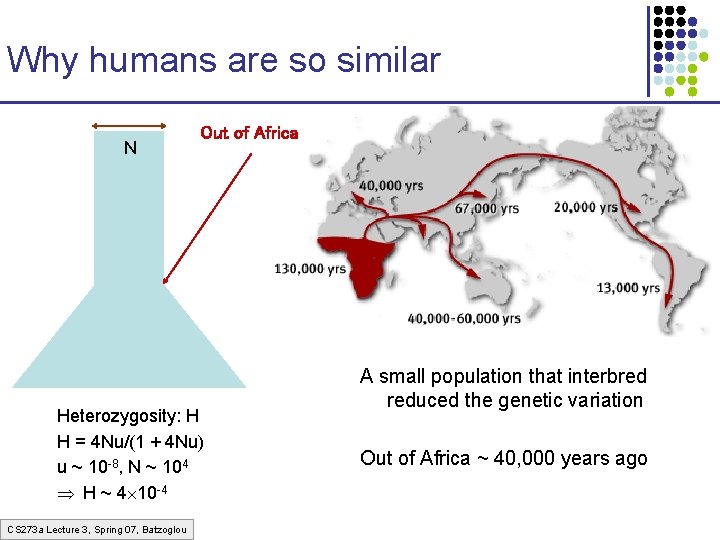
Why humans are so similar N Out of Africa Heterozygosity: H H = 4 Nu/(1 + 4 Nu) u ~ 10 -8, N ~ 104 H ~ 4 10 -4 CS 273 a Lecture 3, Spring 07, Batzoglou A small population that interbred reduced the genetic variation Out of Africa ~ 40, 000 years ago

Human population migrations • Out of Africa, Replacement § “Grandma” of all humans (Eve) ~150, 000 yr • Ancestor of all mt. DNA § “Grandpa” of all humans (Adam) ~100, 000 yr • Ancestor of all Y-chromosomes • Multiregional Evolution § Fossil records show a continuous change of morphological features § Proponents of theory doubt mt. DNA and other genetic evidence CS 273 a Lecture 3, Spring 07, Batzoglou
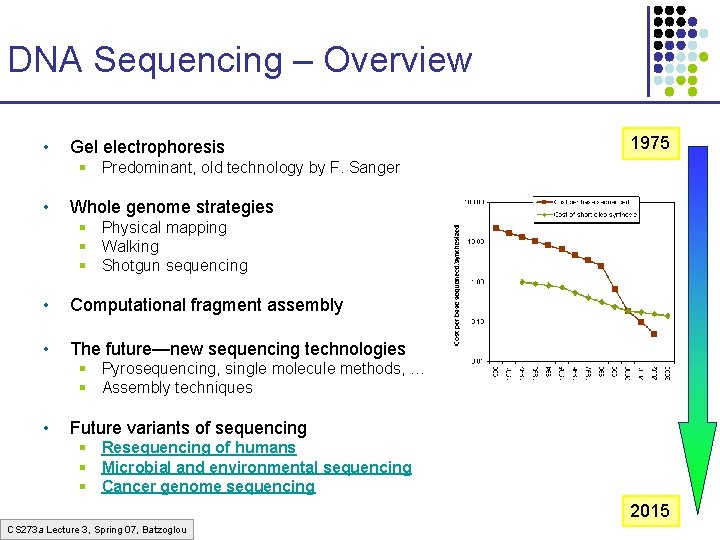
DNA Sequencing – Overview • Gel electrophoresis 1975 § Predominant, old technology by F. Sanger • Whole genome strategies § Physical mapping § Walking § Shotgun sequencing • Computational fragment assembly • The future—new sequencing technologies § Pyrosequencing, single molecule methods, … § Assembly techniques • Future variants of sequencing § Resequencing of humans § Microbial and environmental sequencing § Cancer genome sequencing 2015 CS 273 a Lecture 3, Spring 07, Batzoglou

DNA Sequencing Goal: Find the complete sequence of A, C, G, T’s in DNA Challenge: There is no machine that takes long DNA as an input, and gives the complete sequence as output Can only sequence ~800 letters at a time CS 273 a Lecture 3, Spring 07, Batzoglou
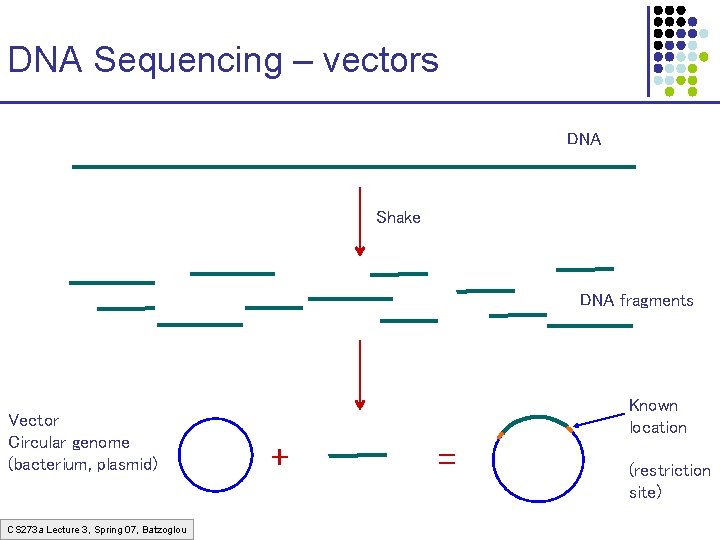
DNA Sequencing – vectors DNA Shake DNA fragments Vector Circular genome (bacterium, plasmid) CS 273 a Lecture 3, Spring 07, Batzoglou Known location + = (restriction site)

