DNA BASICS History and Structure History of DNA

DNA BASICS History and Structure

History of DNA � � Early scientists thought protein was the cell’s hereditary material because it was more complex than DNA Proteins are composed of 20 different amino acids in long polypeptide chains

History of DNA Griffith and Transformation In 1928, British scientist Fredrick Griffith was trying to learn how certain types of bacteria caused pneumonia. He isolated two different strains of pneumonia bacteria from mice and grew them in his lab.
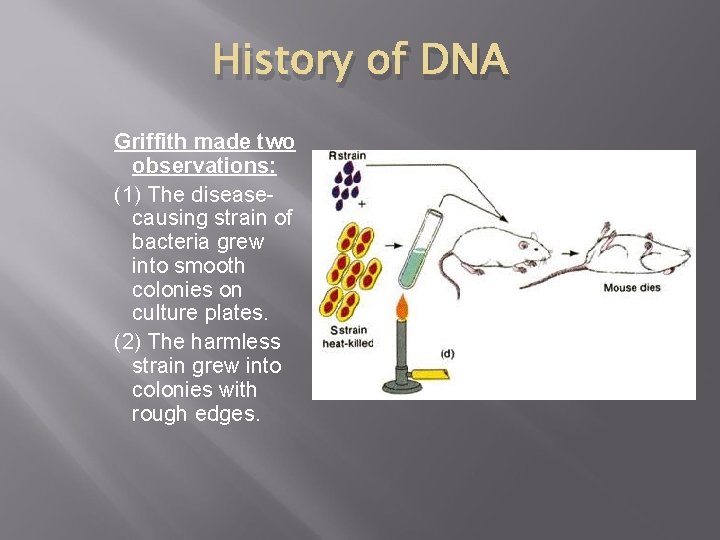
History of DNA Griffith made two observations: (1) The diseasecausing strain of bacteria grew into smooth colonies on culture plates. (2) The harmless strain grew into colonies with rough edges.
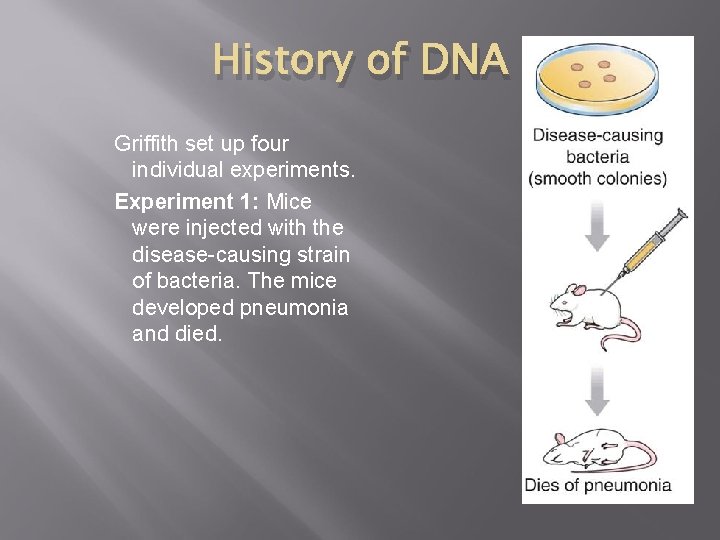
History of DNA Griffith set up four individual experiments. Experiment 1: Mice were injected with the disease-causing strain of bacteria. The mice developed pneumonia and died.
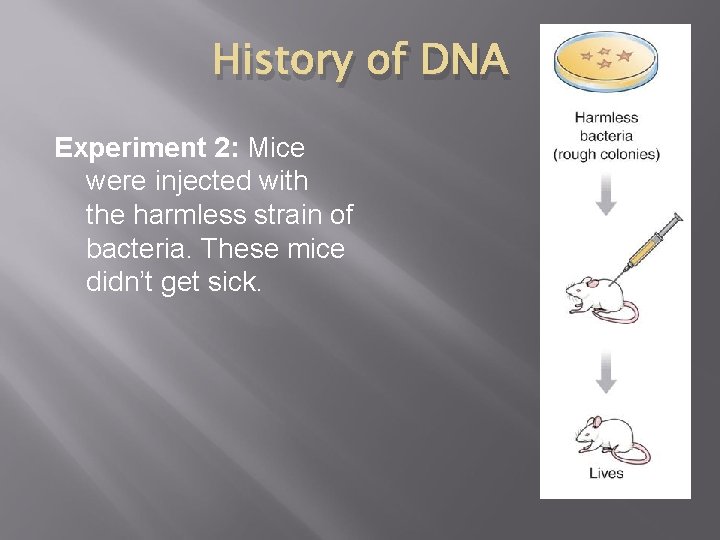
History of DNA Experiment 2: Mice were injected with the harmless strain of bacteria. These mice didn’t get sick.
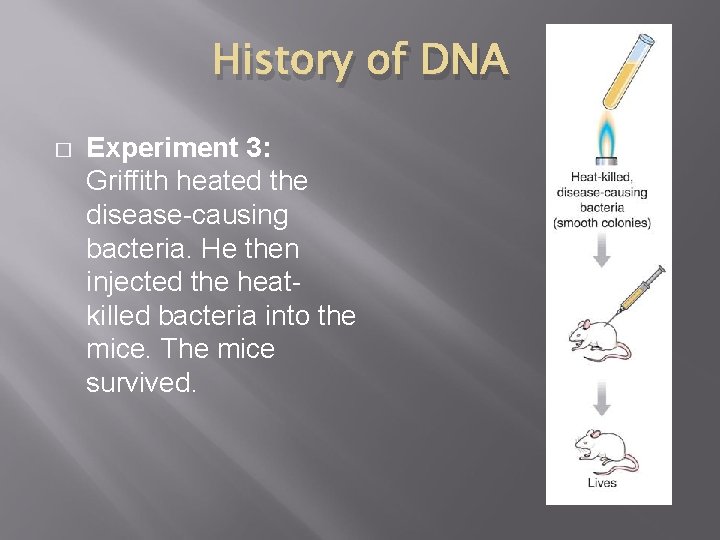
History of DNA � Experiment 3: Griffith heated the disease-causing bacteria. He then injected the heatkilled bacteria into the mice. The mice survived.
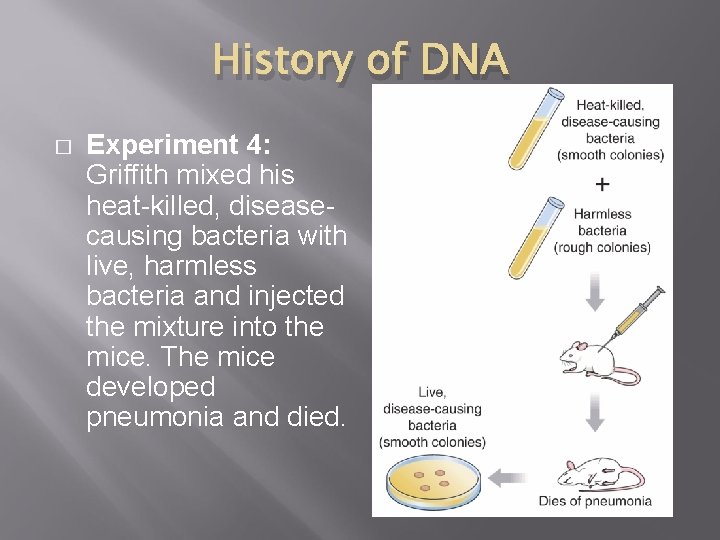
History of DNA � Experiment 4: Griffith mixed his heat-killed, diseasecausing bacteria with live, harmless bacteria and injected the mixture into the mice. The mice developed pneumonia and died.
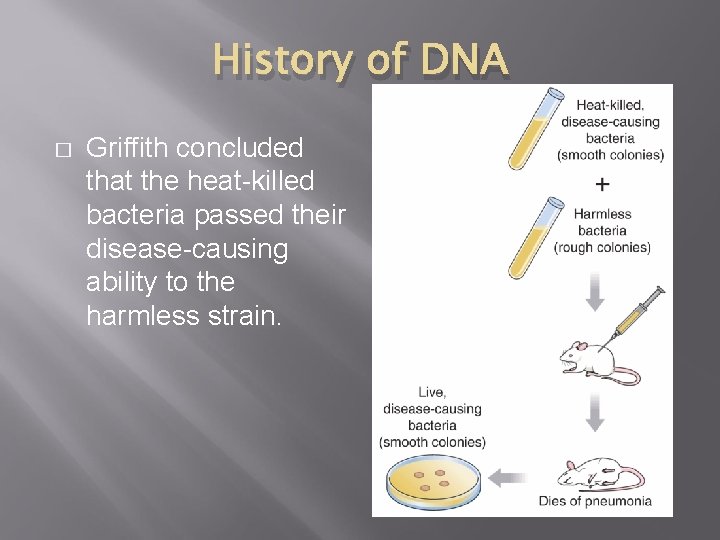
History of DNA � Griffith concluded that the heat-killed bacteria passed their disease-causing ability to the harmless strain.

