DNA Recombination Roles Types Homologous recombination in E

DNA Recombination • • Roles Types Homologous recombination in E. coli Transposable elements

Biological Roles for Recombination 1. Generating new gene/allele combinations (crossing over during meiosis) 2. Generating new genes (e. g. , Immunoglobulin rearrangement) 3. Integration of a specific DNA element (or virus) 4. DNA repair

Practical Uses of Recombination 1. Used to map genes on chromosomes - recombination frequency proportional to distance between genes 2. Making transgenic cells and organisms
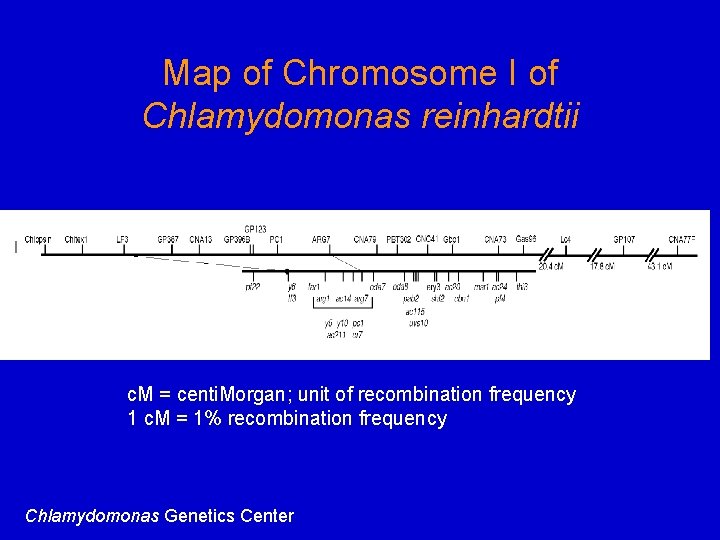
Map of Chromosome I of Chlamydomonas reinhardtii c. M = centi. Morgan; unit of recombination frequency 1 c. M = 1% recombination frequency Chlamydomonas Genetics Center

Types of Recombination 1. Homologous - occurs between sequences that are nearly identical (e. g. , during meiosis) 2. Site-Specific - occurs between sequences with a limited stretch of similarity; involves specific sites 3. Transposition – DNA element moves from one site to another, usually little sequence similarity involved
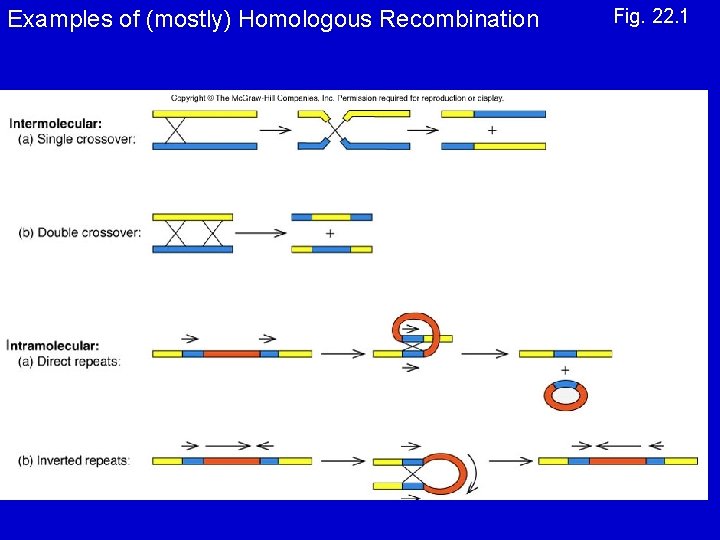
Examples of (mostly) Homologous Recombination Fig. 22. 1
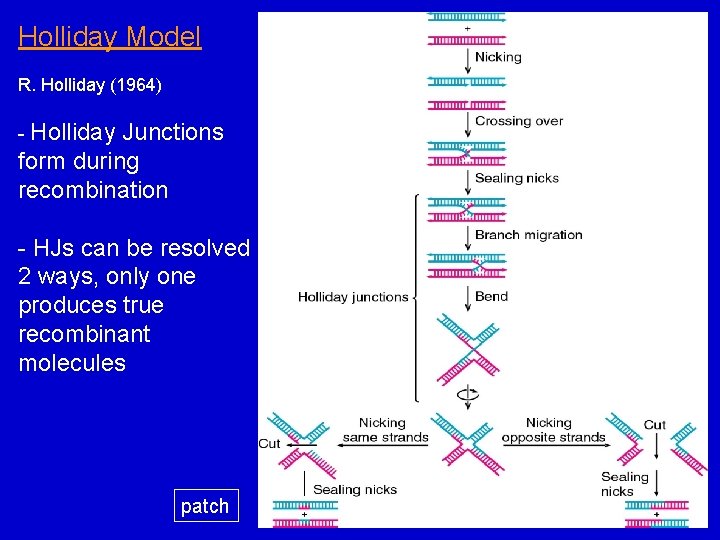
Holliday Model R. Holliday (1964) - Holliday Junctions form during recombination - HJs can be resolved 2 ways, only one produces true recombinant molecules patch