Different types of vectors VECTOR Size of insert Plasmid 2, 000 -10, 000 Can control the size Cosmid 40, 000 BAC (Bacterial Artificial Chromosome) 70, 000 -300, 000 YAC (Yeast Artificial Chromosome) > 300, 000 Not used much recently CS 273 a Lecture 3, Spring 07, Batzoglou
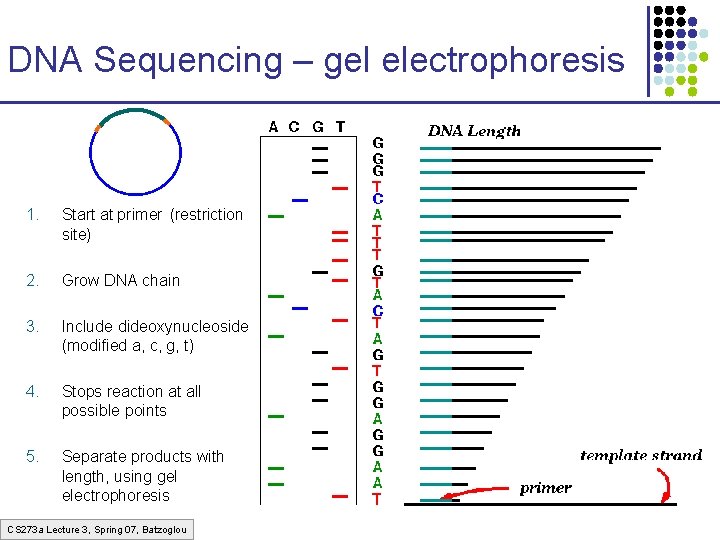
DNA Sequencing – gel electrophoresis 1. Start at primer (restriction site) 2. Grow DNA chain 3. Include dideoxynucleoside (modified a, c, g, t) 4. Stops reaction at all possible points 5. Separate products with length, using gel electrophoresis CS 273 a Lecture 3, Spring 07, Batzoglou
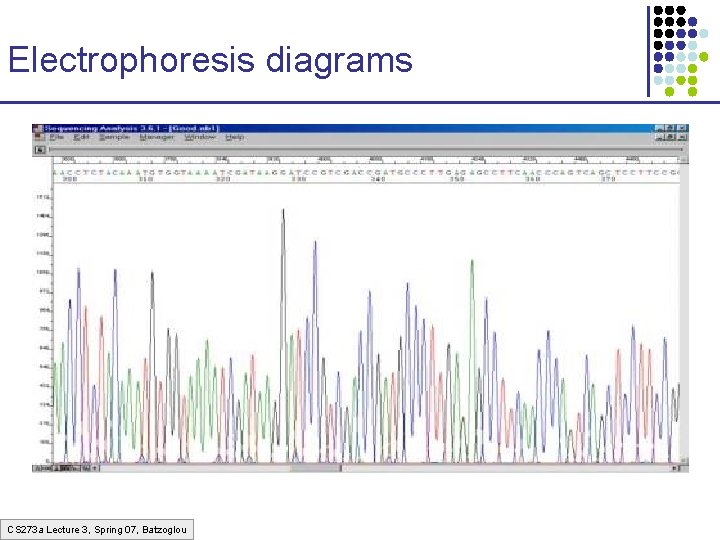
Electrophoresis diagrams CS 273 a Lecture 3, Spring 07, Batzoglou

Reading an electropherogram 1. 2. 3. 4. Filtering Smoothening Correction for length compressions A method for calling the letters – PHRED – PHil’s Read EDitor (by Phil Green) Newer methods may be better, but labs are reluctant to change CS 273 a Lecture 3, Spring 07, Batzoglou

