DNA History Structure Function DNA Deoxyribonucleic Acid Functions

DNA History Structure Function

DNA Deoxyribonucleic Acid Functions to control the production of proteins within the cell thus controlling all chemical processes within the cell

Discovery of DNA Function Fred Griffith Transformation Experiments Transfer of hereditary material from dead S cells (virulent) to living R cells (nonvirulent) of a pneumonia-causing bacterium Concluded something from the S cells (proteins or nucleic acids) had transformed the R cells
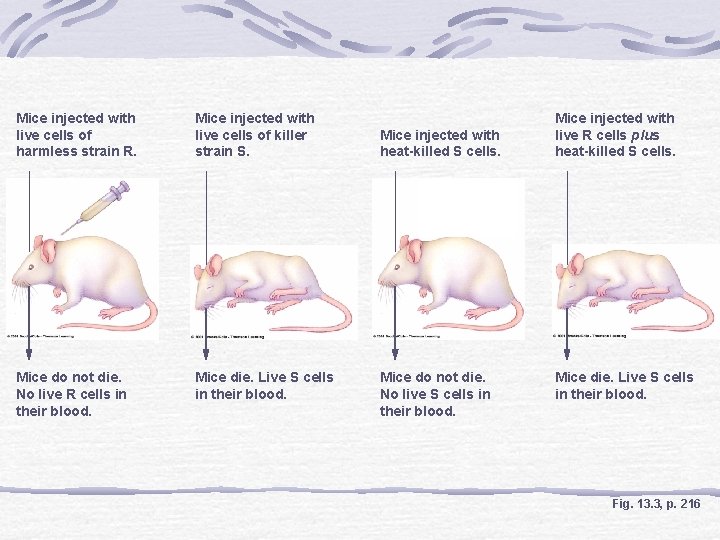
Mice injected with live cells of harmless strain R. Mice injected with live cells of killer strain S. Mice do not die. No live R cells in their blood. Mice die. Live S cells in their blood. Mice injected with heat-killed S cells. Mice do not die. No live S cells in their blood. Mice injected with live R cells plus heat-killed S cells. Mice die. Live S cells in their blood. Fig. 13. 3, p. 216

Discovery of DNA Oswald Avery Showed that Griffith’s “substance” was DNAase blocked the transformation

Discovery of DNA Function Hershey-Chase Bacteriophage injects DNA--not protein-into bacterium Radioisotope labeled protein and DNA of bacteriophage Radioactive phosphorus is detected inside of bacteria
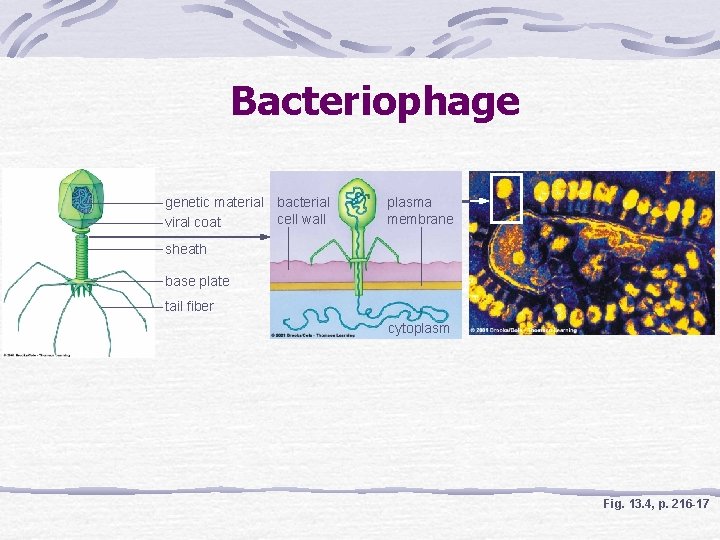
Bacteriophage genetic material bacterial cell wall viral coat plasma membrane sheath base plate tail fiber cytoplasm Fig. 13. 4, p. 216 -17
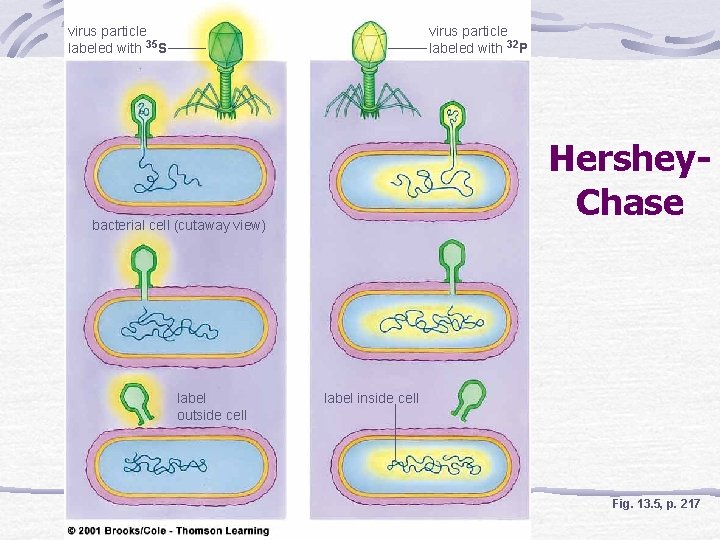
virus particle labeled with 35 S virus particle labeled with 32 P Hershey. Chase bacterial cell (cutaway view) label outside cell label inside cell Fig. 13. 5, p. 217

