DNA repair RNA polymerases Prepared By Dr Meenakshi

DNA repair; RNA polymerases Prepared By: Dr. Meenakshi Bhatt Asst. Professor Dept. of Pharmaceutical Sciences SGRRITS Dehradun Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Key learninggoals: • Understand mechanisms through which mutations arise. • Understand the major DNA repair systems • Know examples of diseases caused by DNA repair deficiency • Understand nomenclature of RNA polymerization • Understand RNAP mechanism of synthesis • Understand division of labor among 3 eukaryotic RNAPs • Understand that RNAP doesn’t know where to start — other proteins to position and initiate it. Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Example: Cancer A major cause of cancer is mutation. Gain-of-function mutations in protooncogenes can cause cancer. These genes generally promote cell division. Usually dominant (one bad copy causes disease) Loss-of-function mutations in antioncogenes can cause cancer. Usually recessive (one good copy is usually enough to stay healthy) It is generally thought that it takes multiple mutational “hits” to cause cancer. 2 Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
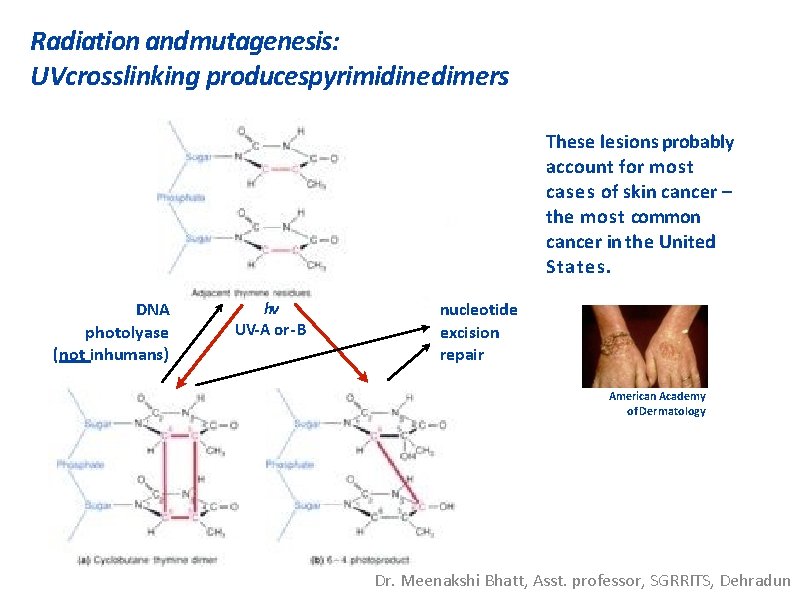
Radiation andmutagenesis: UVcrosslinking producespyrimidine dimers These lesions probably account for most cases of skin cancer – the most common cancer in the United States. DNA photolyase (not in humans) hv UV-A or -B nucleotide excision repair American Academy of Dermatology Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