EM of a Holliday Junction w/a few melted base pairs around junction Fig. 22. 3
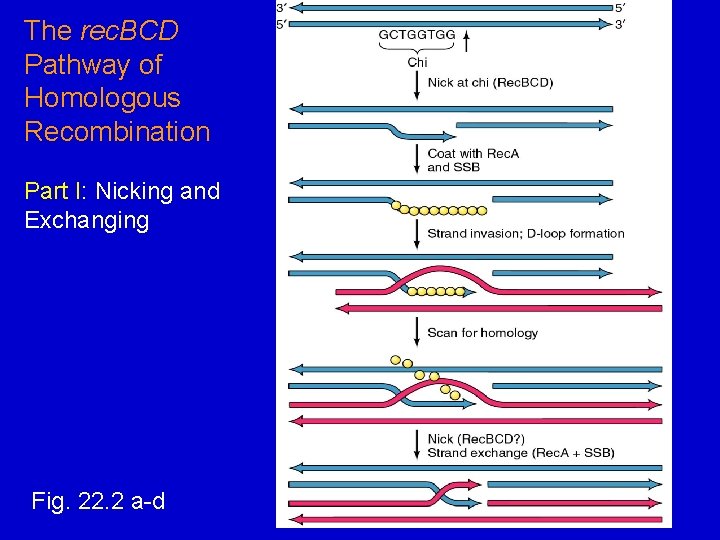
The rec. BCD Pathway of Homologous Recombination Part I: Nicking and Exchanging Fig. 22. 2 a-d

rec. BCD Pathway of Homologous Recomb. Part I: Nicking and Exchanging 1. 2. 3. 4. 5. A nick is created in one strand by rec. BCD at a Chi sequence (GCTGGTGG), found every 5000 bp. Unwinding of DNA containing Chi sequence by rec. BCD allows binding of SSB and rec. A promotes strand invasion into homologous DNA, displacing one strand. The displaced strand base-pairs with the single strand left behind on the other chromosome. The displaced and now paired strand is nicked (by rec. BCD? ) to complete strand exchange.
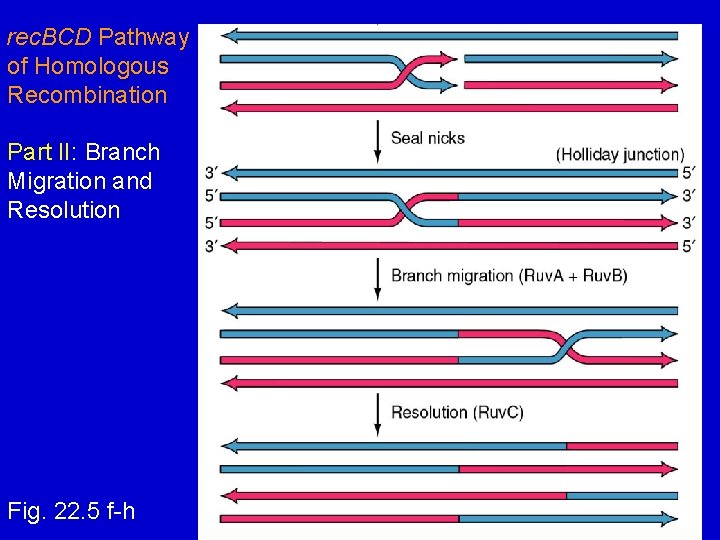
rec. BCD Pathway of Homologous Recombination Part II: Branch Migration and Resolution Fig. 22. 5 f-h

rec. BCD Pathway of Homologous Recom. Part II: Branch Migration and Resolution 1. Nicks are sealed Holliday Junction 2. Branch migration (ruv. A + ruv. B) 3. Resolution of Holliday Junction (ruv. C)

