DNA Replication DNA by Watson Crick Franklin When

DNA Replication

DNA by Watson & Crick & Franklin When discovered, the structure suggested how DNA was able to replicate: n The H-bonds between complementary bases break n This n allows the DNA to unzip Each DNA strand then acts as a template to build the complimentary strand n This results in two identical DNA molecules; one for each daughter cell

The big question one time was the method of DNA replication; was it: 1) Conservative: the original ds. DNA was conserved as one molecule for one daughter, and a completely new one for the other daughter, or 2) Semi-conservative: the original molecule is split in half, and the other side is filledin) 3) Dispersive: Each new molecule is comprised of bits and pieces of both new DNA and the original strand

Models of DNA Replication n Alternative models n become experimental predictions conservative P 1 2 Can you design a nifty experiment to verify? semiconservative dispersive
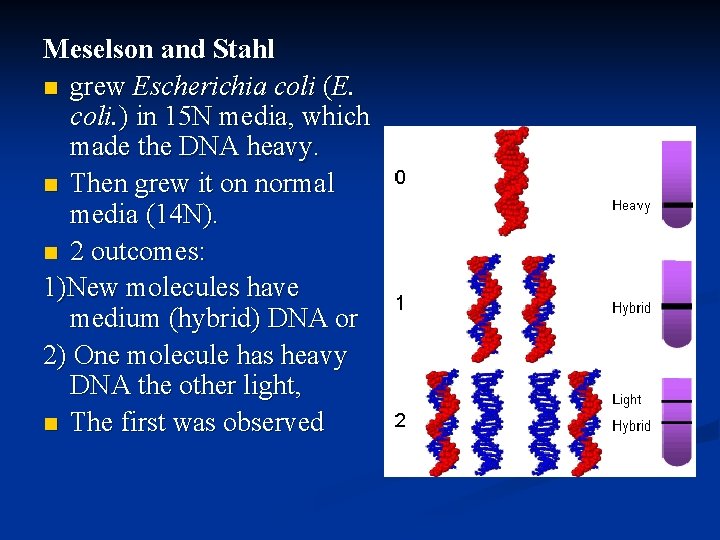
Meselson and Stahl n grew Escherichia coli (E. coli. ) in 15 N media, which made the DNA heavy. n Then grew it on normal media (14 N). n 2 outcomes: 1)New molecules have medium (hybrid) DNA or 2) One molecule has heavy DNA the other light, n The first was observed

1958: Matthew Meselson & Frank Stahl’s Experiment
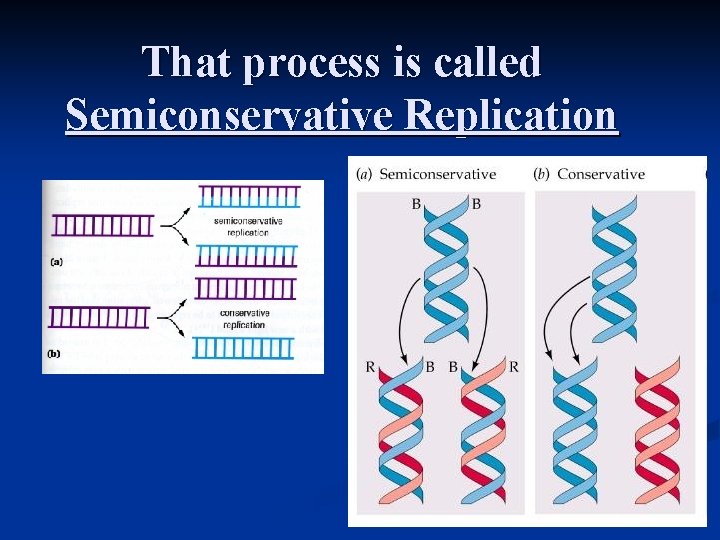
That process is called Semiconservative Replication

Overview. n DNA replication has many steps, but in general: Initiation and unzipping. n Priming n Elongation n Proof-reading. n