Nature of the Genetic Material Property 1 - it must contain, in a stable form, information encoding the organism’s structure, function, development and reproduction Property 2 - it must replicate accurately so progeny cells have the same genetic makeup Property 3 - it must be capable of some variation (mutation) to permit evolution

Discovery of DNA Rosalind Franklin used x-ray defraction to predict structure

X-ray diffraction photograph of the DNA double helix

Discovery of DNA James Watson and Francis Crick 1953 described the helical structure of DNA Double helix 2 strands of hydrogen bonded nucleotides twisted around a central axis

1 6 2 7 13 14 19 20 3 8 4 9 15 10 16 21 22 5 11 12 17 18 X Y

Structure of a nucleotide A nucleotide is made of 3 components: A Pentose sugar This is a 5 carbon sugar The sugar in DNA is deoxyribose. The sugar in RNA is ribose.
Structure of a nucleotide A Phosphate groups are important because they link the sugar on one nucleotide onto the phosphate of the next nucleotide to make a polynucleotide.
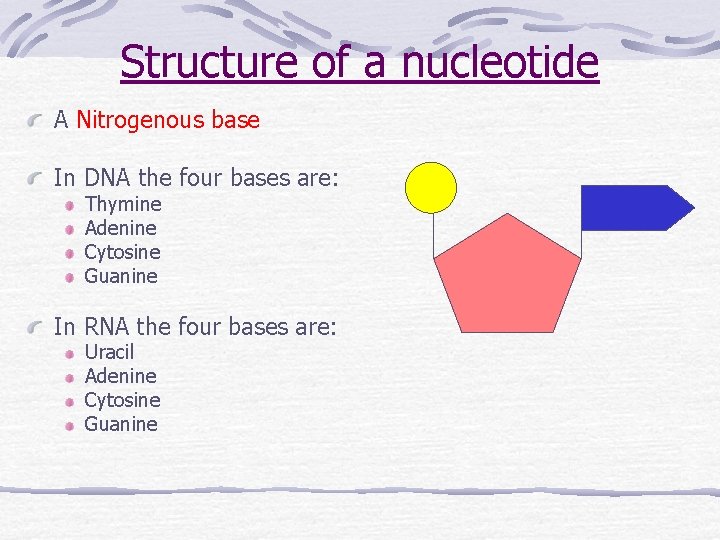
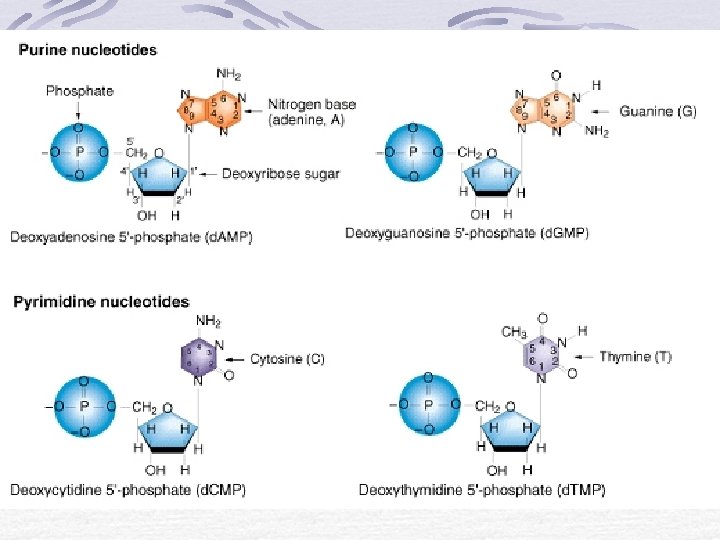
Structure of a nucleotide A Nitrogenous base In DNA the four bases are: Thymine Adenine Cytosine Guanine In RNA the four bases are: Uracil Adenine Cytosine Guanine