Rec. BCD : A Complex Enzyme • Rec. BCD has: 1. Endonuclease subunits (rec. BC) that cut one DNA strand close to Chi sequence. 2. DNA helicase activity (rec. D subunit) and a DNA-dependent ATPase activity – unwinds DNA to generate the 3’ SS tails
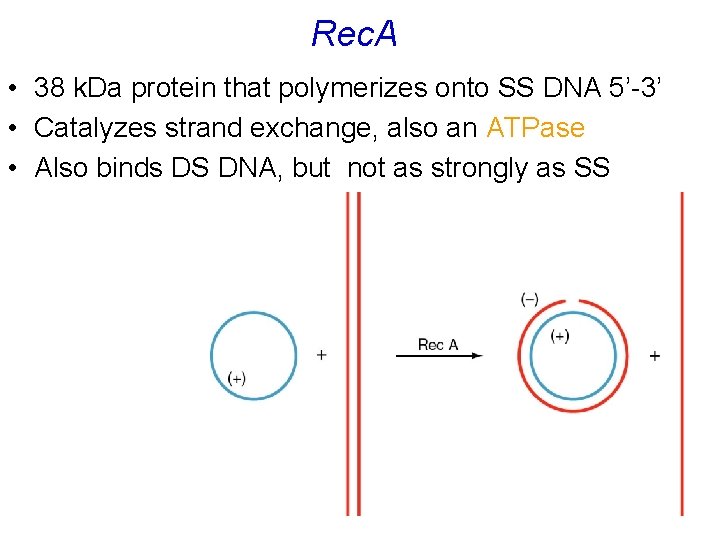
Rec. A • 38 k. Da protein that polymerizes onto SS DNA 5’-3’ • Catalyzes strand exchange, also an ATPase • Also binds DS DNA, but not as strongly as SS
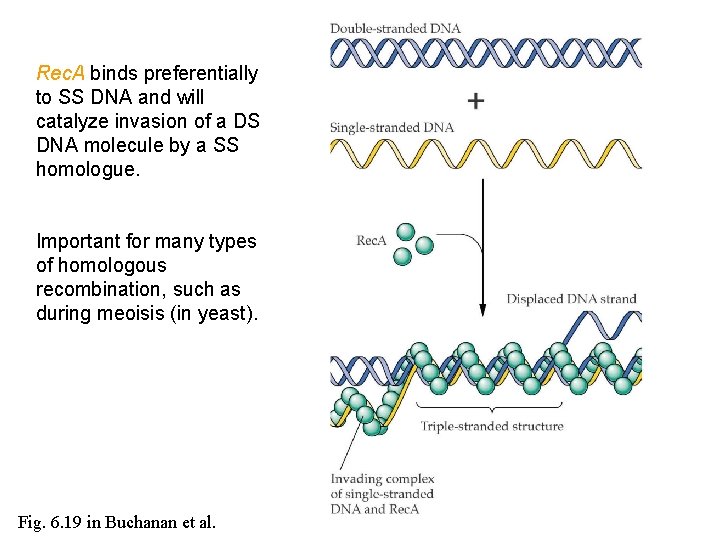
Rec. A binds preferentially to SS DNA and will catalyze invasion of a DS DNA molecule by a SS homologue. Important for many types of homologous recombination, such as during meoisis (in yeast). Fig. 6. 19 in Buchanan et al.

Rec. A Function Dissected • 3 steps of strand exchange: 1. Pre-synapsis: rec. A coats single-stranded DNA (accelerated by SSB, so get more relaxed structure). 2. Synapsis: alignment of complementary sequences in SS and DS DNA (paranemic or side-by-side structure). 3. Post-synapsis or strand-exchange: SS DNA replaces the same strand in the duplex to form a new DS DNA (requires ATP hydrolysis).