Output of PHRED: a read A read: 500 -1000 nucleotides A C G A A T C A G …A 16 18 21 23 25 15 28 30 32 … 21 Quality scores: -10 log 10 Prob(Error) Reads can be obtained from leftmost, rightmost ends of the insert Double-barreled sequencing: (1990) Both leftmost & rightmost ends are sequenced, reads are paired CS 273 a Lecture 3, Spring 07, Batzoglou
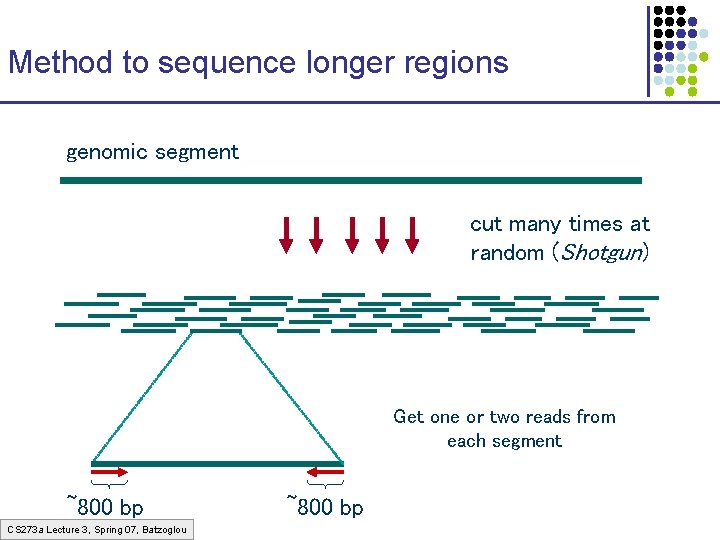
Method to sequence longer regions genomic segment cut many times at random (Shotgun) Get one or two reads from each segment ~800 bp CS 273 a Lecture 3, Spring 07, Batzoglou ~800 bp
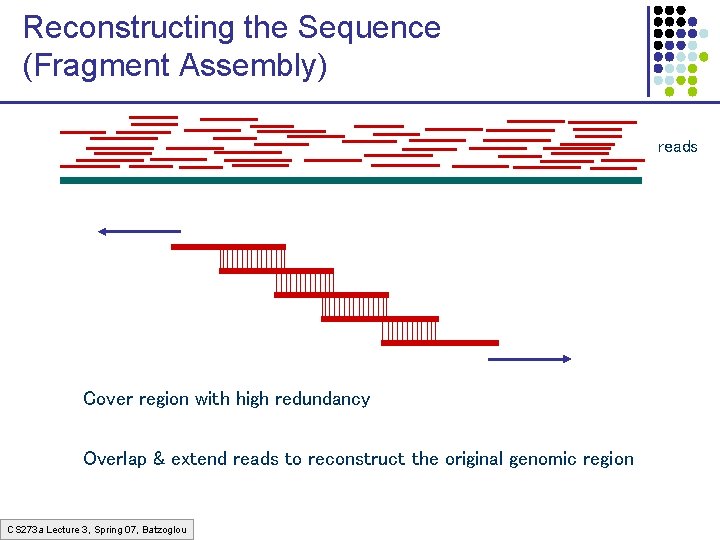
Reconstructing the Sequence (Fragment Assembly) reads Cover region with high redundancy Overlap & extend reads to reconstruct the original genomic region CS 273 a Lecture 3, Spring 07, Batzoglou

Definition of Coverage C Length of genomic segment: Number of reads: Length of each read: L n l Definition: C=nl/L Coverage How much coverage is enough? Lander-Waterman model: Assuming uniform distribution of reads, C=10 results in 1 gapped region /1, 000 nucleotides CS 273 a Lecture 3, Spring 07, Batzoglou

