DNA Recombination Dr Abdul Qayyum Rao Associate Professor

DNA Recombination Dr Abdul Qayyum Rao Associate Professor CEMB , PU

Definition • The process in which a DNA segment moves from one DNA molecule to another DNA molecule

• DNA recombination is the exchange of DNA strands to produce new nucleotide sequence arrangements • Recombination occurs typically, though not exclusively, between regions of similar sequence by breaking and rejoining DNA segments • It is essential for generating genetic diversity and for maintaining genome integrity
Exchange of DNA strands between similar regions
Breaking and rejoining DNA fragments to produce new nucleotide/sequence combinations or arrangements

Biological roles of Recombination • Generating new gene/allele combinations • Generating new genes (e. g. Immuno-globulin rearrangement) • DNA repair (stalled replication forks) • Integration of a specific DNA element

§ This process scrambles the genes of maternal and paternal chromosomes , so nonparental combinations occur in the offspring § This scrambling is valuable because new progeny have new genetic make up and better chances of survival then their parents § Physical links between homologous chromosomes allow their segregation and separation properly during meiosis § Allow cells to deal with the DNA damage and repairing

Practical uses of Recombination • Used to map genes on chromosomes (recombination frequency proportional to distance between genes) • Making transgenic cells and organisms (Genetic engineering)

Classes of Recombination
Classes of recombination Homologous Recombination Site-specific Recombination Transposition/Replicative Recombination
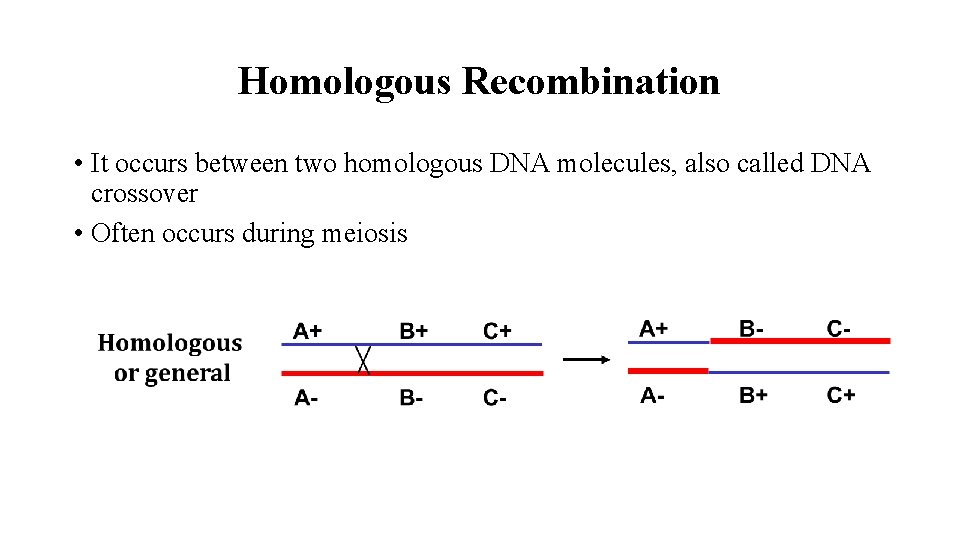
Homologous Recombination • It occurs between two homologous DNA molecules, also called DNA crossover • Often occurs during meiosis

It is also called as General Recombination. Homologous Recombination During meiosis, two homologous pairs of sister chromatids align side by side. The DNA crossover is very likely to occur. It could be as often as several times per meiosis.


Characteristics of Homologous Recombination

1. Two homologous DNA molecules that were originally part of different chromosomes “cross over; ” Their double helices break, and the two broken ends join to their opposite partners to re-form two intact double helices, each composed of parts of the two initial DNA molecules