History of DNA � � Griffith called this process transformation because one strain of bacteria (the harmless strain) had changed permanently into another (the disease-causing strain). Griffith hypothesized that a factor must contain information that could change harmless bacteria into disease-causing ones.

History of DNA Avery and DNA Oswald Avery repeated Griffith’s work to determine which molecule was most important for transformation. Avery and his colleagues made an extract from the heat -killed bacteria that they treated with enzymes.

History of DNA The enzymes destroyed proteins, lipids, carbohydrates, and other molecules, including the nucleic acid RNA. Transformation still occurred.

History of DNA Avery and other scientists repeated the experiment using enzymes that would break down DNA. When DNA was destroyed, transformation did not occur. Therefore, they concluded that DNA was the transforming factor.

History of DNA The Hershey-Chase Experiment Alfred Hershey and Martha Chase studied viruses— nonliving particles smaller than a cell that can infect living organisms.
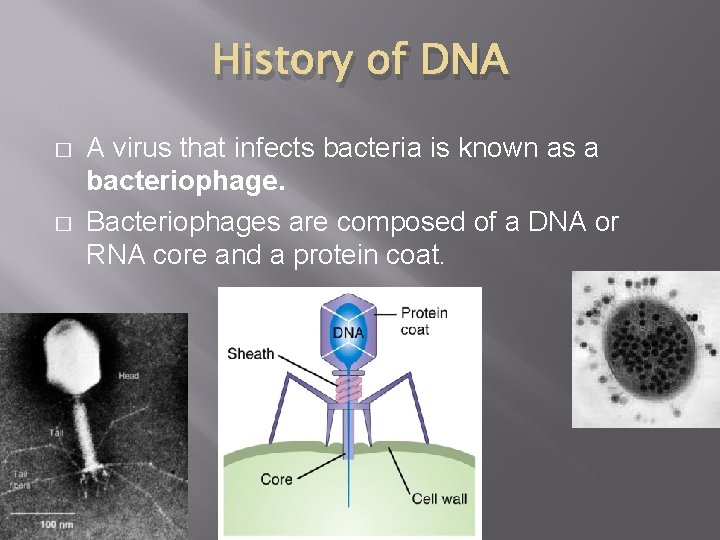
History of DNA � � A virus that infects bacteria is known as a bacteriophage. Bacteriophages are composed of a DNA or RNA core and a protein coat.
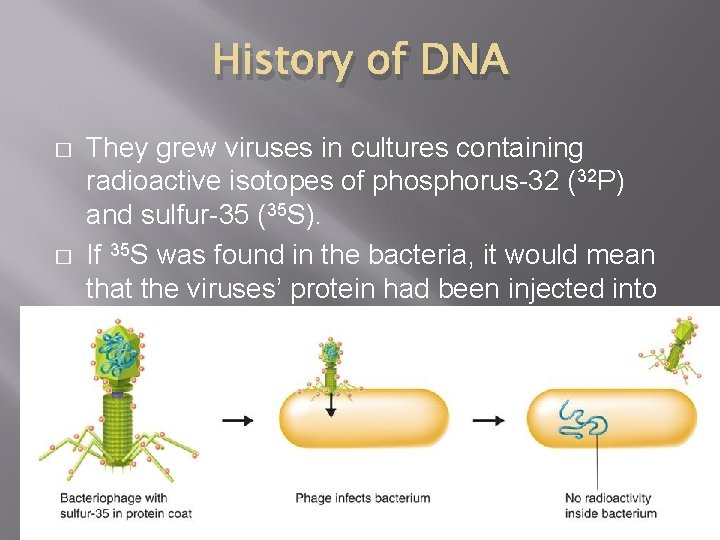
History of DNA � � They grew viruses in cultures containing radioactive isotopes of phosphorus-32 (32 P) and sulfur-35 (35 S). If 35 S was found in the bacteria, it would mean that the viruses’ protein had been injected into the bacteria.
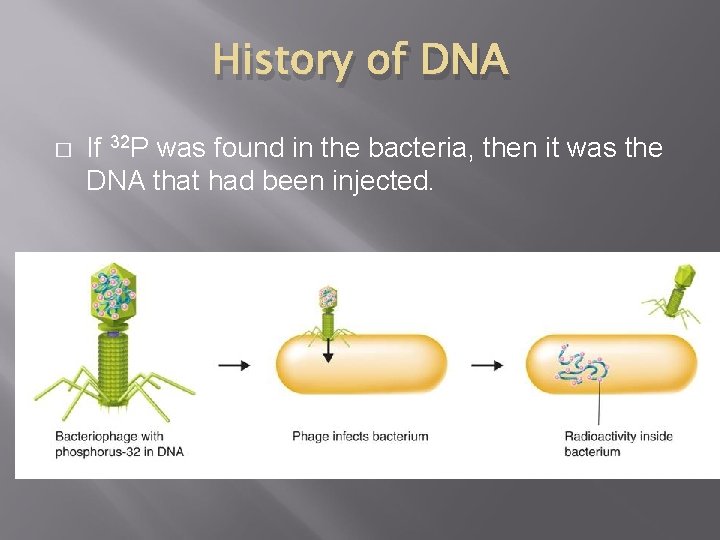
History of DNA � If 32 P was found in the bacteria, then it was the DNA that had been injected.