Repeats Bacterial genomes: Mammals: 5% 50% Repeat types: • Low-Complexity DNA (e. g. ATATACATA…) • Microsatellite repeats • Transposons § SINE (a 1…ak)N where k ~ 3 -6 (e. g. CAGCAGTAGCAGCACCAG) (Short Interspersed Nuclear Elements) e. g. , ALU: ~300 -long, 106 copies § LINE § LTR retroposons (Long Interspersed Nuclear Elements) ~4000 -long, 200, 000 copies (Long Terminal Repeats (~700 bp) at each end) cousins of HIV • Gene Families genes duplicate & then diverge (paralogs) • Recent duplications ~100, 000 -long, very similar copies CS 273 a Lecture 3, Spring 07, Batzoglou
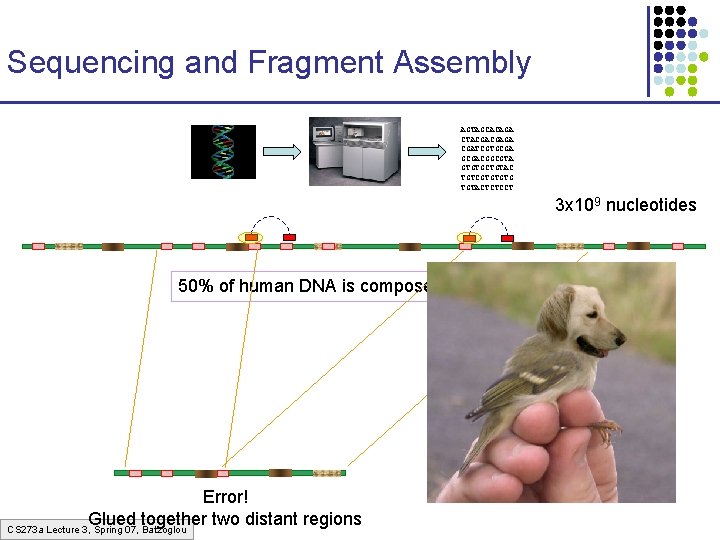
Sequencing and Fragment Assembly AGTAGCACAGA CTACGACGAGA CGATCGTGCGACGGCGTA GTGTGCTGTAC TGTCGTGTGTG TGTACTCTCCT 3 x 109 nucleotides 50% of human DNA is composed of repeats Error! Glued together two distant regions CS 273 a Lecture 3, Spring 07, Batzoglou
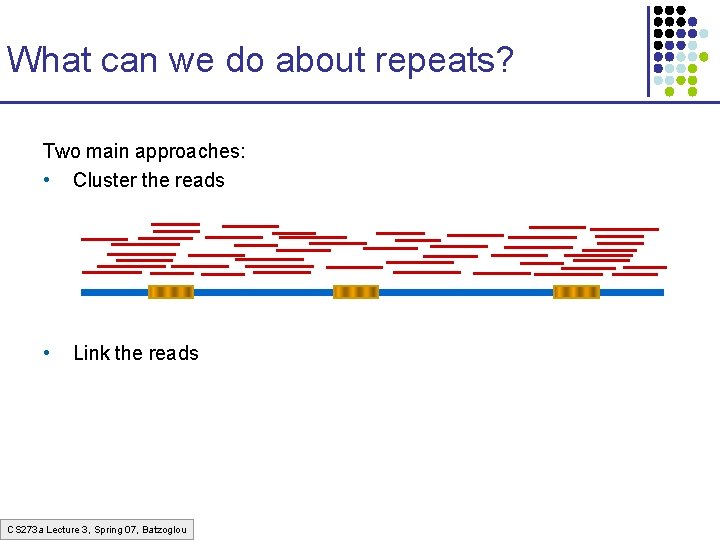
What can we do about repeats? Two main approaches: • Cluster the reads • Link the reads CS 273 a Lecture 3, Spring 07, Batzoglou