2. The site of exchange can occur anywhere in the homologous nucleotide sequences of the two participating DNA molecules. Characteristics of homologous recombination 3. At the site of exchange, a strand of one DNA molecule has become basepaired to a strand of the second DNA molecule to create a heteroduplex joint that links the two double helices
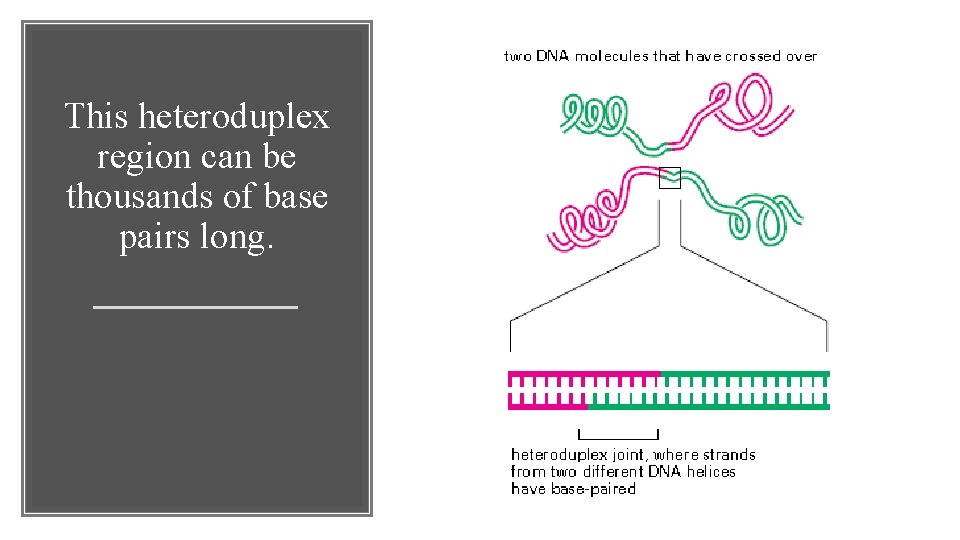
This heteroduplex region can be thousands of base pairs long.
4. No nucleotide sequences are altered at the site of exchange. Characteristics of homologous recombination The cleavage and rejoining events occur so precisely that not a single nucleotide is lost or gained.

Mechanism of homologous recombination

Mechanism of homologous recombination • Recombination is initiated by a special endonuclease that simultaneously cuts both strands of the double helix, creating a complete break in the DNA molecule. • The 5’ ends at the break are then chewed back by an exonuclease, creating protruding single-stranded 3’ ends. • These single strands search for a homologous DNA helix with which to pair—leading to the formation of a “joint molecule” between a maternal and a paternal chromosome.