Initiation and Unzipping n Replication begins when proteins bind at a specific site on the DNA known as the origin of replication (ori). n n Eukaryotic replication is similar to prokaryotic replication but more complex The closed circular DNA of prokaryotes usually only has one origin of replication (ori) n Linear eukaryotic DNA has multiple ori’s
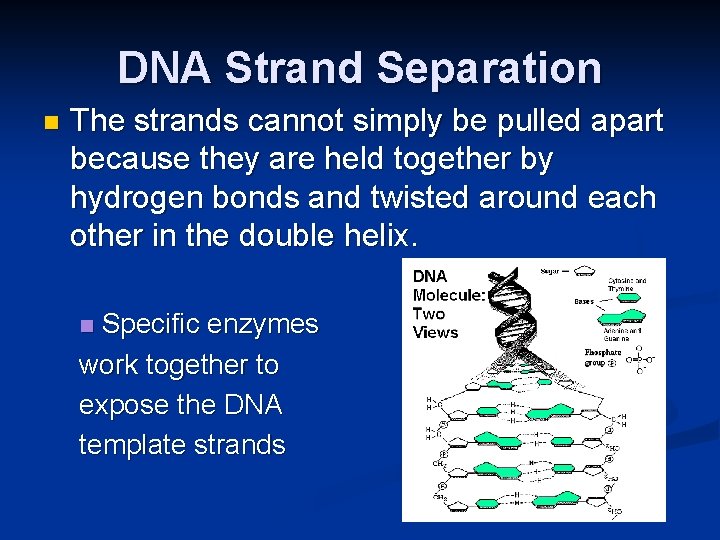
DNA Strand Separation n The strands cannot simply be pulled apart because they are held together by hydrogen bonds and twisted around each other in the double helix. Specific enzymes work together to expose the DNA template strands n

DNA Strand Separation n What is involved? n DNA helicase: unwinds the double helix by breaking the H-bonds at the replication fork. Replication fork: region where enzymes replicating DNA bind to an untwisted, s. s. DNA strand.
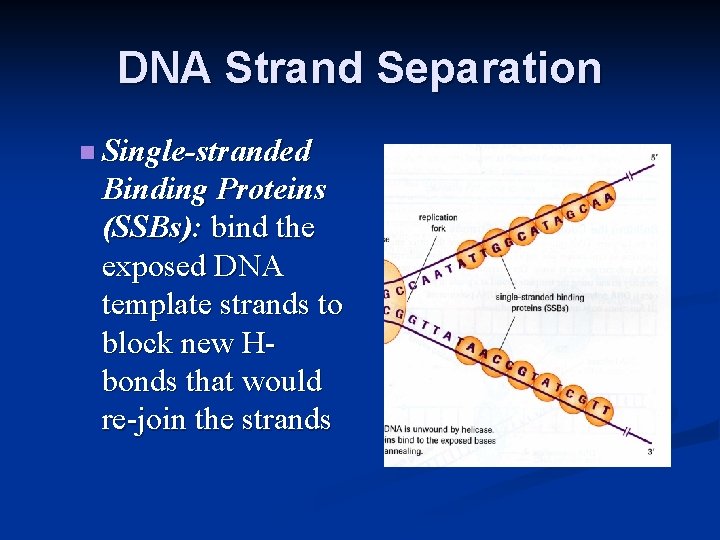
DNA Strand Separation n Single-stranded Binding Proteins (SSBs): bind the exposed DNA template strands to block new Hbonds that would re-join the strands

DNA Strand Separation n DNA gyrase: relieves tension from the unwinding of the DNA strands. It cuts nicks in both strands of DNA, allowing them to swivel around one another and then resealing the cut strands.

Replication begins in 2 directions from the ori ’s as a region of the DNA is unwound. n DNA replication proceeds toward the direction of the replication fork (leading strand) on one strand away from the fork (lagging strand) on the other strand. n In eukaryotes, when 2 replication forks are too near, a replication bubble forms n