Nitrogenous bases Two types Pyramidines Purines Thymine - T Cytosine - C Uracil - U Adenine - A Guanine - G
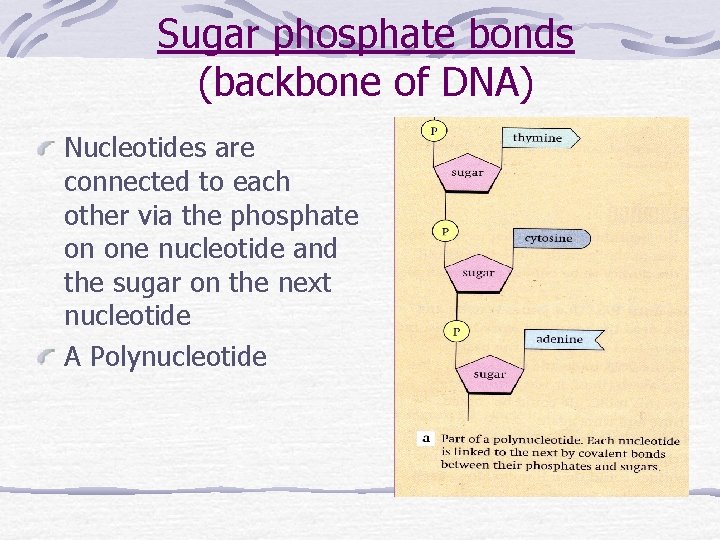
Sugar phosphate bonds (backbone of DNA) Nucleotides are connected to each other via the phosphate on one nucleotide and the sugar on the next nucleotide A Polynucleotide


Patterns of Base Pairing DNA consists of two strands of nucleotides held together at bases by hydrogen bonds ü Purines bond to Pyrimidines This is because there is exactly enough room for one purine and one pyrimidine base between the two polynucleotide strands of DNA. Amount of A=T and C=G

Double Helix of DNA

DNA Replication and Repair DNA helicase causes the two strands of the double helix to unwind at one "origin" (viral/bacterial) or many origins (eukaryotic); Unwinding and assembly proceeding simultaneously in both directions at replication forks exposing the bases of the two strands of DNA

DNA Replication and Repair Unattached nucleotides pair with exposed bases DNA polymerases assemble the nucleotides into nucleic acids and "proofread" the new bases for mismatched pairs, which are replaced with correct bases DNA ligase seals the bases up Gyrase winds the DNA molecules Replication resulting in 2 molecules that consist of one "old" strand one "new" strand
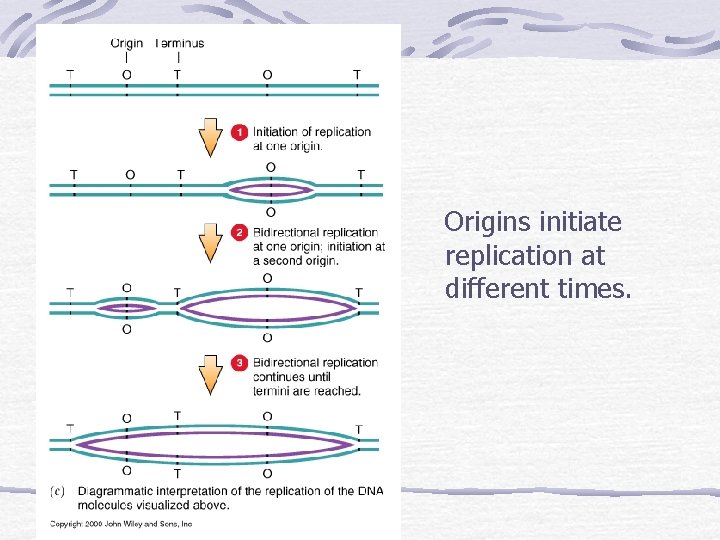
Origins initiate replication at different times.