Homologous Recombination in prokaryotes
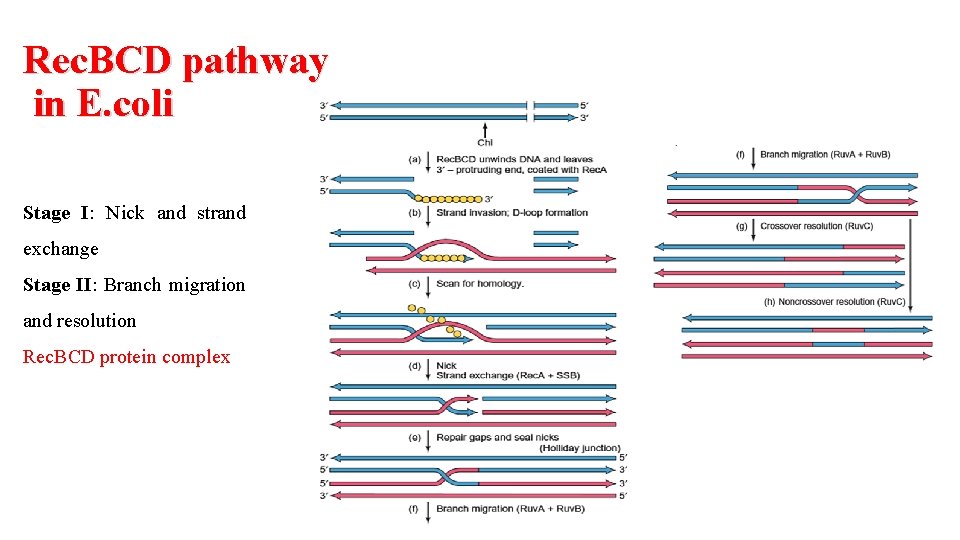
Rec. BCD pathway in E. coli Stage I: Nick and strand exchange Stage II: Branch migration and resolution Rec. BCD protein complex
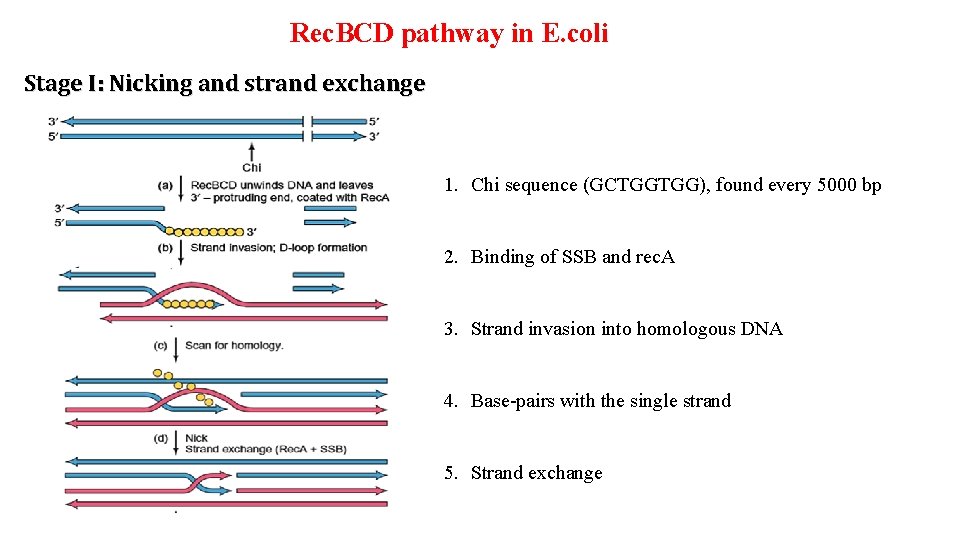
Rec. BCD pathway in E. coli Stage I: Nicking and strand exchange 1. Chi sequence (GCTGGTGG), found every 5000 bp 2. Binding of SSB and rec. A 3. Strand invasion into homologous DNA 4. Base-pairs with the single strand 5. Strand exchange
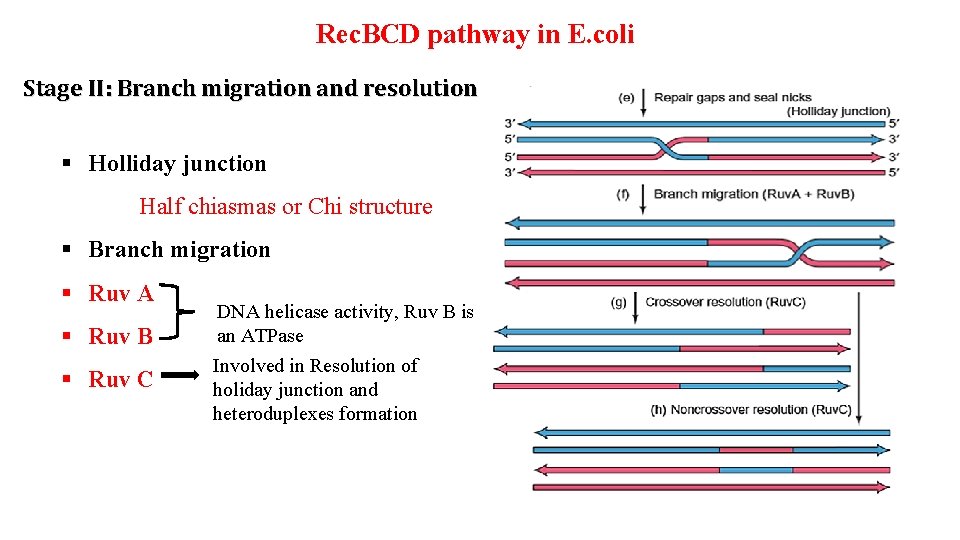
Rec. BCD pathway in E. coli Stage II: Branch migration and resolution § Holliday junction Half chiasmas or Chi structure § Branch migration § Ruv A § Ruv B DNA helicase activity, Ruv B is an ATPase § Ruv C Involved in Resolution of holiday junction and heteroduplexes formation
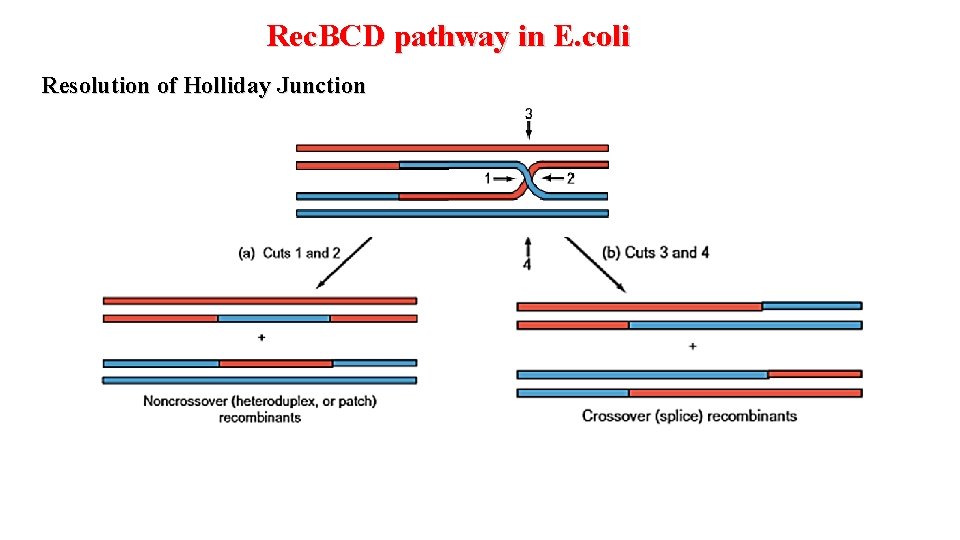
Rec. BCD pathway in E. coli Resolution of Holliday Junction
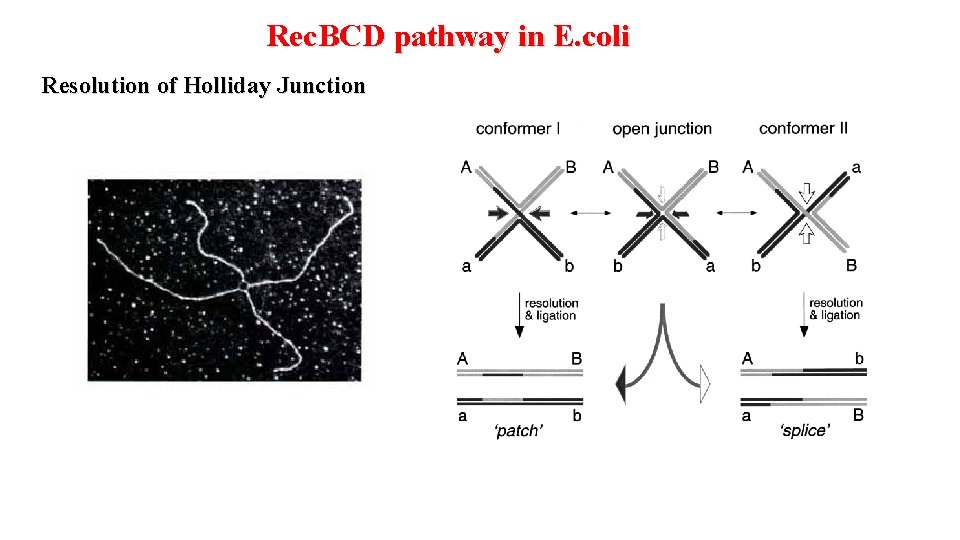
Rec. BCD pathway in E. coli Resolution of Holliday Junction