History of DNA � � Nearly all the radioactivity in the bacteria was from phosphorus (32 P). Hershey and Chase concluded that the genetic material of the bacteriophage was DNA, not protein.

History of DNA The Components and Structure of DNA is made up of nucleotides. A nucleotide is a monomer of nucleic acids made up of: Deoxyribose – 5 -carbon Sugar Phosphate Group Nitrogenous Base
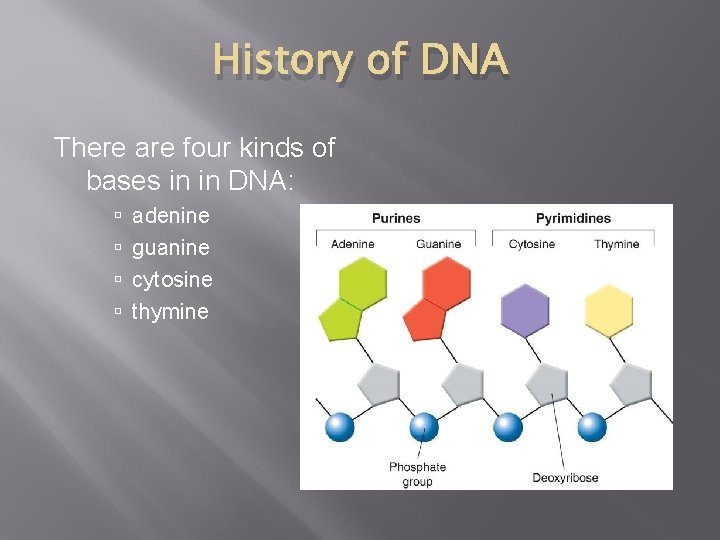
History of DNA There are four kinds of bases in in DNA: adenine guanine cytosine thymine

History of DNA � Chargaff's Rules Erwin Chargaff discovered that: The percentages of guanine [G] and cytosine [C] bases are almost equal in any sample of DNA. The percentages of adenine [A] and thymine [T] bases are almost equal in any sample of DNA.
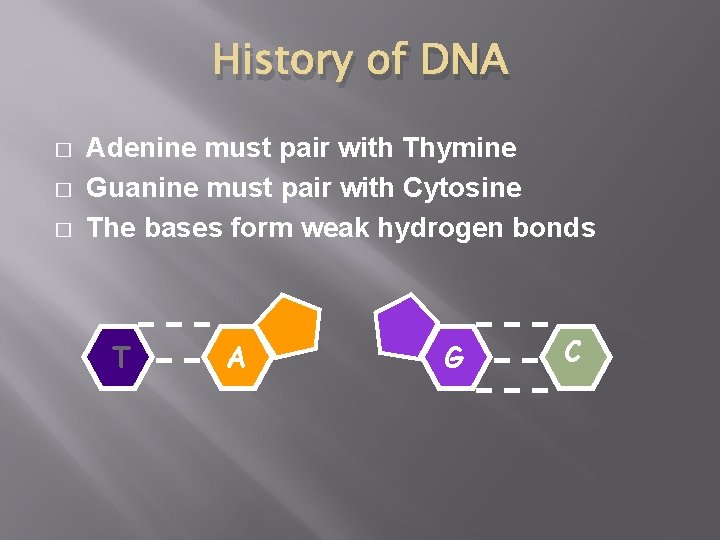
History of DNA � � � Adenine must pair with Thymine Guanine must pair with Cytosine The bases form weak hydrogen bonds T A G C

History of DNA � X-Ray Evidence Rosalind Franklin used X-ray diffraction to get information about the structure of DNA. She aimed an X-ray beam at concentrated DNA samples and recorded the scattering pattern of the X-rays on film.

History of DNA � Using clues from Franklin’s pattern, James Watson and Francis Crick built a model that explained how DNA carried information and could be copied. � Watson and Crick's model of DNA was a double helix, in which two strands were wound around each other.
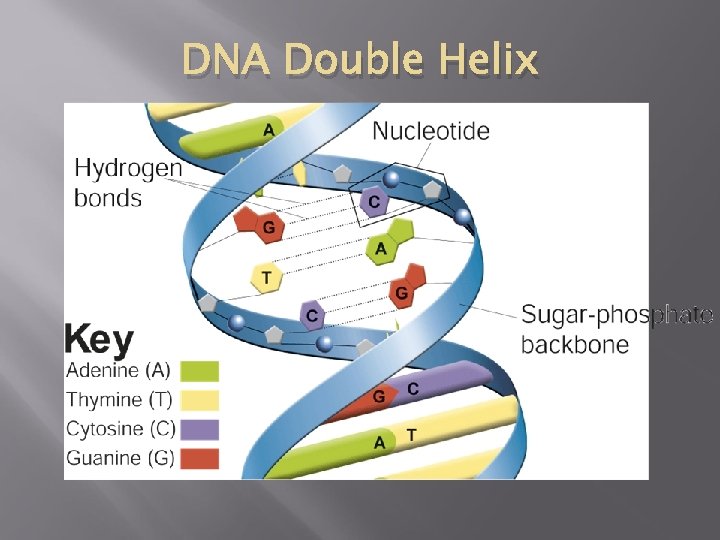
DNA Double Helix