Ruv. A and Ruv. B • DNA helicase that catalyzes branch migration • Ruv. A tetramer binds to HJ (each DNA helix between subunits), forces it into square planar conformation • 2 copies of Ruv. B bind at the HJ (to Ruv. A and 2 of the DNA helices) – Ruv. B is a hexamer ring, has helicase & ATPase activity • Branch migration is in the direction of rec. A mediated strand-exchange
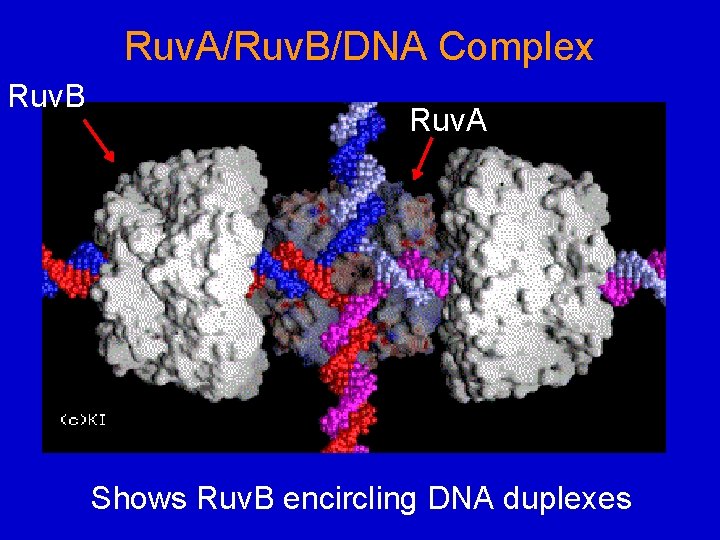
Ruv. A/Ruv. B/DNA Complex Ruv. B Ruv. A Shows Ruv. B encircling DNA duplexes
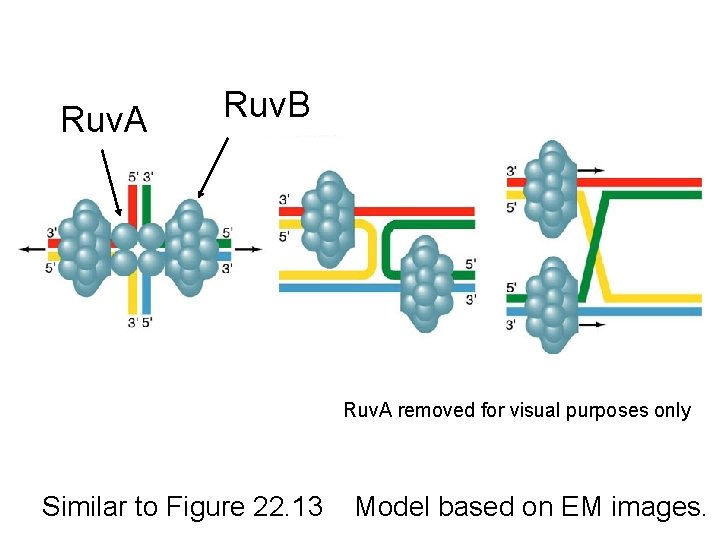
Ruv. A Ruv. B Ruv. A removed for visual purposes only Similar to Figure 22. 13 Model based on EM images.