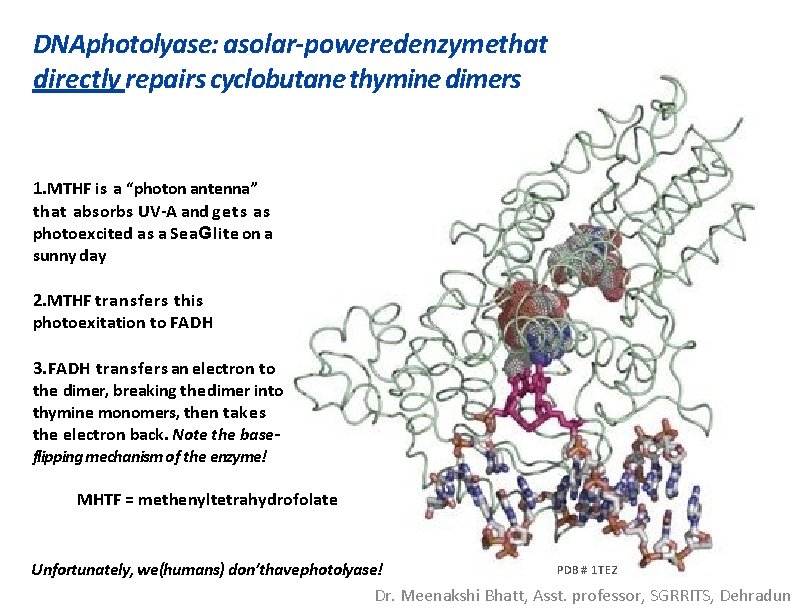
DNAphotolyase: asolar-poweredenzymethat directly repairs cyclobutane thymine dimers 1. MTHF is a “photon antenna” that absorbs UV-A and gets as photoexcited as a Sea. Glite on a sunny day 2. MTHF transfers this photoexitation to FADH 3. FADH transfers an electron to the dimer, breaking the dimer into thymine monomers, then takes the electron back. Note the baseflipping mechanism of the enzyme! MHTF = methenyltetrahydrofolate PDB # 1 TEZ Unfortunately, we(humans) don’thavephotolyase! Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
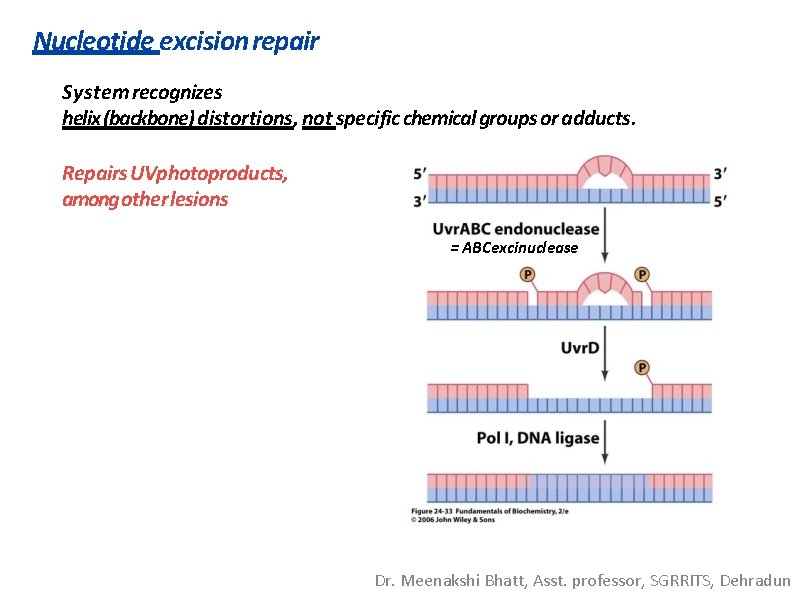
Nucleotide excision repair System recognizes helix (backbone) distortions, not specific chemical groups or adducts. Repairs UVphotoproducts, among other lesions = ABCexcinuclease Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
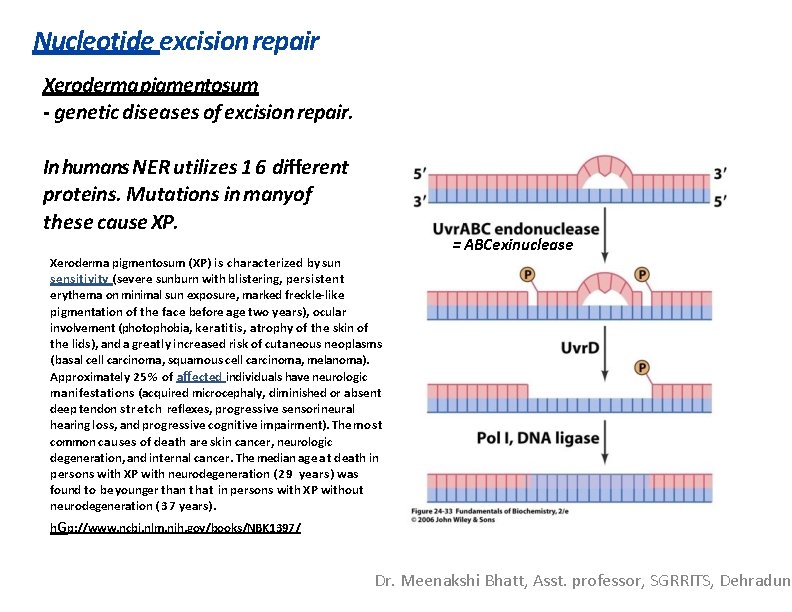
Nucleotide excision repair Xeroderma pigmentosum - genetic diseases of excision repair. In humans NER utilizes 1 6 different proteins. Mutations in manyof these cause XP. Xeroderma pigmentosum (XP) is characterized by sun sensitivity (severe sunburn with blistering, persistent erythema on minimal sun exposure, marked freckle-like pigmentation of the face before age two years), ocular involvement (photophobia, keratitis, atrophy