STRUCTURE OF DNA
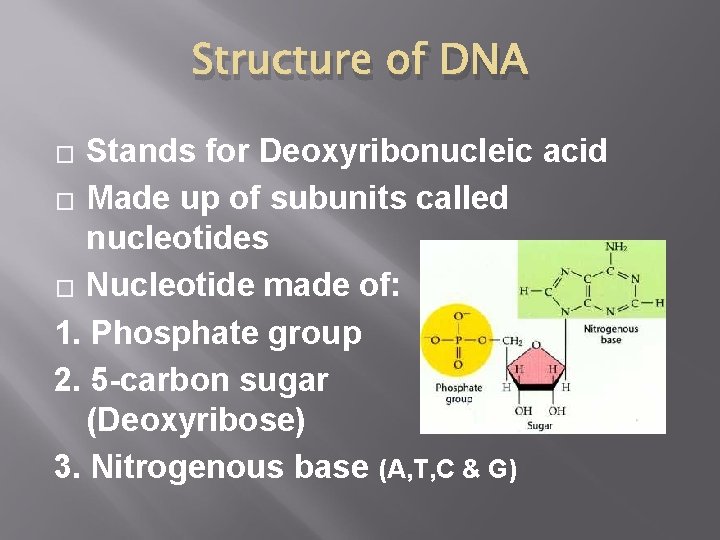
Structure of DNA Stands for Deoxyribonucleic acid � Made up of subunits called nucleotides � Nucleotide made of: 1. Phosphate group 2. 5 -carbon sugar (Deoxyribose) 3. Nitrogenous base (A, T, C & G) �
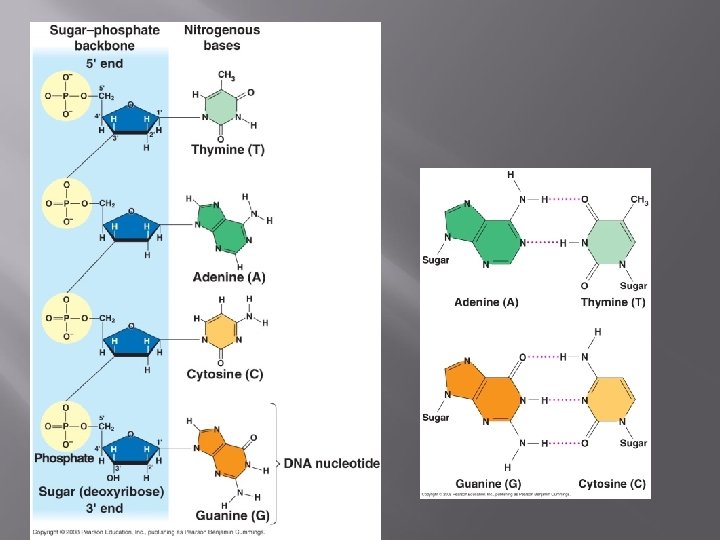