Holliday Junction Resolution https: //www. youtube. com/watch? v=5638 I 3 f. Gu. W 4 https: //www. youtube. com/watch? v=Mvn. Wx. N 81 Qps

Rec. BCD Enzyme complex Rec. BCD has: § Endonuclease subunits (rec. BC) that cut one DNA strand close to Chi sequence 5’-GCTGGTGGGTT*G*CCT-3’ § DNA helicase activity (rec. D subunit) § DNA-dependent ATPase activity, unwinds DNA to generate the 3’ single stranded tails 10/29/2020
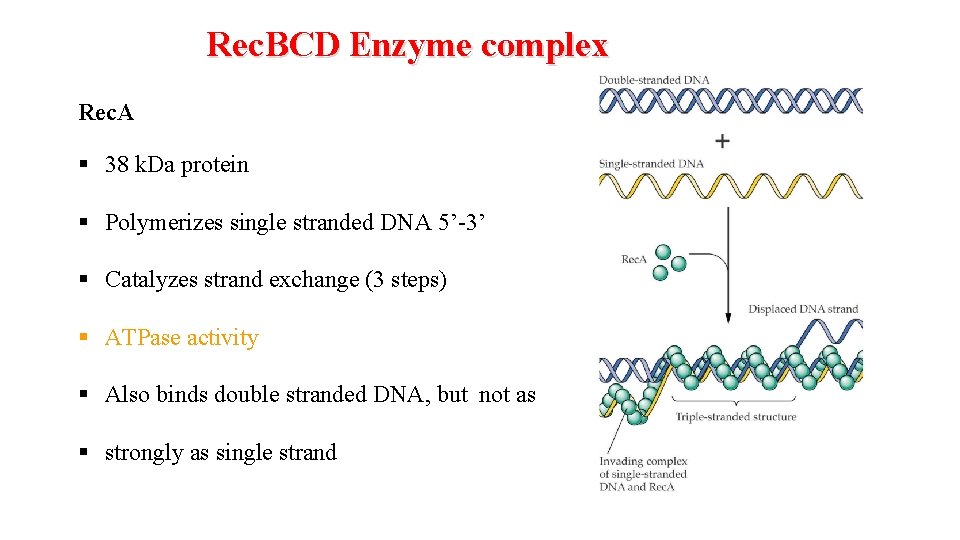
Rec. BCD Enzyme complex Rec. A § 38 k. Da protein § Polymerizes single stranded DNA 5’-3’ § Catalyzes strand exchange (3 steps) § ATPase activity § Also binds double stranded DNA, but not as § strongly as single strand