of the skin of the lids), and a greatly increased risk of cutaneous neoplasms (basal cell carcinoma, squamous cell carcinoma, melanoma). Approximately 25% of affected individuals have neurologic manifestations (acquired microcephaly, diminished or absent deep tendon stretch reflexes, progressive sensorineural hearing loss, and progressive cognitive impairment). The most common causes of death are skin cancer, neurologic degeneration, and internal cancer. The median age at death in persons with XP with neurodegeneration ( 2 9 years) was found to be younger than that in persons with XP without neurodegeneration ( 3 7 years). = ABCexinuclease h. Gp: //www. ncbi. nlm. nih. gov/books/NBK 1397/ Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
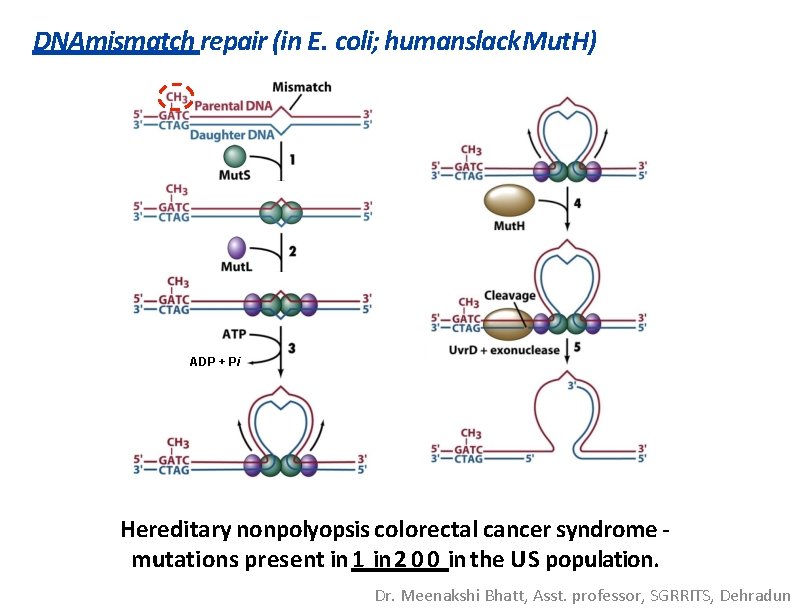
DNAmismatch repair (in E. coli; humanslack Mut. H) ADP + Pi Hereditary nonpolyopsis colorectal cancer syndrome mutations present in 1 in 2 0 0 in the US population. Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
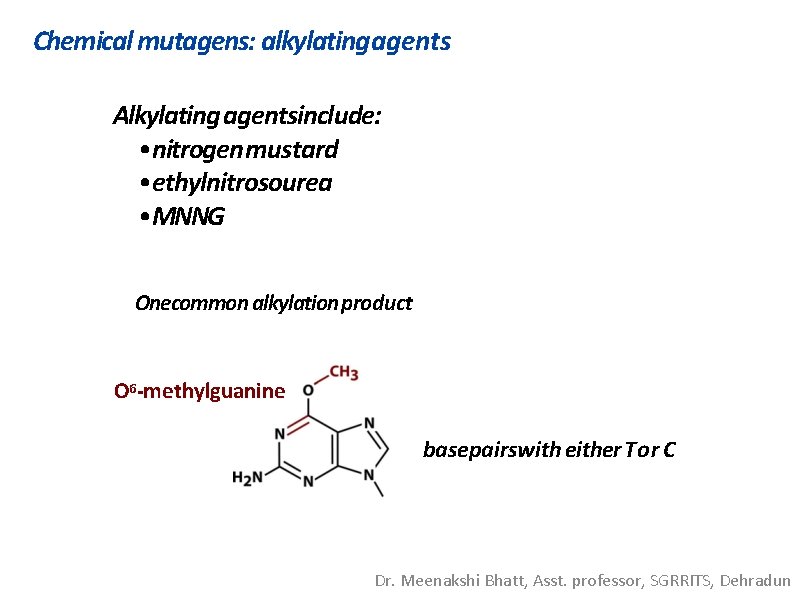
Chemical mutagens: alkylating agents Alkylating agentsinclude: • nitrogen mustard • ethylnitrosourea • MNNG Onecommon alkylation product O 6 -methylguanine basepairswith either Tor C Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Chemical mutagens: oxidizers >100 different oxidative DNA modifications Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
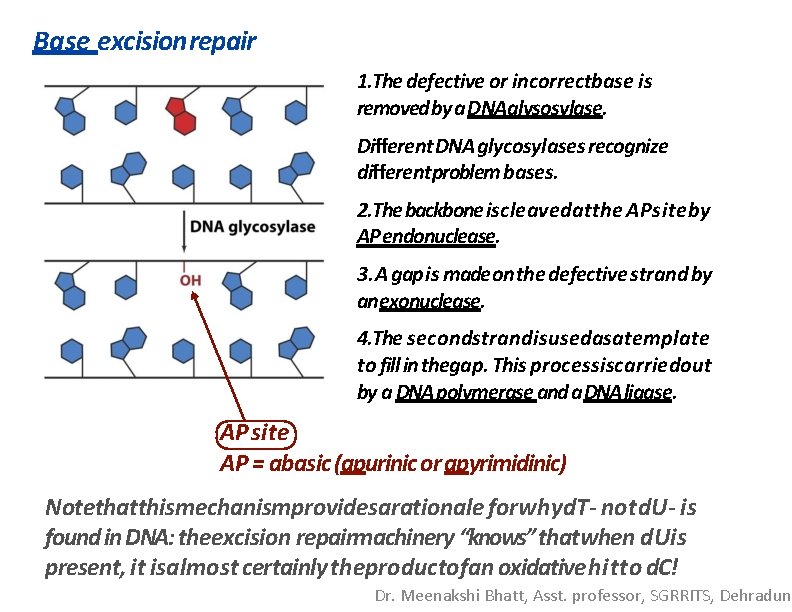