Structure of DNA � � � Two strands coiled called a double helix Sides made of a pentose (5 carbon) sugar deoxyribose bonded to phosphate (PO 4) groups – for support Center made of nitrogen bases bonded together by weak hydrogen bonds – genetic information (A, T, C & G).
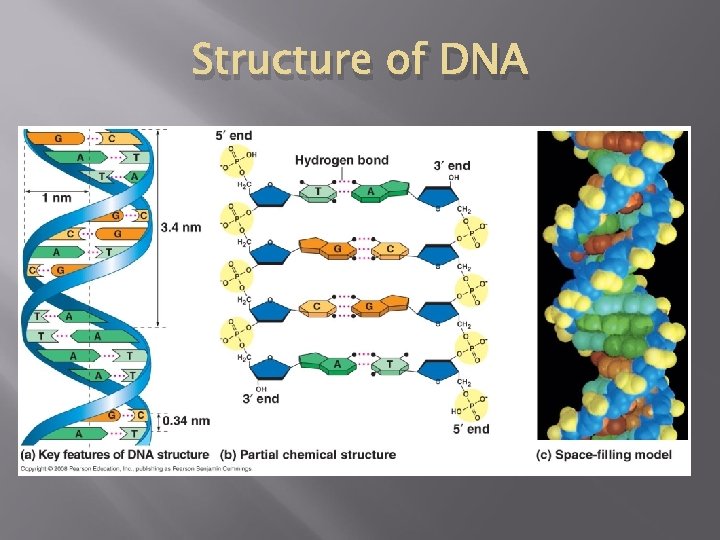
Structure of DNA
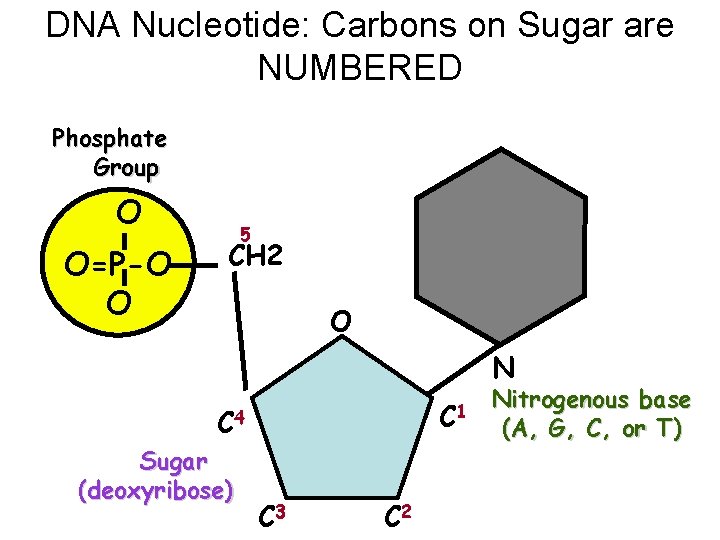
DNA Nucleotide: Carbons on Sugar are NUMBERED Phosphate Group O O=P-O O 5 CH 2 O N C 1 C 4 Sugar (deoxyribose) C 3 C 2 Nitrogenous base (A, G, C, or T)
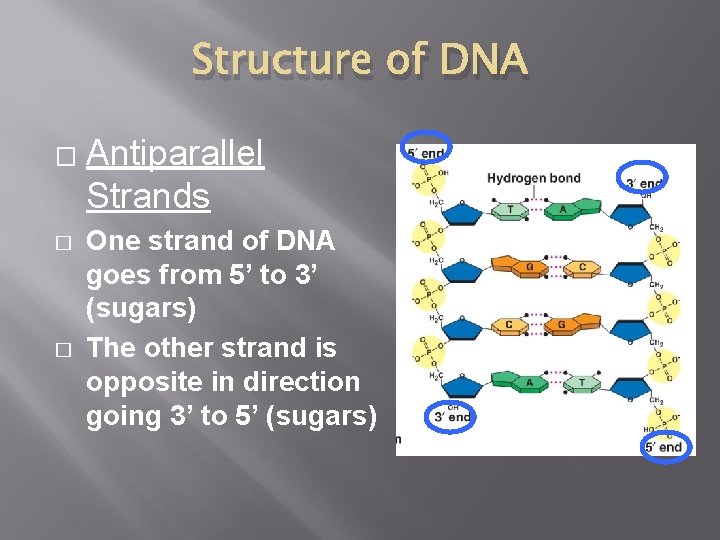
Structure of DNA � � � Antiparallel Strands One strand of DNA goes from 5’ to 3’ (sugars) The other strand is opposite in direction going 3’ to 5’ (sugars)
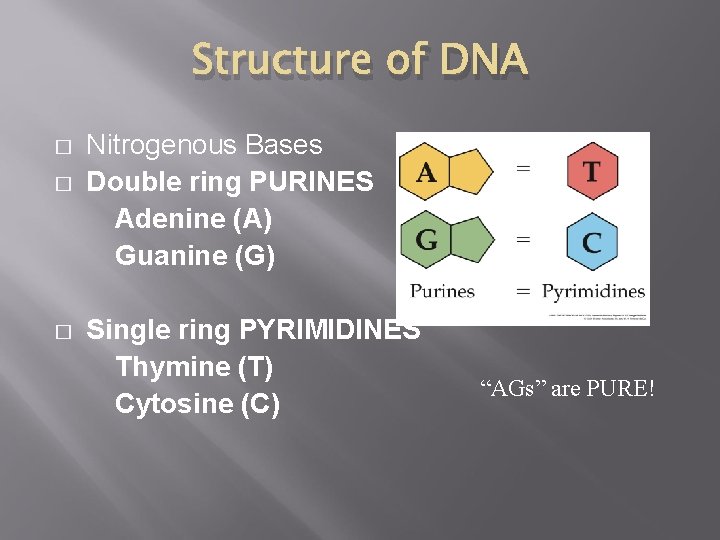
Structure of DNA � � � Nitrogenous Bases Double ring PURINES Adenine (A) Guanine (G) Single ring PYRIMIDINES Thymine (T) Cytosine (C) “AGs” are PURE!
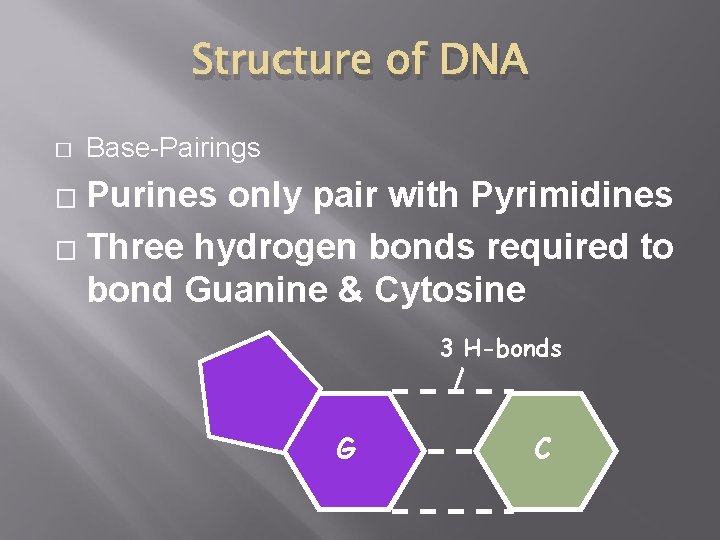
Structure of DNA � Base-Pairings Purines only pair with Pyrimidines � Three hydrogen bonds required to bond Guanine & Cytosine � 3 H-bonds G C
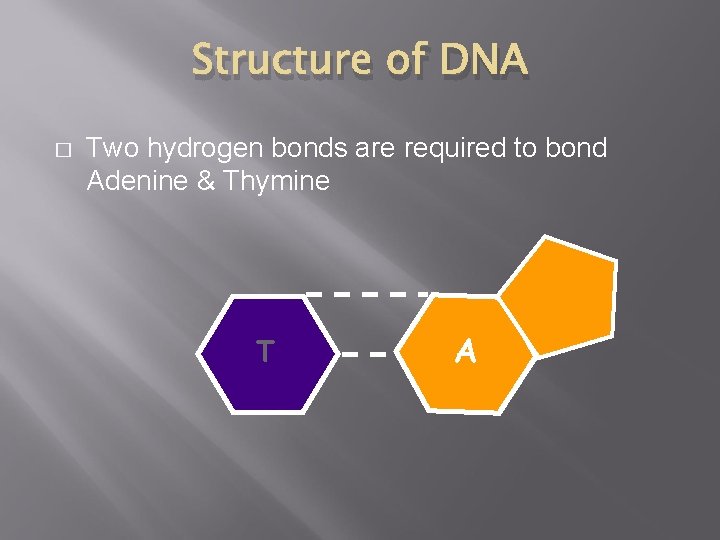
Structure of DNA � Two hydrogen bonds are required to bond Adenine & Thymine T A

Question: � If there is 30% Adenine, how much Cytosine is present?

Answer: � There would be 20% Cytosine Adenine (30%) = Thymine (30%) � Guanine (20%) = Cytosine (20%) � Therefore, 60% A-T and 40% C-G �

DNA REPLICATION
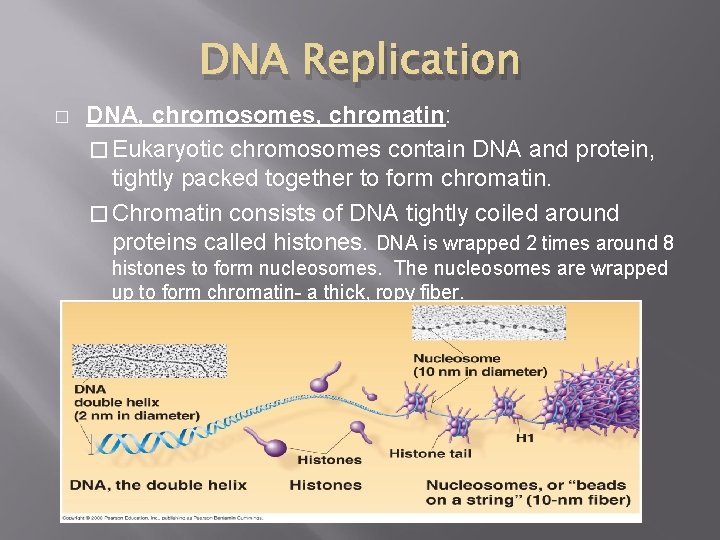
DNA Replication � DNA, chromosomes, chromatin: � Eukaryotic chromosomes contain DNA and protein, tightly packed together to form chromatin. � Chromatin consists of DNA tightly coiled around proteins called histones. DNA is wrapped 2 times around 8 histones to form nucleosomes. The nucleosomes are wrapped up to form chromatin- a thick, ropy fiber.