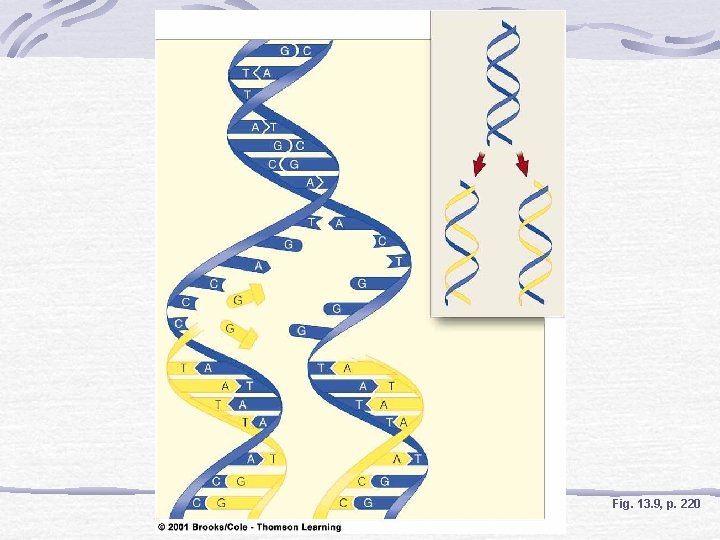
Fig. 13. 9, p. 220
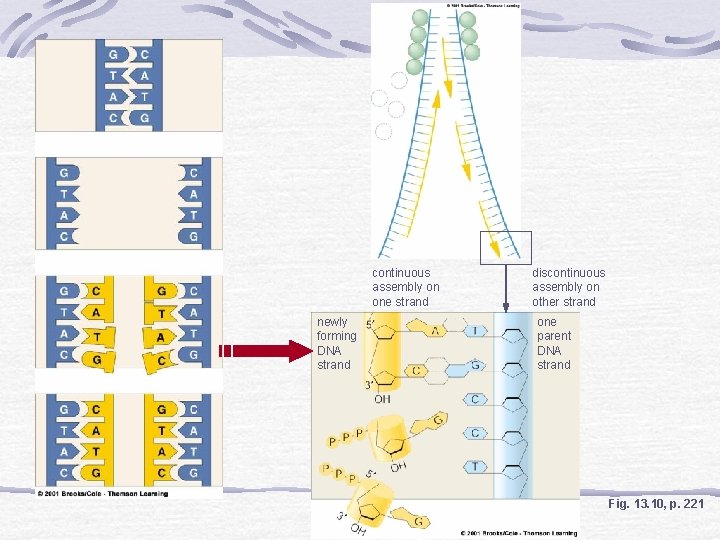
continuous assembly on one strand newly forming DNA strand discontinuous assembly on other strand one parent DNA strand Fig. 13. 10, p. 221

Replication of DNA and Chromosomes Speed of DNA replication: 3, 000 nucleotides/min in human 30, 000 nucleotides/min in E. coli Accuracy of DNA replication: Very precise (1 error/1, 000, 000 nt)

Questions? Of the 4 bases, which other base does adenine most closely resemble? List the 4 different nucleotides. Which 2 molecules of a nucleotide form the sides of a DNA ladder?


Questions? If 30% of a DNA molecule is Adenine, what percent is Cytosine? What does the term replication mean? What is another name for adenine, ribose and three phosphate molecules attached to it?

From DNA to Proteins

The Three Classes of RNA Messenger RNA (m. RNA) carries the "blueprint" for protein assembly to the ribosomes Ribosome RNA (r. RNA) combines with proteins to form ribosomes upon which polypeptides are assembled Transfer RNA (t. RNA) brings the correct amino acid to the ribosome and pairs up with an m. RNA code for that amino acid