Base excisionrepair 1. The defective or incorrectbase is removed by a DNAglysosylase. Different DNA glycosylases recognize differentproblem bases. 2. The backbone iscleavedatthe APsiteby AP endonuclease. 3. A gap is made on the defective strand by an exonuclease. 4. The secondstrandisusedasatemplate to fill in thegap. This processiscarriedout by a DNA polymerase and a. DNA ligase. AP site AP = abasic (apurinic or apyrimidinic) Notethatthismechanismprovidesarationale forwhyd. T- notd. U- is found in DNA: theexcision repairmachinery “knows” thatwhen d. Uis present, it isalmost certainly theproductofan oxidative hitto d. C! Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
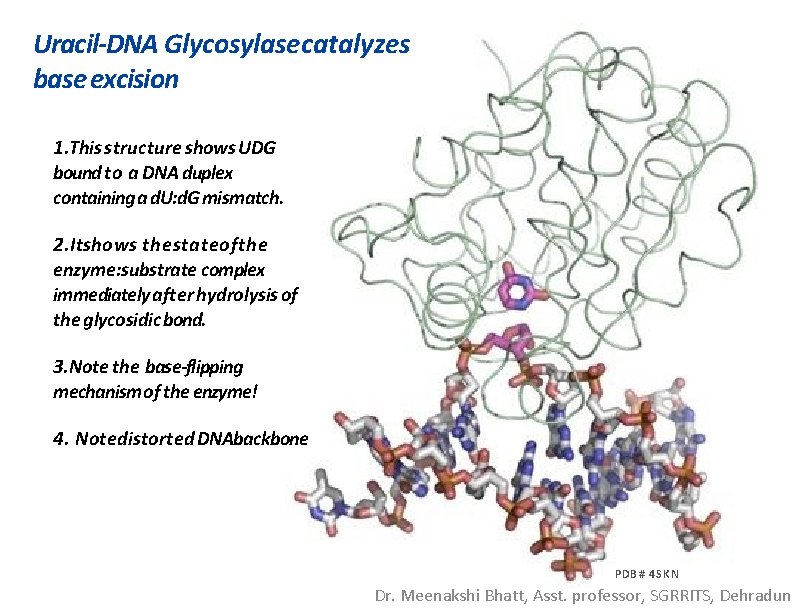
Uracil-DNA Glycosylase catalyzes base excision 1. This structure shows UDG bound to a DNA duplex containing a d. U: d. G mismatch. 2. Itshows thestateofthe enzyme: substrate complex immediately after hydrolysis of the glycosidic bond. 3. Note the base-flipping mechanism of the enzyme! 4. Notedistorted DNAbackbone PDB # 4 SKN Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Genetic mutagens: transposons, retrotransposons, andviruses that insert their genomes into the host’s genome Transposons are found in all organisms. ≥ 45% of our genome is composed of retrotransposons and retrotransposon debris! Because many of these elements can insert into almost any sequence, they often disrupt