DNA Replication � � � DNA has to be copied before a cell divides – in the form of chromatin New cells will need a complete copy of identical DNA Chromatin does not condense into chromosomes until the first phase of cell division– MOST OF THE TIME it is in chromatin form.
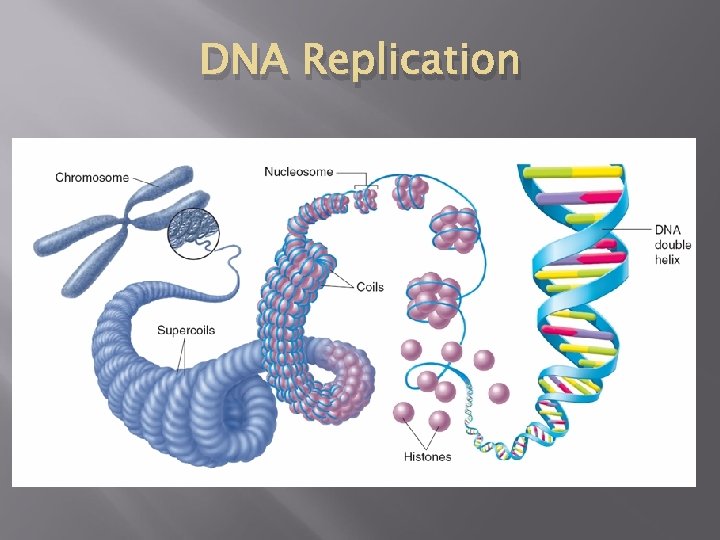
DNA Replication

DNA Replication � DNA Replication Each strand of the DNA double helix has all the information needed to reconstruct the other half by the mechanism of base pairing. � � Semi – Conservative Hypothesis– a parent strand is used as a model while new nucleotides attach: Meselsson and Stahl’s experiment showed Watson & Crick’s semi-conservative hypothesis to be true, by labeling some bacterial DNA with “heavy” nitrogen. DNA Template Parental DNA New DNA
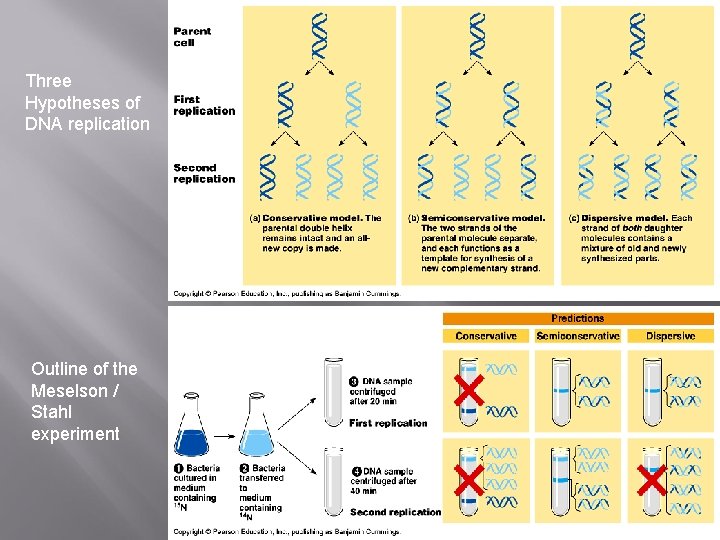
Three Hypotheses of DNA replication Outline of the Meselson / Stahl experiment

DNA Replication � � � In prokaryotic cells, DNA is located in the cytoplasm. Most prokaryotes have a single DNA molecule containing nearly all of the cell’s genetic information. In prokaryotes – replication begins at a single point and proceeds in two directions simultaneously until the DNA is copied.
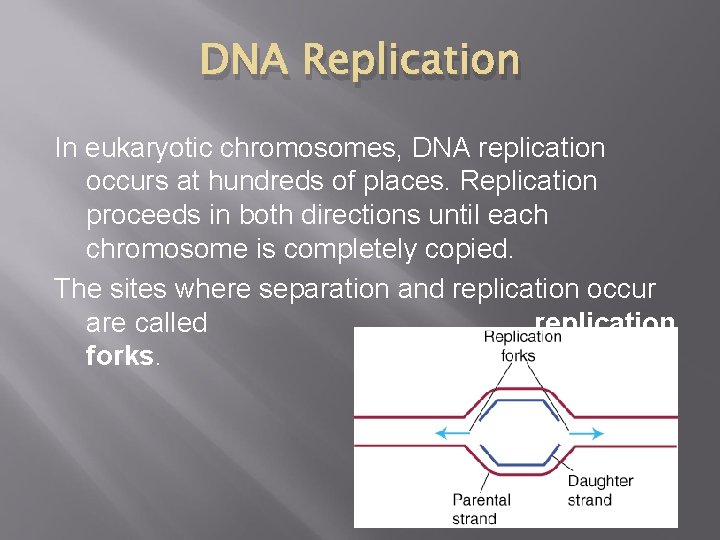
DNA Replication In eukaryotic chromosomes, DNA replication occurs at hundreds of places. Replication proceeds in both directions until each chromosome is completely copied. The sites where separation and replication occur are called replication forks.

DNA Replication � � DNA Replication Begins at Origins of Replication Two strands open forming Replication Forks (Y-shaped region) New strands grow at the forks
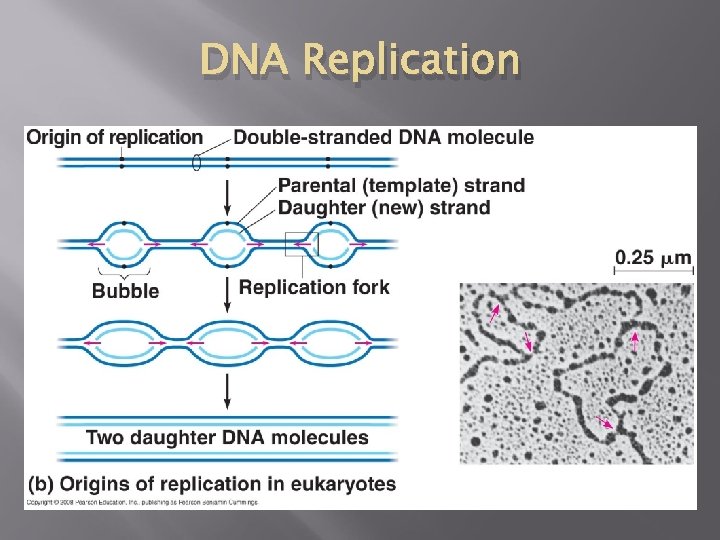
DNA Replication