Structure of an RNA Nucleotide RNA composed of nucleotides Ribose Phosphate group Bases Adenine Cytosine Guanine Uracil
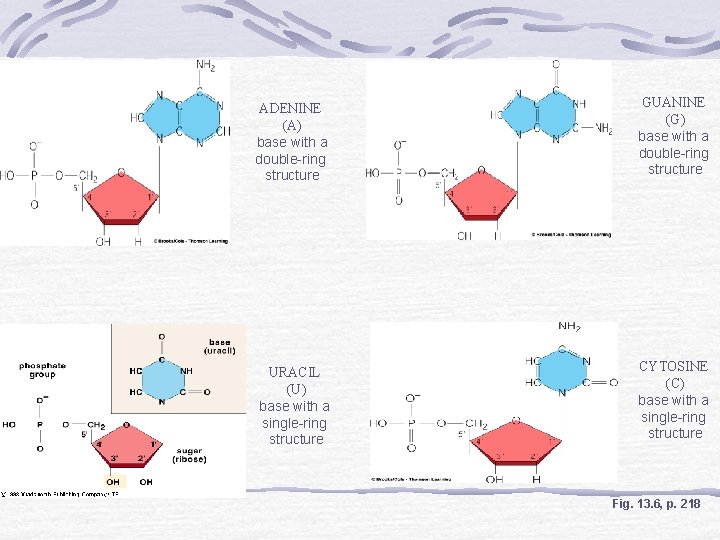
ADENINE (A) base with a double-ring structure GUANINE (G) base with a double-ring structure URACIL (U) base with a single-ring structure CYTOSINE (C) base with a single-ring structure Fig. 13. 6, p. 218
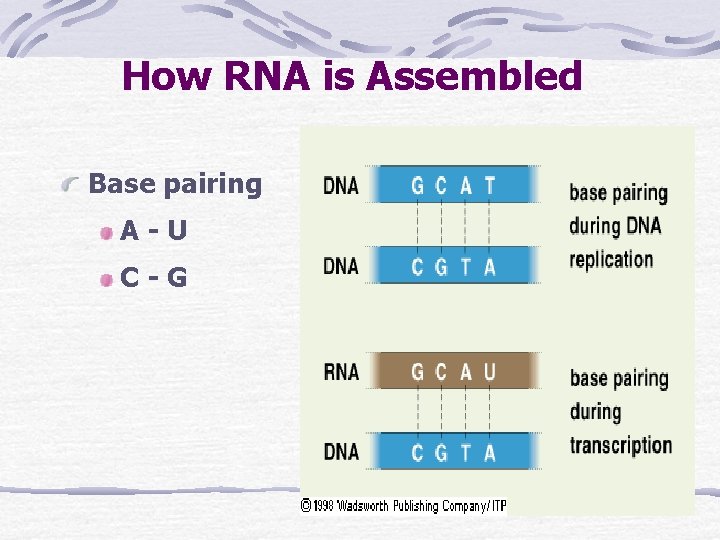
How RNA is Assembled Base pairing A-U C-G

Transcription Nucleotide assembled Portion of DNA serves as template (5’ ---> 3’ direction) RNA polymerase 3 types in eukaryotes to make the 3 types of RNA Promoter signals start of a gene