gene function. Tn 3, abacterial transposon responsible for the spread of penicillin resistance Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Summary of DNA repair systems Type of Damage Surveillance/ Repair System Key Proteins Required DNA backbone breaks (nicks; spontaneous hydrolysis) Direct Repair DNA ligase UV photoproducts: cyclobutane pyrimidine dimer, 6 - 4 photoproduct Direct Repair DNA photolyase [but not in humans : -( ] Base misincorporation (e. g. , from tautomerization) Proofreading DNA Polymerase with 3´→ 5´ exonuclease activity Abnormal bases in DNA (d. U); Covalently modified bases (spontaneous deamination; alkylation products) Base Excision Repair DNA glycosidases Apurinic (AP) endonuclease DNA Pol I, DNA ligase UV photoproducts and other lesions that cause major structural changes Nucleotide Excision Repair ABC exinuclease DNA polymerase I DNA ligase Base mismatches; Short deletions and insertions; Base modifications that distort double helix DNA Mismatch Repair Methylases; Mut. H, Mut. L, Mut. S; exonuclease I; Helicase, SSB, DNA Pol I or III, DNA ligase Major template damage to duplex DNA; Double-strand backbone breaks Recombination Repair Not covered in this course 13 Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

TRANS CRIPTION Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
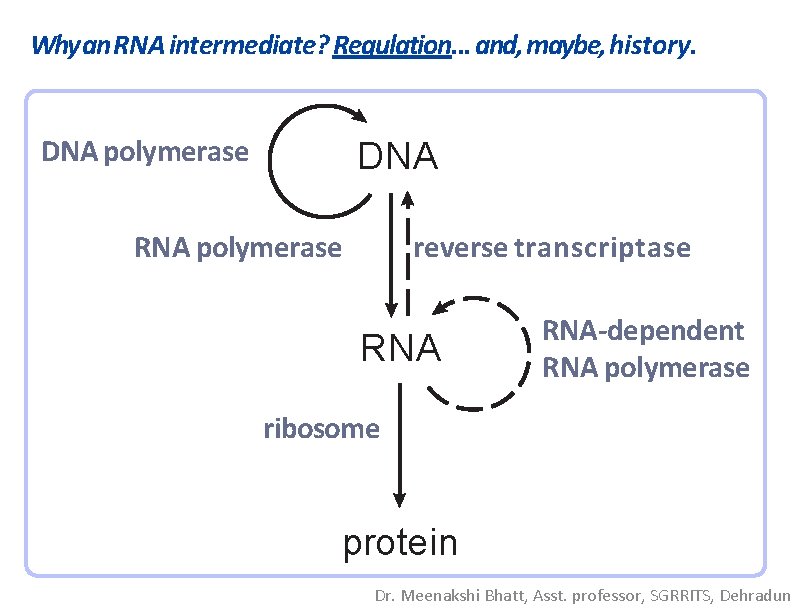