DNA Replication � � � As the 2 DNA strands open at the origin, Replication Bubbles form Eukaryotic chromosomes have MANY bubbles Prokaryotes (bacteria) have a single bubble
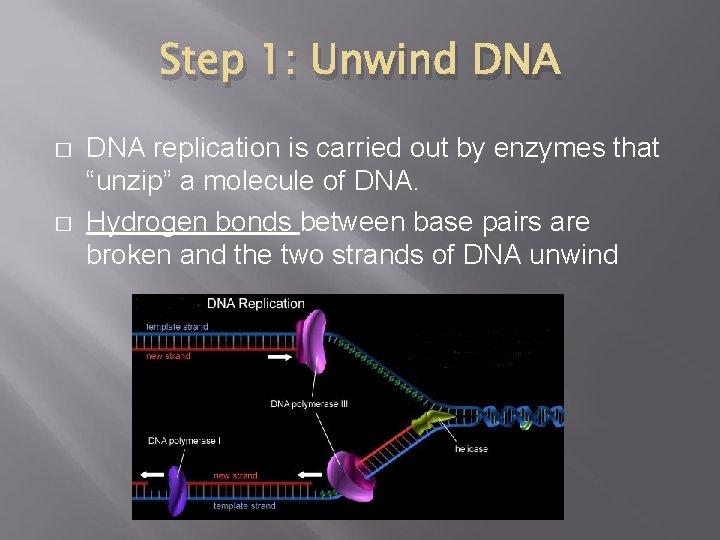
Step 1: Unwind DNA � � DNA replication is carried out by enzymes that “unzip” a molecule of DNA. Hydrogen bonds between base pairs are broken and the two strands of DNA unwind

Step 1: Unwind DNA 1. Enzyme Helicase unwinds and separates the 2 DNA strands by breaking the weak hydrogen bonds 2. Single-Strand Binding Proteins attach and keep the 2 DNA strands separated and untwisted (like thumb tacks)

Step 2: ADD RNA Primers 3. Before new DNA strands can form, there must be RNA primers (short RNA sequences) present to start the addition of new nucleotides � Primase is the enzyme that synthesizes the RNA Primer. The primer is about 10 nucleotides long. 4. DNA polymerase can then add the new nucleotides � The primer will be removed and replaced with DNA nucleotides at the end of replication.
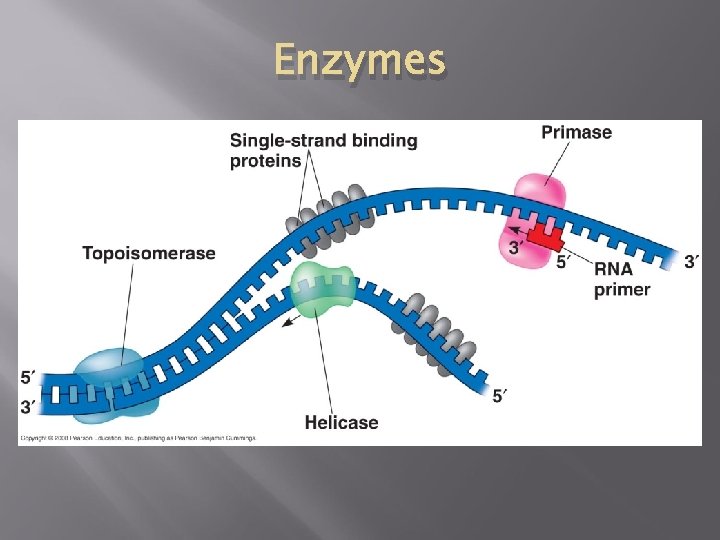
Enzymes

Step 3: Adding DNA Nucleotides � � � DNA polymerase can only add nucleotides to the 3’ end of the DNA This causes the NEW strand to be built in a 5’ to 3’ direction Remember each DNA molecule has two strands of nucleotides ANTI-PARALLEL
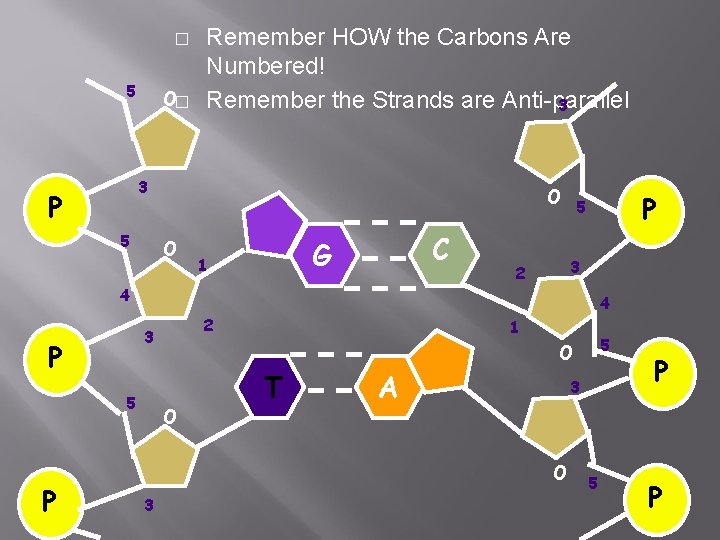
� 5 O� Remember HOW the Carbons Are Numbered! Remember the Strands are Anti-parallel 3 3 P 5 O O C G 1 4 P 5 P 3 2 4 2 3 1 5 O O T A 3 O 3 P 5 5 P P
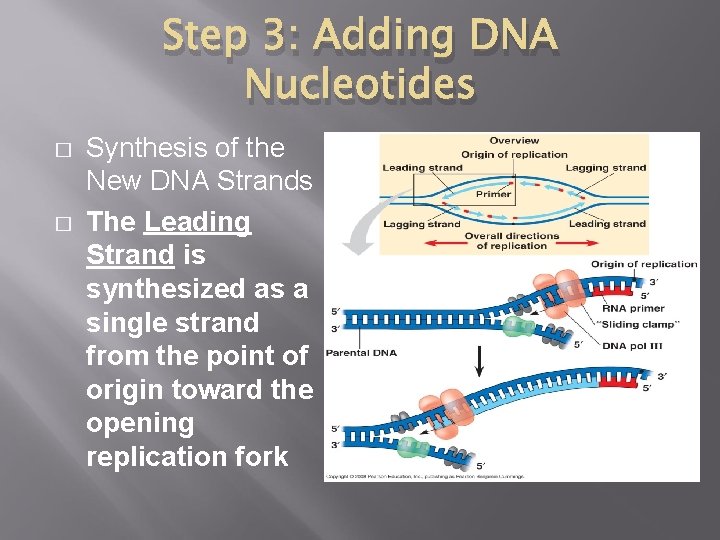
Step 3: Adding DNA Nucleotides � � Synthesis of the New DNA Strands The Leading Strand is synthesized as a single strand from the point of origin toward the opening replication fork
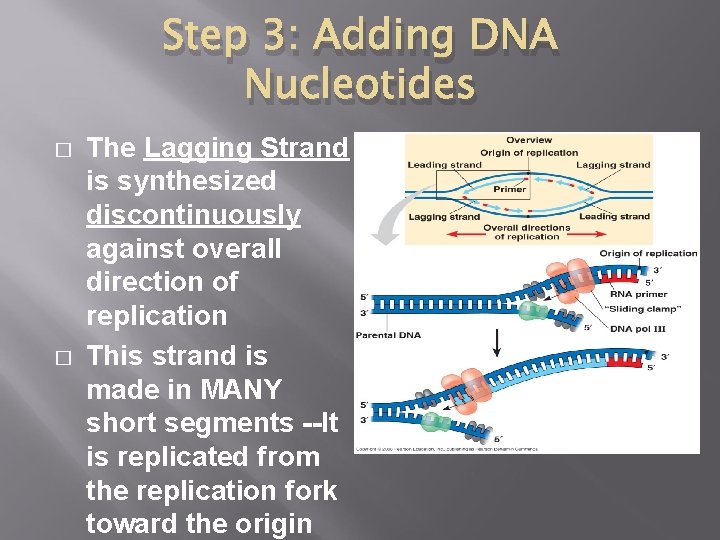
Step 3: Adding DNA Nucleotides � � The Lagging Strand is synthesized discontinuously against overall direction of replication This strand is made in MANY short segments --It is replicated from the replication fork toward the origin
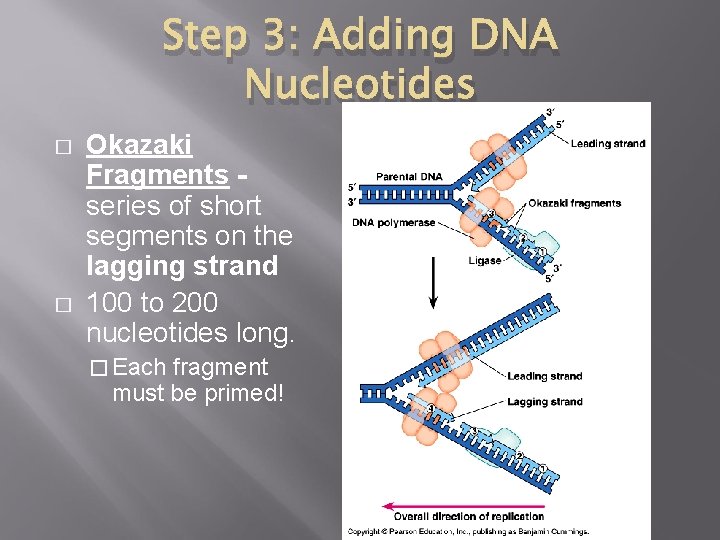
Step 3: Adding DNA Nucleotides � � Okazaki Fragments - series of short segments on the lagging strand 100 to 200 nucleotides long. � Each fragment must be primed!