Transcription Helicase unwinds the DNA molecule RNA polymerase matches down a RNA base with the appropriate DNA base Ligase seals the RNA strand RNA now leaves the DNA molecule and the nucleus Gyrase winds up the DNA strand
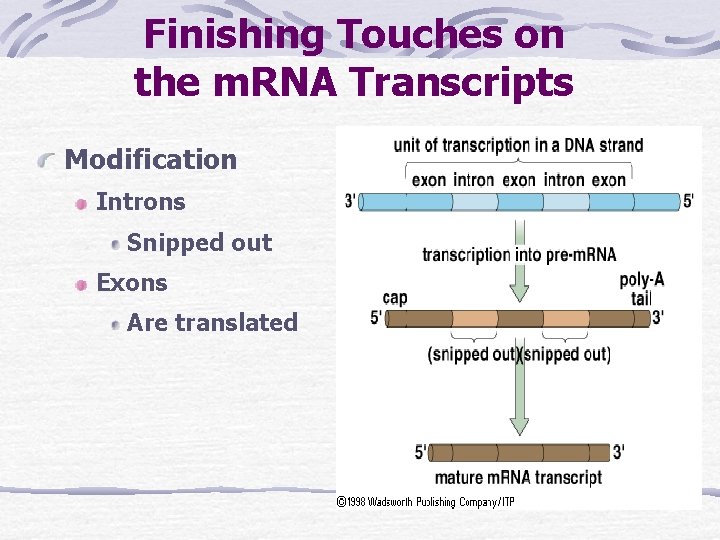
Finishing Touches on the m. RNA Transcripts Modification Introns Snipped out Exons Are translated
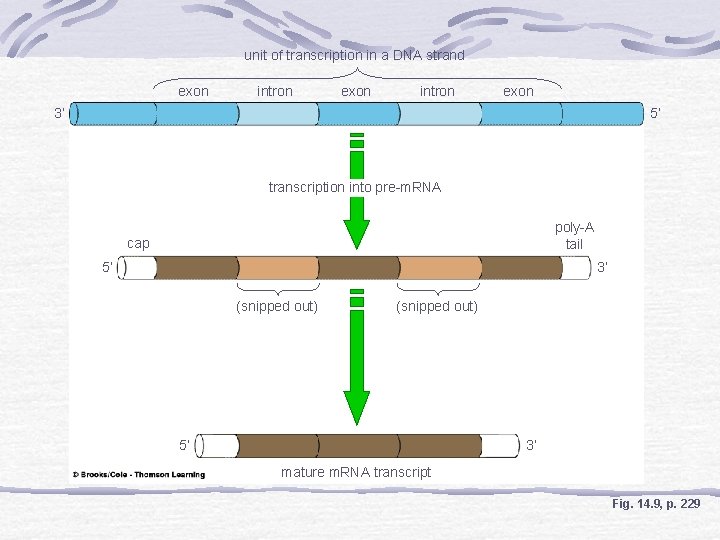
unit of transcription in a DNA strand exon intron exon 3’ 5’ transcription into pre-m. RNA poly-A tail cap 5’ 3’ (snipped out) 5’ 3’ mature m. RNA transcript Fig. 14. 9, p. 229

An Overview • DNA unwinds • m. RNA copy is made of one of the DNA strands. • m. RNA copy moves out of nucleus into cytoplasm. • t. RNA molecules are activated as their complementary amino acids are attached to them.

An Overview • m. RNA copy attaches to the small subunit of the ribosomes in cytoplasm. 6 of the bases in the m. RNA are exposed in the ribosome. • A t. RNA bonds complementarily with the m. RNA via its anticodon. • A second t. RNA bonds with the next three bases of the m. RNA, the amino acid joins onto the amino acid of the first t. RNA via a peptide bond.

An Overview • The ribosome moves along. The first t. RNA leaves the ribosome. • A third t. RNA brings a third amino acid • Eventually a stop codon is reached on the m. RNA. The newly synthesised polypeptide leaves the ribosome.

The Genetic Code Codons On m. RNA Triplets RNA polymerases read nucleotide bases Cells have 64 kinds of codons Only 20 amino acids
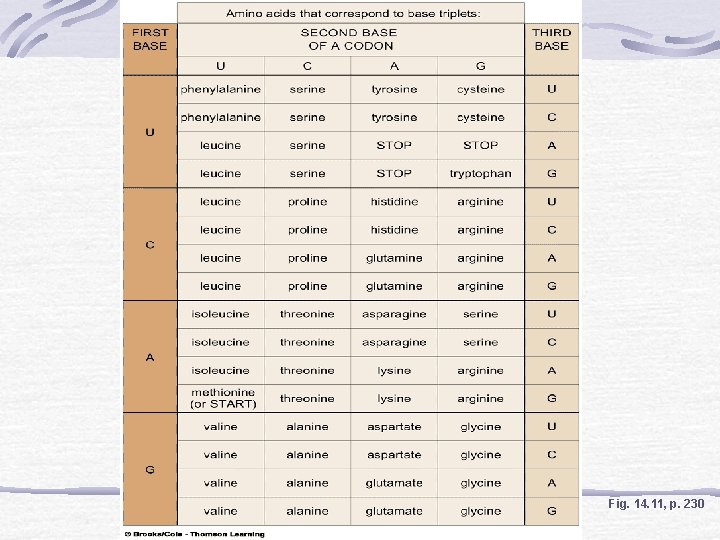
Fig. 14. 11, p. 230