Why an RNA intermediate? Regulation. . . and, maybe, history. DNA polymerase DNA RNA polymerase reverse transcriptase RNA-dependent RNA polymerase ribosome protein Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
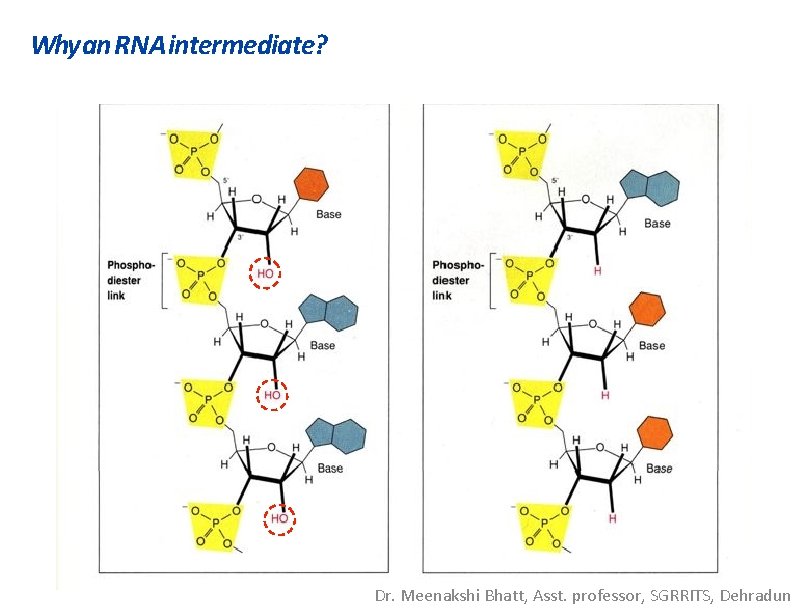
Why an RNA intermediate? Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
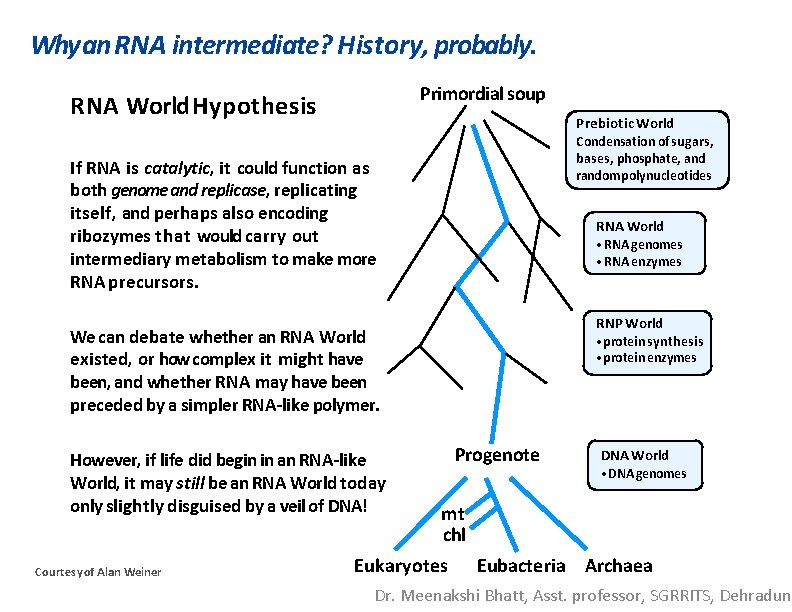
Why an RNA intermediate? History, probably. Primordial soup RNA World Hypothesis Prebiotic World Condensation of sugars, bases, phosphate, and random polynucleotides If RNA is catalytic, it could function as both genome and replicase, replicating itself, and perhaps also encoding ribozymes that would carry out intermediary metabolism to make more RNA precursors. RNA World • RNA genomes • RNA enzymes RNP World We can debate whether an RNA World existed, or how complex it might have been, and whether RNA may have been preceded by a simpler RNA-like polymer. However, if life did begin in an RNA-like World, it may still be an RNA World today only slightly disguised by a veil of DNA! Courtesy of Alan Weiner • protein synthesis • protein enzymes Progenote DNA World • DNA genomes mt chl Eukaryotes Eubacteria Archaea Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
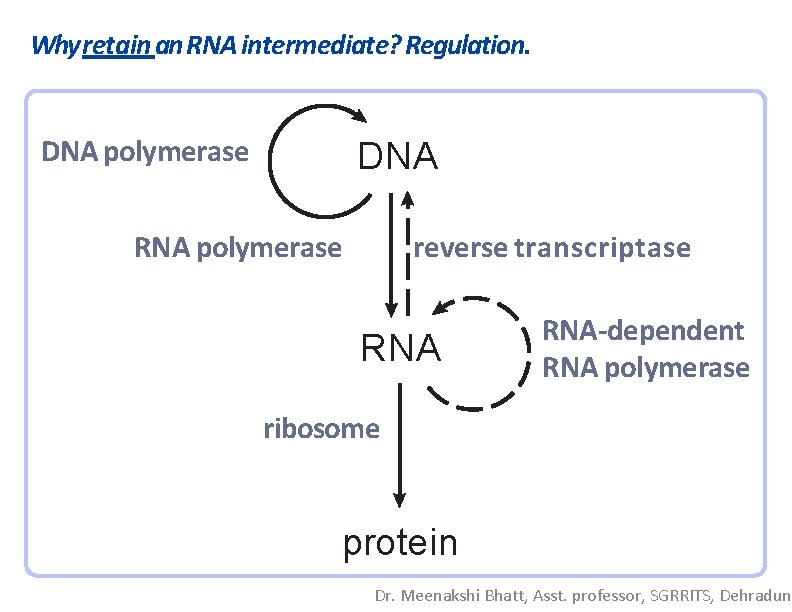