What can we do about repeats? Two main approaches: • Cluster the reads • Link the reads CS 273 a Lecture 3, Spring 07, Batzoglou
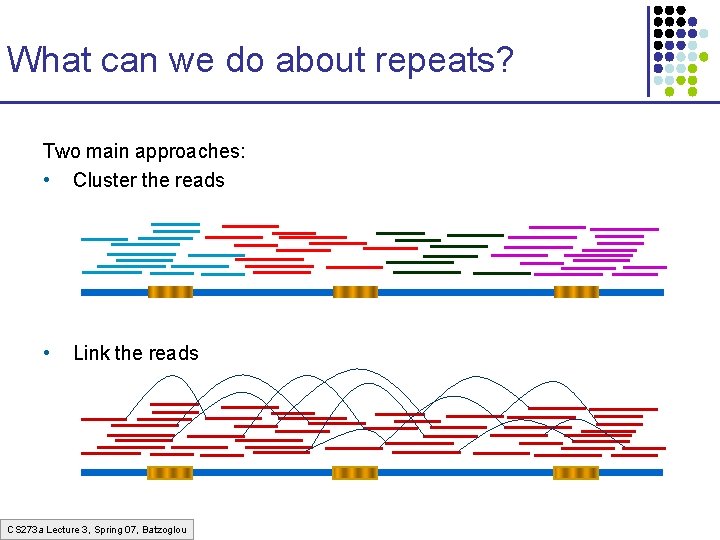
What can we do about repeats? Two main approaches: • Cluster the reads • Link the reads CS 273 a Lecture 3, Spring 07, Batzoglou
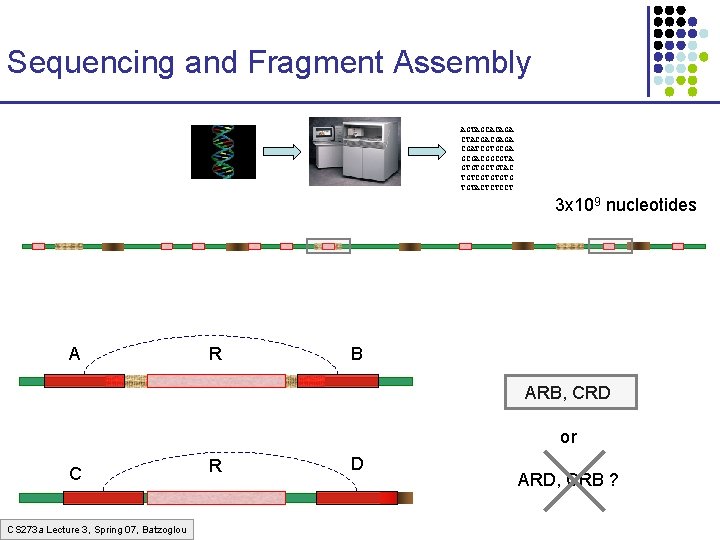
Sequencing and Fragment Assembly AGTAGCACAGA CTACGACGAGA CGATCGTGCGACGGCGTA GTGTGCTGTAC TGTCGTGTGTG TGTACTCTCCT 3 x 109 nucleotides A R B ARB, CRD or C CS 273 a Lecture 3, Spring 07, Batzoglou R D ARD, CRB ?
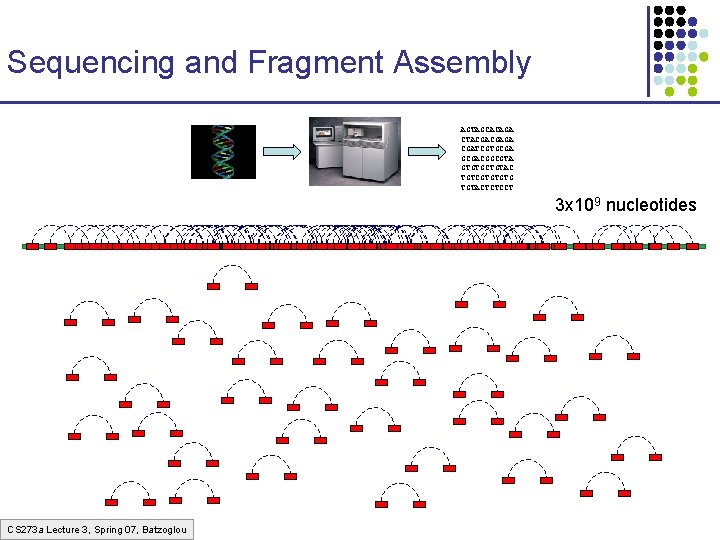
Sequencing and Fragment Assembly AGTAGCACAGA CTACGACGAGA CGATCGTGCGACGGCGTA GTGTGCTGTAC TGTCGTGTGTG TGTACTCTCCT 3 x 109 nucleotides CS 273 a Lecture 3, Spring 07, Batzoglou
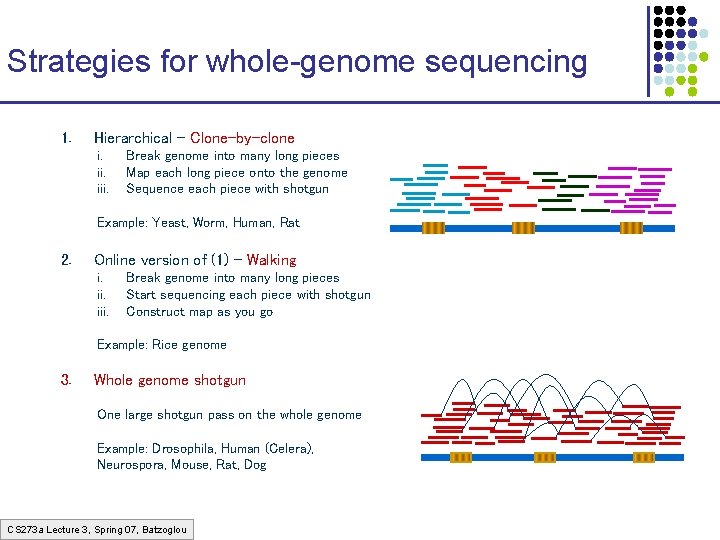
Strategies for whole-genome sequencing 1. Hierarchical – Clone-by-clone i. iii. Break genome into many long pieces Map each long piece onto the genome Sequence each piece with shotgun Example: Yeast, Worm, Human, Rat 2. Online version of (1) – Walking i. iii. Break genome into many long pieces Start sequencing each piece with shotgun Construct map as you go Example: Rice genome 3. Whole genome shotgun One large shotgun pass on the whole genome Example: Drosophila, Human (Celera), Neurospora, Mouse, Rat, Dog CS 273 a Lecture 3, Spring 07, Batzoglou