Protein Synthesis: An Example DNA Sequence: TAC GGA GGT TGA ACT m. RNA: AUG CCU CCA ACU UGA t. RNA: UAC GGA GGU UGA ACU Amino Acids: met – pro - pro – thr - stop

t. RNA Role of t. RNA Attachment site for amino acid Anticodon
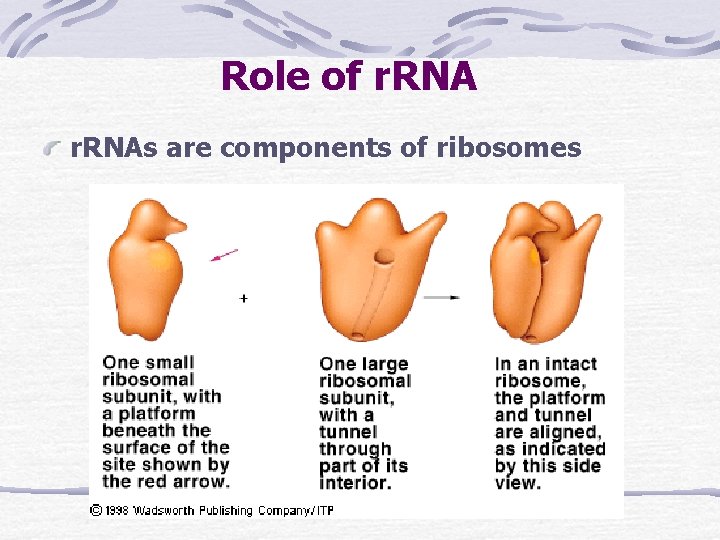
Role of r. RNAs are components of ribosomes
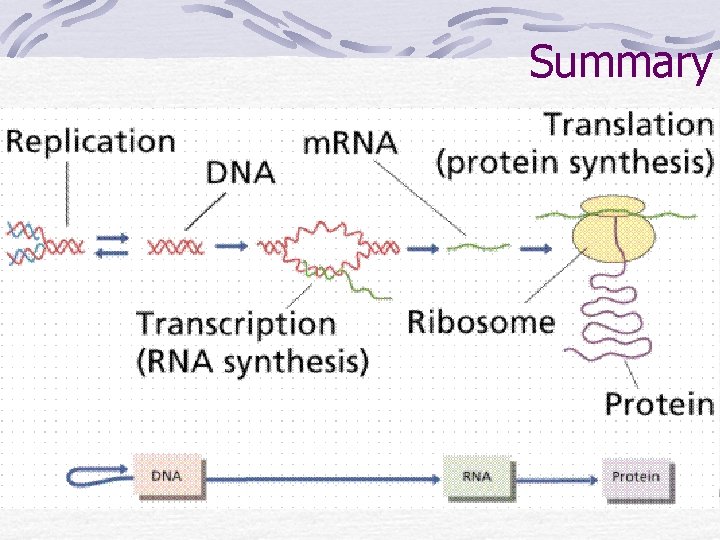
Summary

Translation Initiation t. RNA and m. RNA are loaded onto a ribosome Elongation Polypeptide chain forms as m. RNA passes between ribosome subunits Termination Stop codon and detachment of m. RNA and polypeptide chain
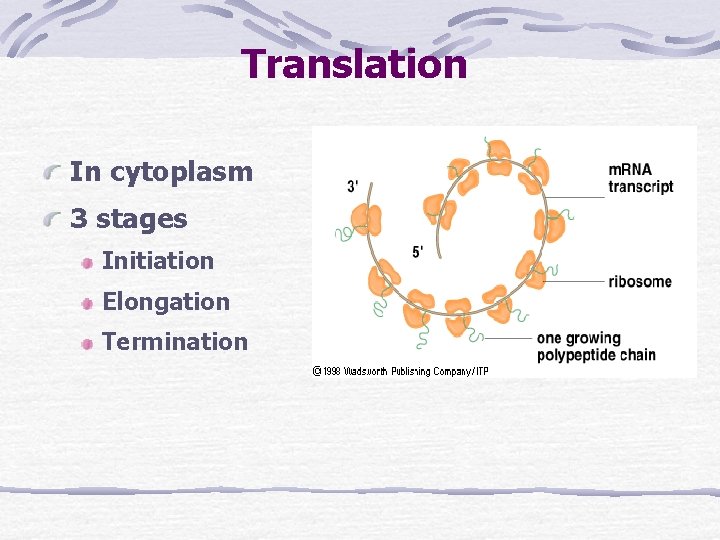
Translation In cytoplasm 3 stages Initiation Elongation Termination