Ruv. C : resolvase • Endonuclease that cuts 2 strands of HJ • Binds to HJ as a dimer (that already has Ruv. A/Ruv. B) • Consensus sequence: (A/T)TT (G/C) - occurs frequently in E. coli genome - branch migration needed to reach consensus sequence!
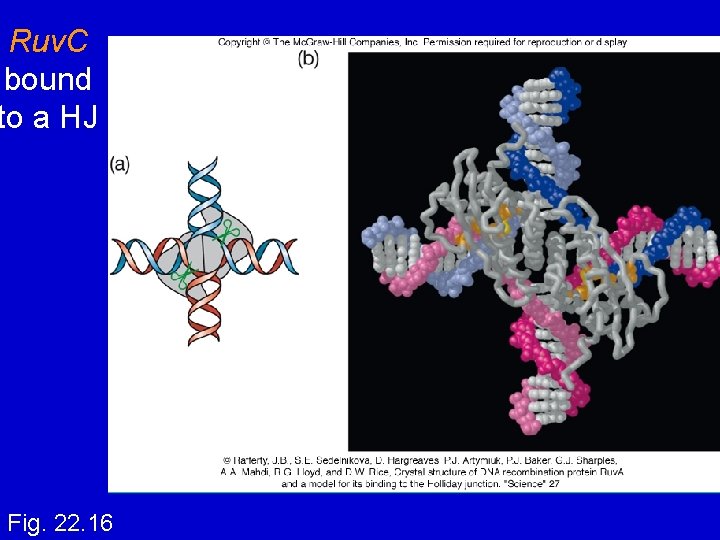
Ruv. C bound to a HJ Fig. 22. 16
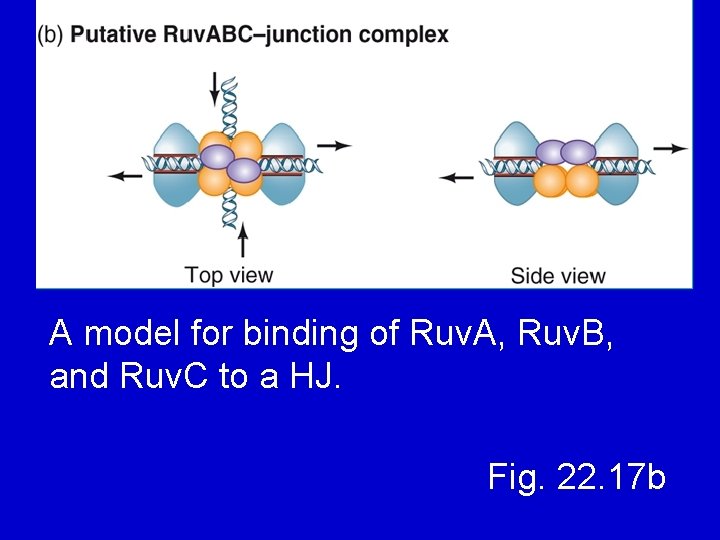
A model for binding of Ruv. A, Ruv. B, and Ruv. C to a HJ. Fig. 22. 17 b
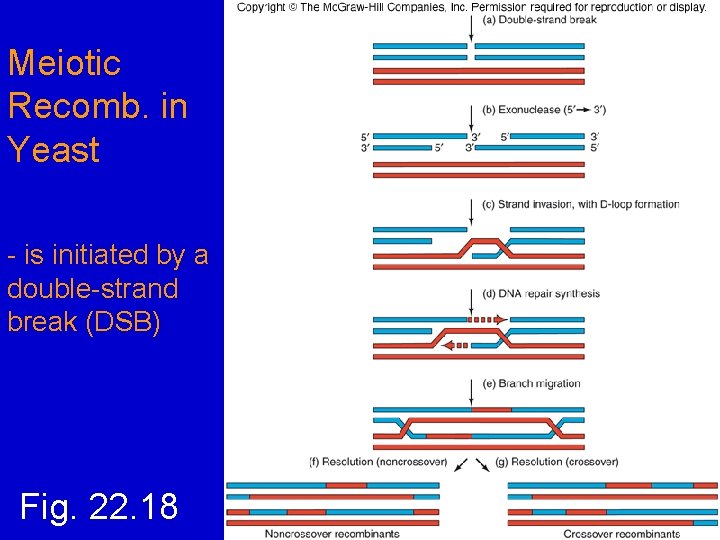
Meiotic Recomb. in Yeast - is initiated by a double-strand break (DSB) Fig. 22. 18

Repair of double-strand breaks (DSBs) in non-dividing or mitotic cells DSBs probably most severe form of DNA damage, can cause loss of genes or even cell death (apoptosis) DSBs caused by: - ionizing radiation - certain chemicals - some enzymes (topoisomerases, endonucleases) - torsional stress

2 general ways to repair DSBs: 1. Homologous recombination (HR) - repair of broken DNA using the intact homologue, very similar to meiotic recombination. Very accurate. 2. Non-homologous end joining (NHEJ) - ligating nonhomologous ends. Prone to errors, ends can be damaged before religation (genetic material lost) or get translocations. (Mechanism in Fig 20. 38) Usage: NHEJ >> HR in plants and animals
- Slides: 25