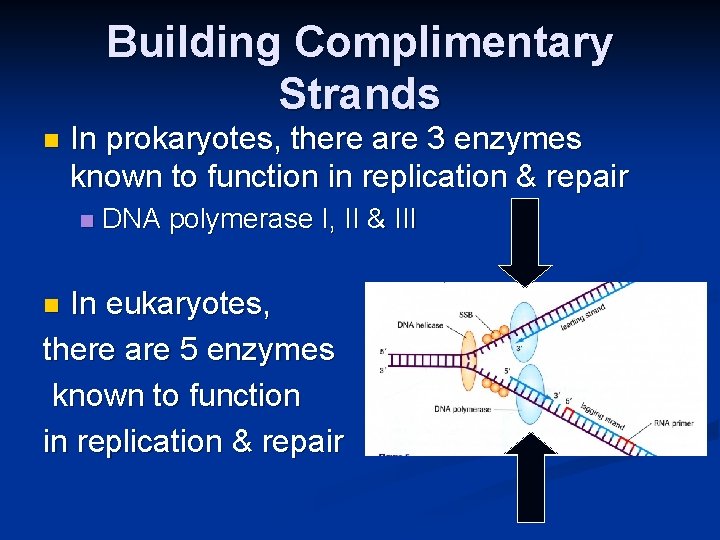
Building Complimentary Strands n In prokaryotes, there are 3 enzymes known to function in replication & repair n DNA polymerase I, II & III In eukaryotes, there are 5 enzymes known to function in replication & repair n
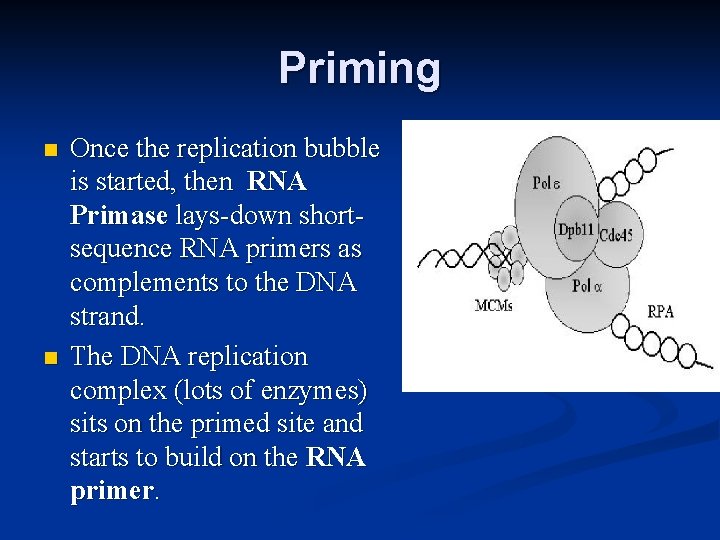
Priming n n Once the replication bubble is started, then RNA Primase lays-down shortsequence RNA primers as complements to the DNA strand. The DNA replication complex (lots of enzymes) sits on the primed site and starts to build on the RNA primer.

RNA primers are synthesized by primase and are temporary n The leading strand (uses 3’-5’ template) is synthesized continuously n The lagging strand (uses 5’-3’ template) is synthesized discontinuously in short fragments n

Elongation DNA polymerase III builds the complimentary strand of DNA n DNA polymerase III adds complimentary nucleotides (deoxyribonucleoside triphosphates) in the 5’ to 3’ direction, using RNA primers as starting points n The RNA primers are needed because DNA Pol III needs a 3’OH to add to. n The lagging strands produces many small segments called Okazaki fragments n


n DNA polymerase I removes the RNA primers from the leading strand fragments from the lagging strand replaces them with the appropriate deoxyribonucleotides. Text page 221, figure 7 b

n DNA ligase joins the Okazaki fragments into one strand on the lagging strand of DNA through the formation of a phosphodiester bond.

As the 2 new strands of DNA are synthesized, 2 d. s. DNA molecules are produced that automatically twist into a helix.



Editing & proofreading DNA n 1000 bases/second = lots of typos! n DNA polymerase I n proofreads & corrects typos n repairs mismatched bases n removes abnormal bases n repairs damage throughout life n reduces error rate from 1 in 10, 000 to 1 in 100 million bases

What does it really look like? 1 2 3 4
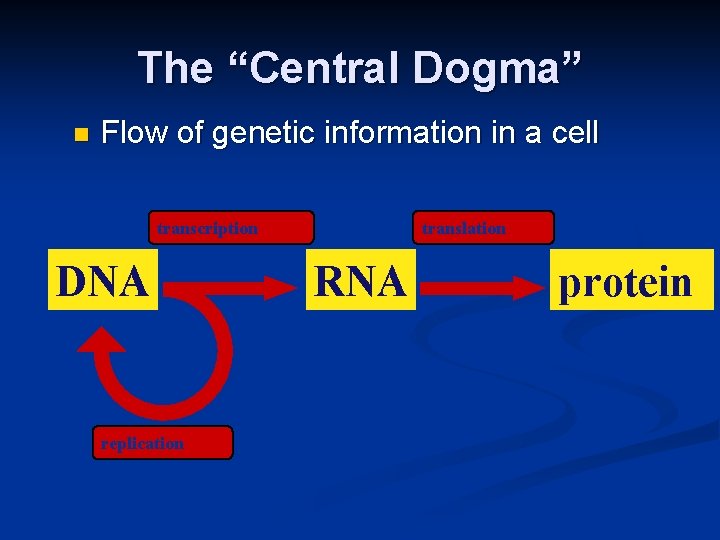
The “Central Dogma” n Flow of genetic information in a cell transcription DNA replication translation RNA protein