Rec. BCD Enzyme complex Ruv. A and Ruv. B § DNA helicase that drive branch migration § Ruv. A ü Tetramer protein binds to Holliday junction ü Square planar conformation § Ruv. B ü 2 copies of Ruv. B bind at the Holliday junction ü Ruv. B is a hexamer ring, has helicase & ATPase activity § Branch migration is in the direction of rec. A mediated strand-exchange
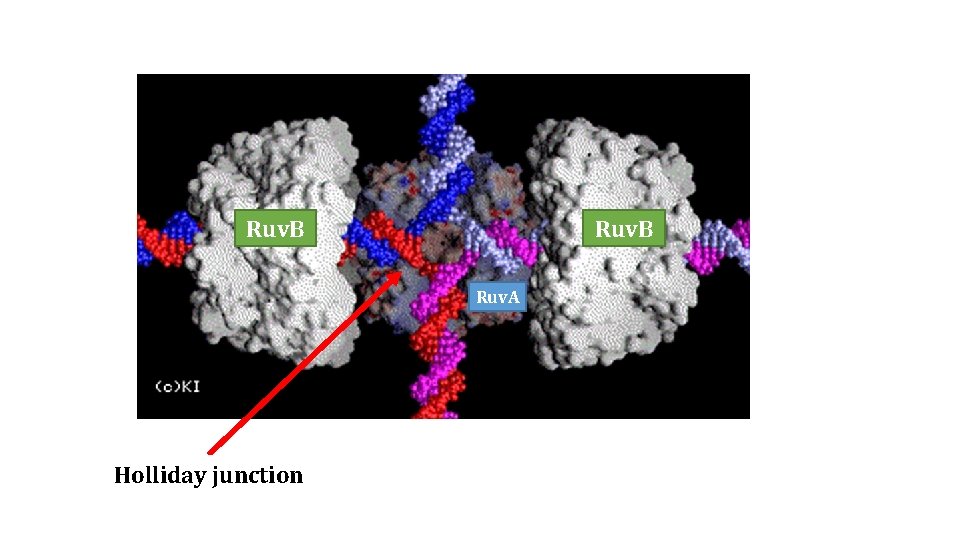
Ruv. B Ruv. A Holliday junction
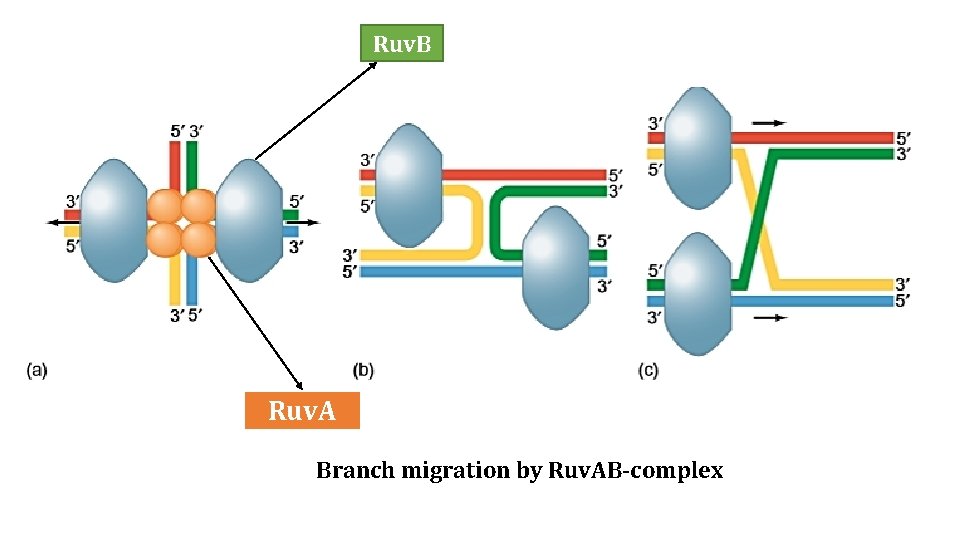
Ruv. B Ruv. A Branch migration by Ruv. AB-complex

Rec. BCD Enzyme complex Ruv. C § Resolvase § Endonuclease cut 2 strands of HJ § Bind as a dimer to the HJ § Recognize consensus sequences 5’-(A/T)TT ↓ (G/C)-3’ § Work with Ruv A and Ruv B as a complex § Frequently present in E. coli genome

- Slides: 35