Why retain an RNA intermediate? Regulation. DNA polymerase DNA RNA polymerase reverse transcriptase RNA-dependent RNA polymerase ribosome protein Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Why an RNA intermediate? Amplification. The cell usesjust-in-time manufacturing: you can think of Ribosomesasfactories, andm. RNA molecules aswork orderssenttothefactory— “I need 25, 000 molecules ofglyceraldehyde-3 phosphate dehydrogenase in the next 2 0 minutes” Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
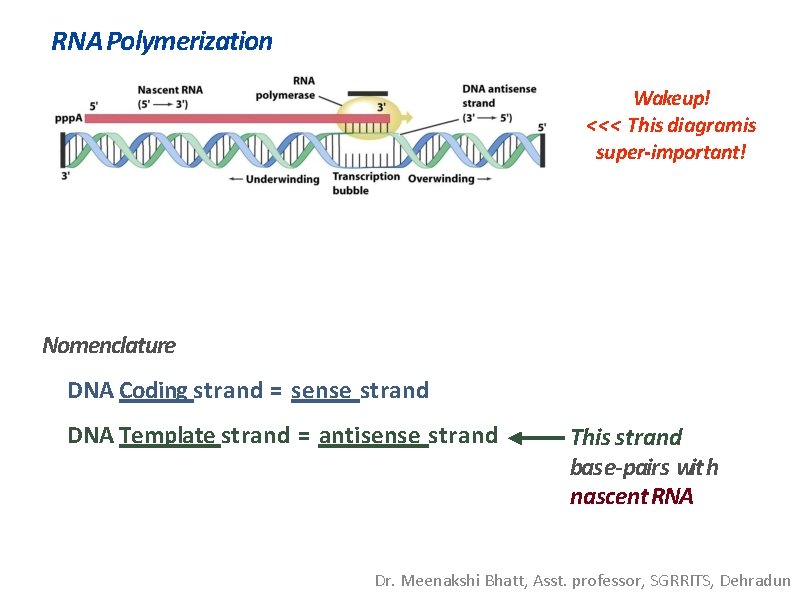
RNA Polymerization Wakeup! <<< This diagramis super-important! Nomenclature DNA Coding strand = sense strand DNA Template strand = antisense strand This strand base-pairs with nascent RNA Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
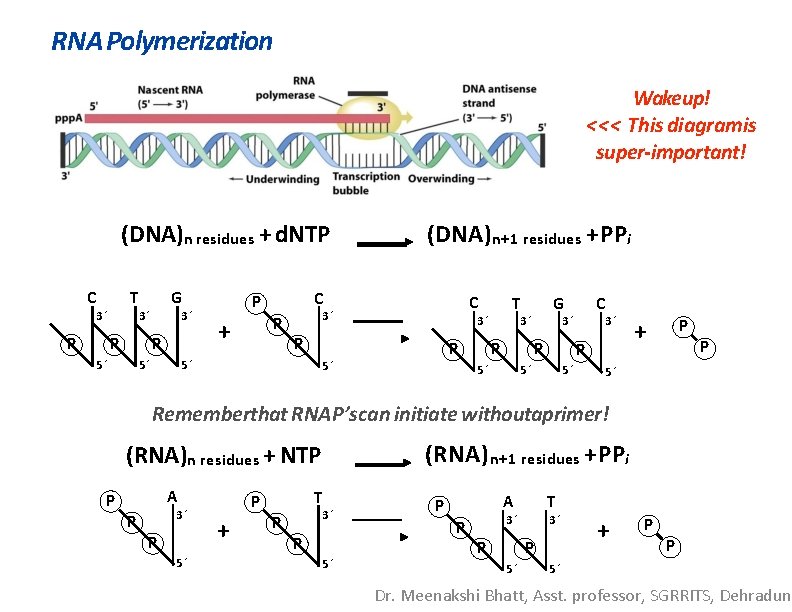
RNA Polymerization Wakeup! <<< This diagramis super-important! (DNA)n residues + d. NTP C T 3´ P G 3´ P 5´ 5´ C P P + (DNA)n+1 residues + PPi C 3´ 3´ P 5´ T P 5´ G 3´ P 5´ C 3´ P 5´ 5´ P + P 5´ Rememberthat RNAP’scan initiate withoutaprimer! (RNA)n residues + NTP A P P 3´ P 5´ T P + (RNA) n+1 residues + PPi P 3´ P 5´ P P A T 3´ 3´ P P 5´ + P P 5´ Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
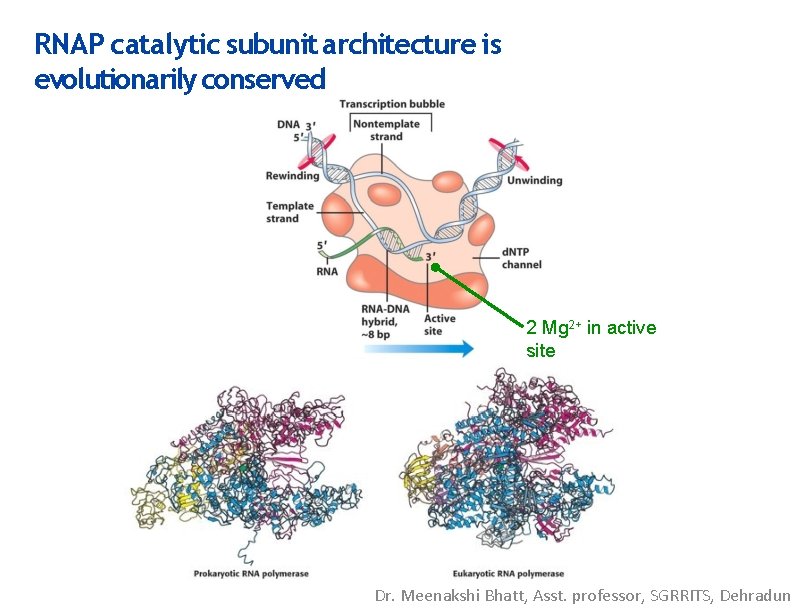
RNAP catalytic subunit architecture is evolutionarily conserved 2 Mg 2+ in active site Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
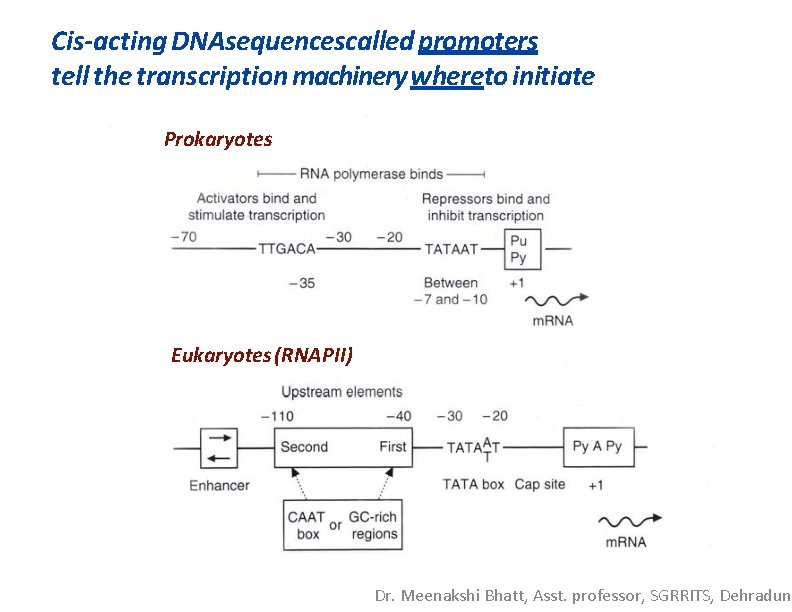
Cis-acting DNAsequencescalled promoters tell the transcription machinery whereto initiate Prokaryotes Eukaryotes (RNAPII) Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