Hierarchical Sequencing CS 273 a Lecture 3, Spring 07, Batzoglou
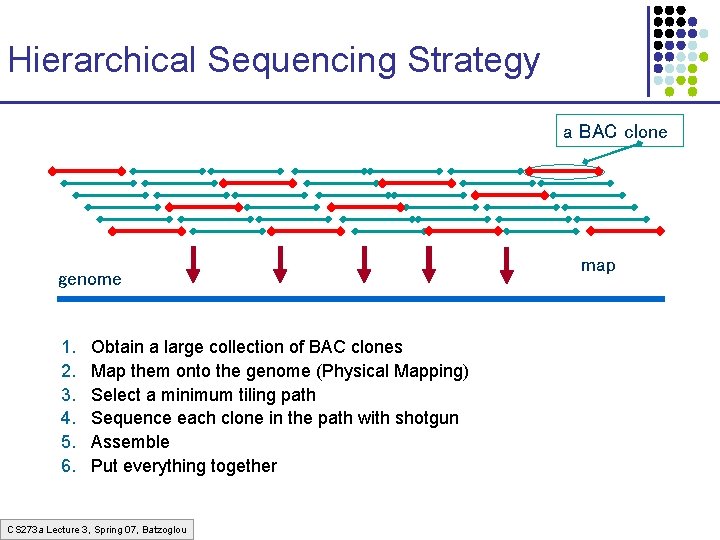
Hierarchical Sequencing Strategy a BAC clone genome 1. 2. 3. 4. 5. 6. Obtain a large collection of BAC clones Map them onto the genome (Physical Mapping) Select a minimum tiling path Sequence each clone in the path with shotgun Assemble Put everything together CS 273 a Lecture 3, Spring 07, Batzoglou map
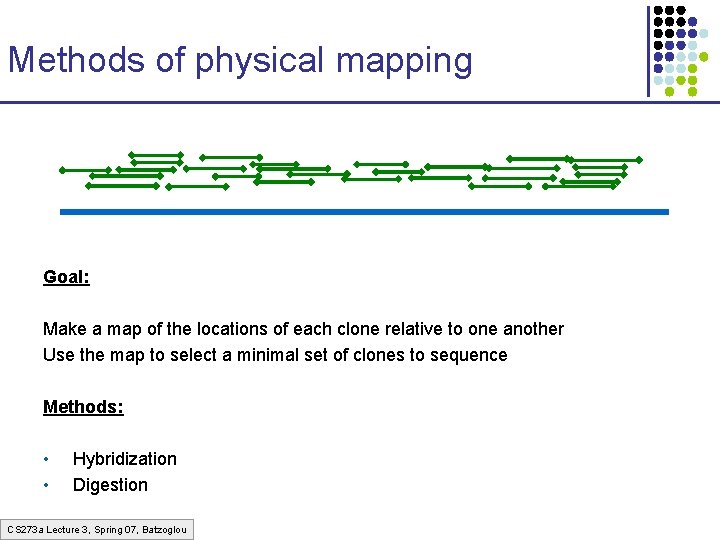
Methods of physical mapping Goal: Make a map of the locations of each clone relative to one another Use the map to select a minimal set of clones to sequence Methods: • • Hybridization Digestion CS 273 a Lecture 3, Spring 07, Batzoglou

1. Hybridization p 1 Short words, the probes, attach to complementary words 1. 2. 3. 4. Construct many probes Treat each BAC with all probes Record which ones attach to it Same words attaching to BACS X, Y overlap CS 273 a Lecture 3, Spring 07, Batzoglou pn

2. Digestion Restriction enzymes cut DNA where specific words appear 1. Cut each clone separately with an enzyme 2. Run fragments on a gel and measure length 3. Clones Ca, Cb have fragments of length { li, lj, lk } overlap Double digestion: Cut with enzyme A, enzyme B, then enzymes A + B CS 273 a Lecture 3, Spring 07, Batzoglou

Online Clone-by-clone The Walking Method CS 273 a Lecture 3, Spring 07, Batzoglou