m. RNA attaches to small ribosomal subunit

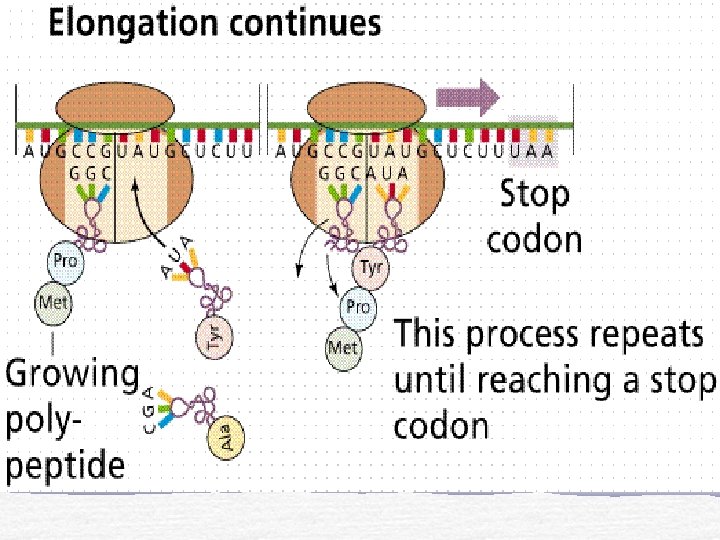
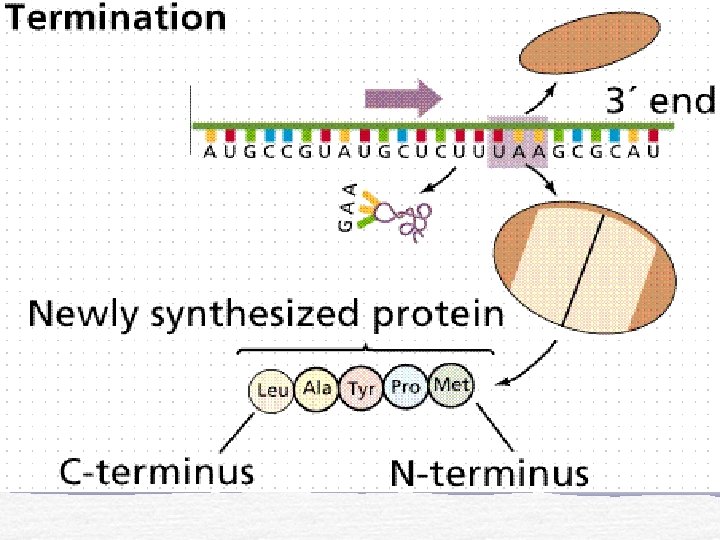
End of Translation

Translation - Animation



One Gene, One Polypeptide Sickle Cell Anemia Hemoglobin S and A Gene mutation Affects protein synthesis Amino acid sequences of polypeptide chains are encoded in genes
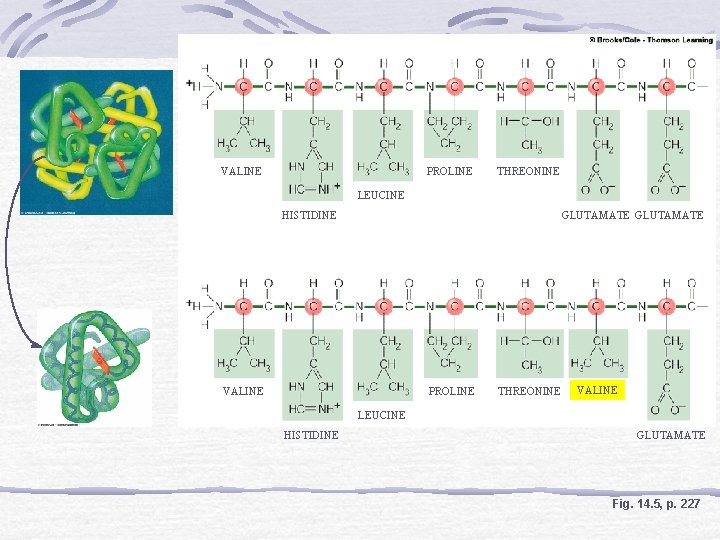
VALINE PROLINE THREONINE LEUCINE HISTIDINE GLUTAMATE VALINE PROLINE THREONINE VALINE LEUCINE HISTIDINE GLUTAMATE Fig. 14. 5, p. 227
- Slides: 61