Step 4: Removal of Primers � � A second DNA polymerase removes the RNA primers These are replaced with the proper DNA nucleotides.

Step 5: Sealing Lagging Fragments � � All segments of the sugar – phosphate backbone are sealed by Ligase This includes: � All fragments of lagging strand � Sections of primer replaced by DNA nucleotides.
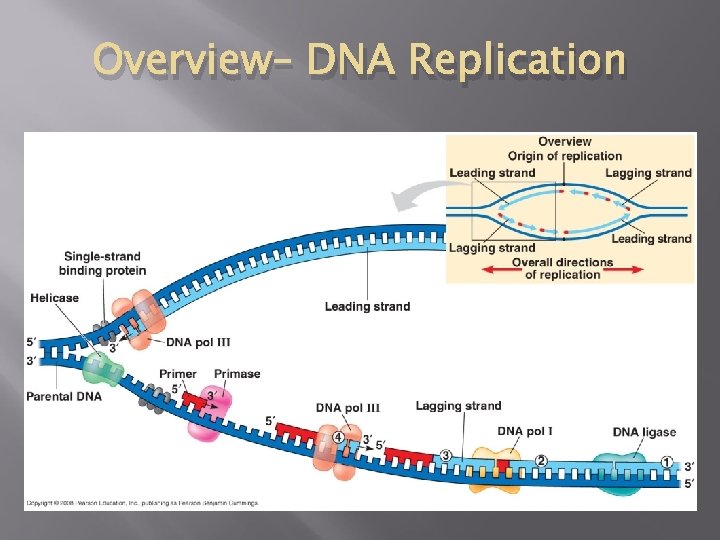
Overview– DNA Replication

Step 6: Proofreading � � Initial error rate: 1 in 10, 000 base pairs. DNA polymerase proofreads – removing and replacing nucleotides if necessary. Some times errors are missed or arise after. Cells must continuously repair DAMAGED DNA. Mismatch repair: special enzymes are used to fix incorrecly paired nucleotides. Nucleotide Excision Repair: nuclease/ exonuclease removes damaged section, DNA polymerase adds bases, Ligase reseals.
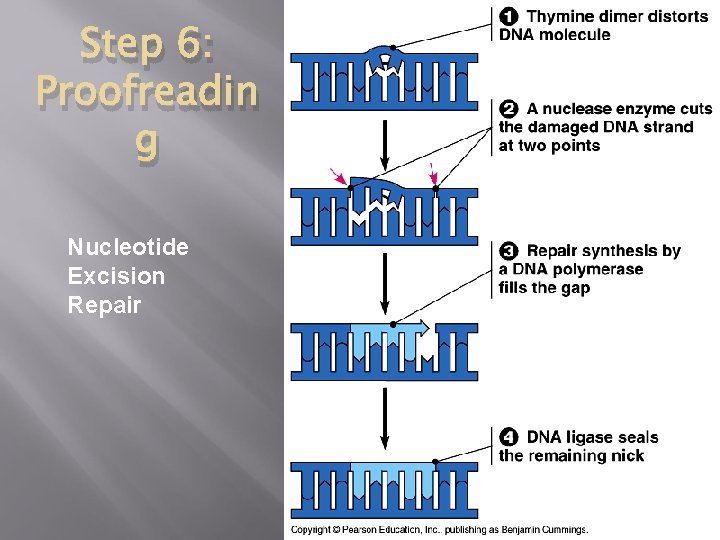
Step 6: Proofreadin g Nucleotide Excision Repair

Rate of Replication / Factors that can affect DNA � � � 500 nucleotide bases per second in bacteria 50 per second in human cells DNA is easily damaged by our environment. � Defect in proofreading mechanisms could lead to cancer. � Radiation, x-rays, UV light can change nucleotides that may affect genetic information. � 100 repair enzymes in E. coli; 130 repair enzymes in humans

Question: � What would be the complementary DNA strand for the following DNA sequence? DNA 5’-CGTATG-3’

Answer: DNA 5’GCGTA 3’ DNA 3’CGCAT 5’
- Slides: 65