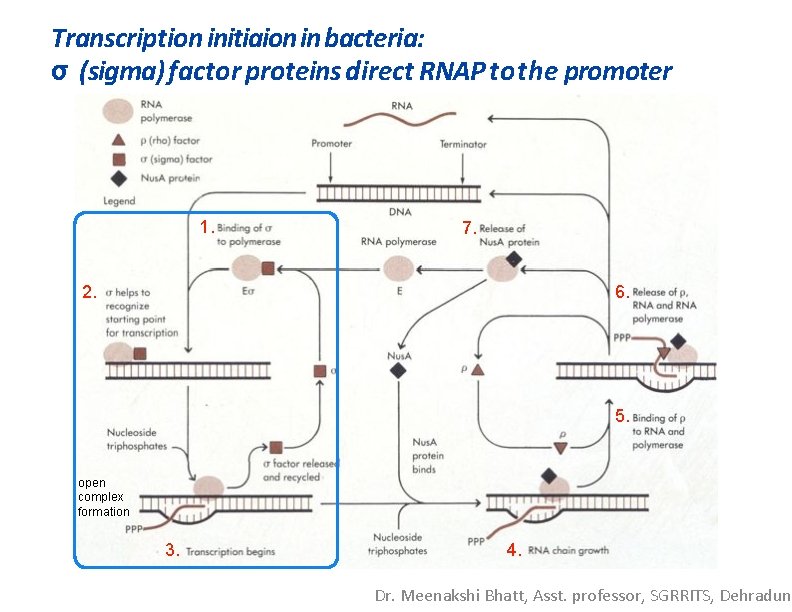
Transcription initiaion in bacteria: σ (sigma) factor proteins direct RNAP tothe promoter 1. 7. 2. 6. 5. open complex formation 3. 4. Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Fromhereon, we’ll focuson transcription ineukaryotes, not bacteria! Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun

Three (3) eukaryotic DNA-dependent RNApolymerases RNAP I synthesizes most r. RNA; resides innucleolus RNAP II synthesizes most m. RNA; resides in nucleoplasm RNAP III synthesizes 5 S r. RNA, t. RNA, and many other RNA types; resides in nucleoplasm. Other dedicated RNAP’sare in mitochondria andcholoroplasts Nature 2 2 4 , 2 3 4 - 2 3 7 ( 1 8 October 1 9 6 9 ) |doi: 10. 1038/224234 a 0; Multiple Forms of DNA-dependent RNA Polymerase in Eukaryotic Organisms ROBERT G. ROEDER* & WILLIAM J. RUTTER† Department of Biochemistry, University of Washington, Sea. Gle 9 8 1 0 5 Abstract Three distinct RNA polymerase activities have been isolated from developing sea urchin embryos. In rat liver nuclei there are two RNA polymerase activities. One polymerase (I) is probably localized in the nucleolus and one (II) in the nucleoplasm. Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
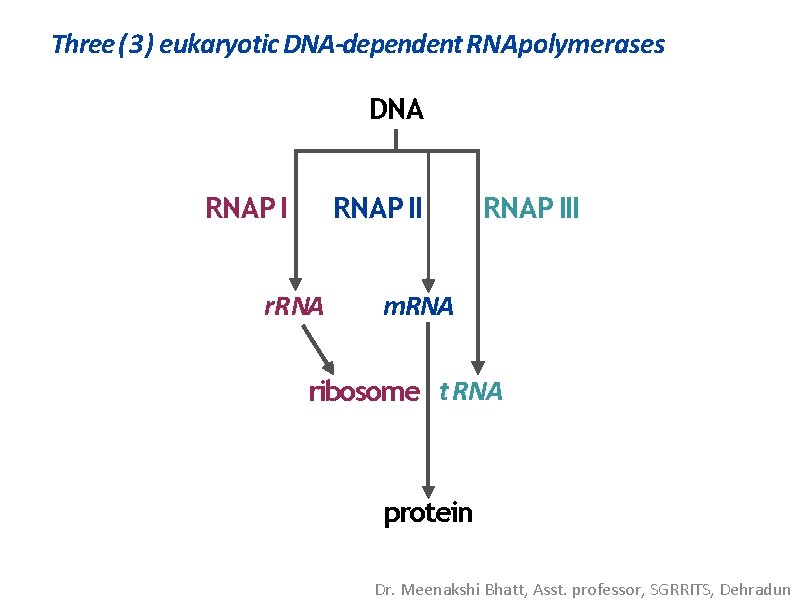
Three (3) eukaryotic DNA-dependent RNApolymerases DNA RNAP II r. RNAP III m. RNA ribosome t. RNA protein Dr. Meenakshi Bhatt, Asst. professor, SGRRITS, Dehradun
- Slides: 28