The Walking Method 1. Build a very redundant library of BACs with sequenced cloneends (cheap to build) 2. Sequence some “seed” clones 3. “Walk” from seeds using clone-ends to pick library clones that extend left & right CS 273 a Lecture 3, Spring 07, Batzoglou
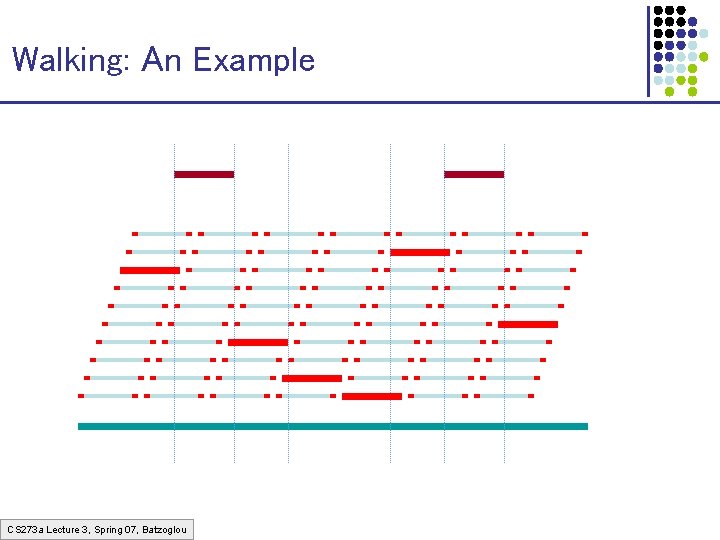
Walking: An Example CS 273 a Lecture 3, Spring 07, Batzoglou
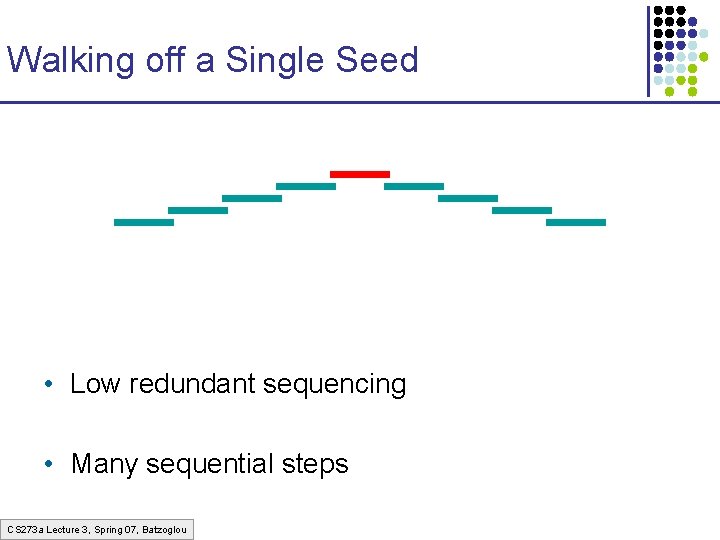
Walking off a Single Seed • Low redundant sequencing • Many sequential steps CS 273 a Lecture 3, Spring 07, Batzoglou

Walking off a single clone is impractical Cycle time to process one clone: 1 -2 months 1. 2. 3. 4. 5. Grow clone Prepare & Shear DNA Prepare shotgun library & perform shotgun Assemble in a computer Close remaining gaps A mammalian genome would need 15, 000 walking steps ! CS 273 a Lecture 3, Spring 07, Batzoglou
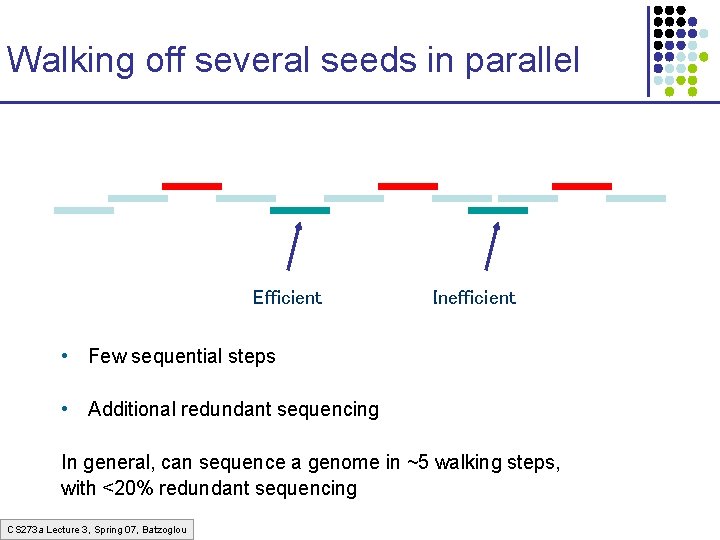
Walking off several seeds in parallel Efficient Inefficient • Few sequential steps • Additional redundant sequencing In general, can sequence a genome in ~5 walking steps, with <20% redundant sequencing CS 273 a Lecture 3, Spring 07, Batzoglou
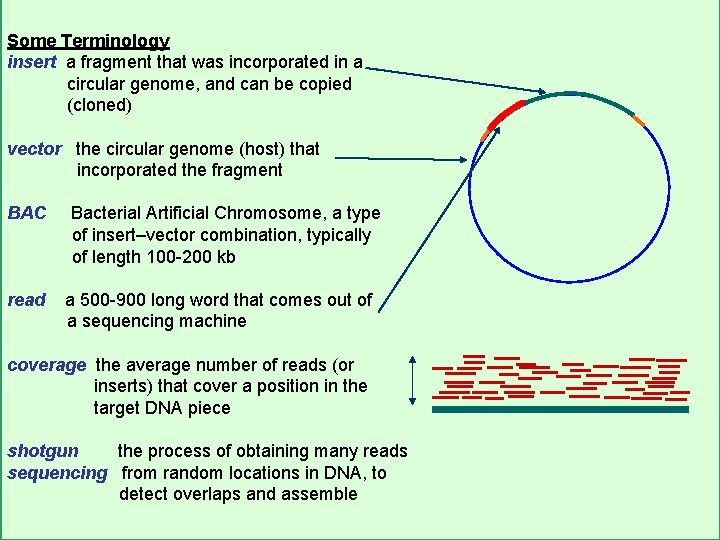
Some Terminology insert a fragment that was incorporated in a circular genome, and can be copied (cloned) vector the circular genome (host) that incorporated the fragment BAC read Bacterial Artificial Chromosome, a type of insert–vector combination, typically of length 100 -200 kb a 500 -900 long word that comes out of a sequencing machine coverage the average number of reads (or inserts) that cover a position in the target DNA piece shotgun the process of obtaining many reads sequencing from random locations in DNA, to detect overlaps and assemble CS 273 a Lecture 3, Spring 07, Batzoglou
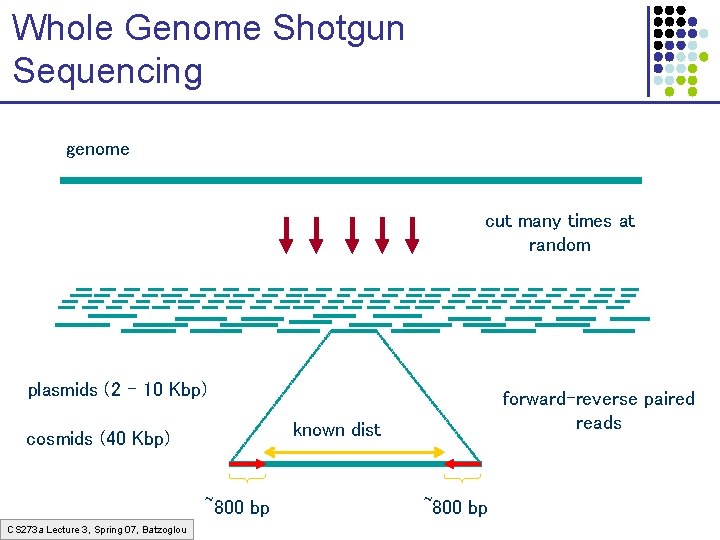
Whole Genome Shotgun Sequencing genome cut many times at random plasmids (2 – 10 Kbp) known dist cosmids (40 Kbp) ~800 bp CS 273 a Lecture 3, Spring 07, Batzoglou forward-reverse paired reads ~800 bp