Review Questions

Base your answers to the following questions on the choices below: A. Helicase B. Polymerase C. Ligase D. Primase 1. 2. 3. Brings together the Okazaki fragments. Adds nucleotides to the leading strand. Recognizes the origin of replication in the DNA.

4. In DNA synthesis, DNA is A. B. C. D. Read 3’ to 5’ and made 5’ to 3’ Read 3’ to 5’ and made 3’ to 5’ Read 5’ to 3’ and made 5’ to 3;
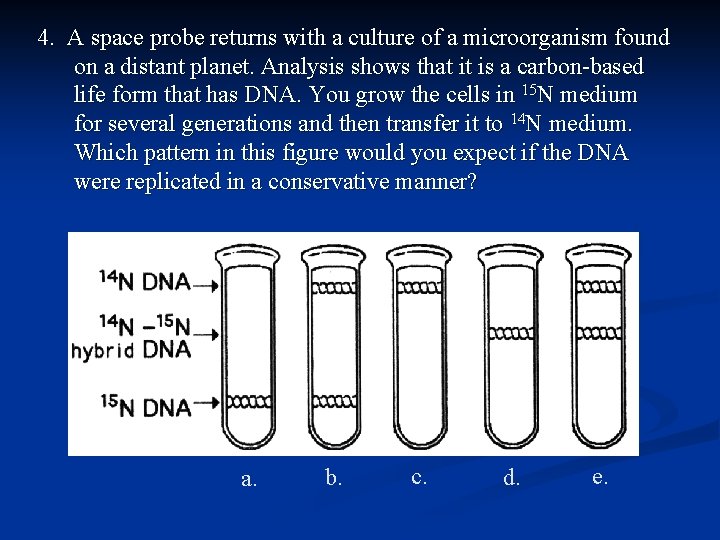
4. A space probe returns with a culture of a microorganism found on a distant planet. Analysis shows that it is a carbon-based life form that has DNA. You grow the cells in 15 N medium for several generations and then transfer it to 14 N medium. Which pattern in this figure would you expect if the DNA were replicated in a conservative manner? a. b. c. d. e.
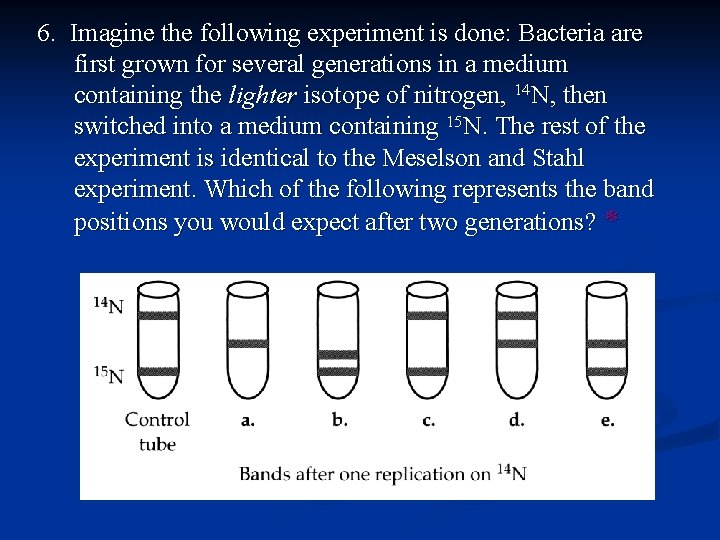
6. Imagine the following experiment is done: Bacteria are first grown for several generations in a medium containing the lighter isotope of nitrogen, 14 N, then switched into a medium containing 15 N. The rest of the experiment is identical to the Meselson and Stahl experiment. Which of the following represents the band positions you would expect after two generations? *
- Slides: 33