Fragment Assembly (in whole-genome shotgun sequencing) CS 273 a Lecture 3, Spring 07, Batzoglou

Fragment Assembly Given N reads… Where N ~ 30 million… We need to use a linear-time algorithm CS 273 a Lecture 3, Spring 07, Batzoglou
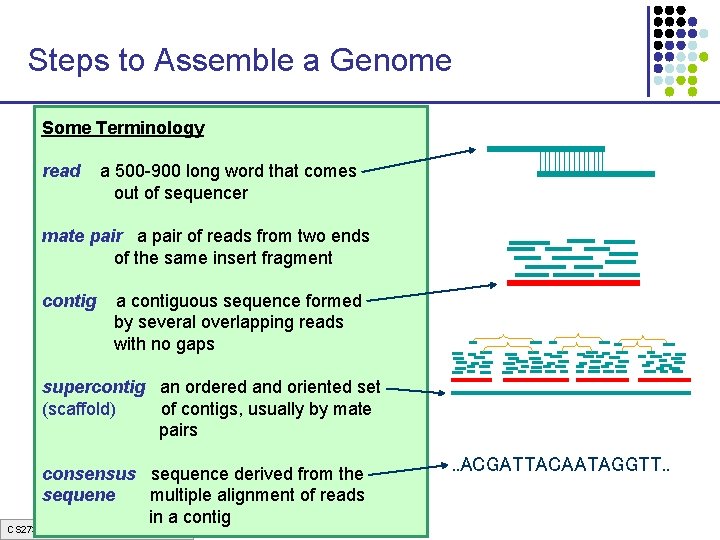
Steps to Assemble a Genome Some Terminology 1. Find overlapping readsthat comes read a 500 -900 long word out of sequencer mate pair a pair of reads from two ends 2. Merge some “good” of reads into of the same insert pairs fragment longer contigs contig a contiguous sequence formed by several overlapping reads with no gaps 3. Link contigs to form supercontigs supercontig an ordered and oriented set (scaffold) of contigs, usually by mate pairs 4. Derive consensus sequence derived from the sequene multiple alignment of reads in a contig CS 273 a Lecture 3, Spring 07, Batzoglou . . ACGATTACAATAGGTT. .

1. Find Overlapping Reads aaactgcagtacggatct aaactgcagt … gtacggatct gggcccaaactgcagtac gggcccaaac … actgcagtac gtacggatctactacaca gtacggatct … ctactacaca CS 273 a Lecture 3, Spring 07, Batzoglou (read, pos. , word, orient. ) (word, read, orient. , pos. ) aaactgcagt actgcagta … gtacggatct gggcccaaac gcccaaact … actgcagtac gtacggatct acggatcta … ctactacaca aaactgcagt acggatcta actgcagta cccaaactg cggatctactacac ctgcagtac gcccaaact ggcccaaac gggcccaaa gtacggatct tactacaca
- Slides: 41