DNA Barcoding and Molecular Phylogeny of Glyptothorax Species

DNA Barcoding and Molecular Phylogeny of Glyptothorax Species (Siluriformes, sisoridae) Mahender Singh 1*, W S Lakra 2, Y Ph Kartavtsev 3, A K Singh 1, S N Bahuguna 4 1 National Bureau of Fish Genetic Resources, Lucknow, India; Central Institute of Fisheries Education, Mumbai, India 3 A. V. Zhirmunsky Institute of Marine Biology, Vladivostok, Russia 4 Hemwati Nandan Bahuguna Garhwal University, Srinagar, Uttrakhand, India; ISO: 9001: 2008 http: //www. nbfgr. res. in/ 2

DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION Glyptothorax G. garhwali http: //www. nbfgr. res. in/ Siluriformes, sisoridae § Attached to rocks, boulders, stones at the bottom of the water bodies where they live by means of a thoracic adhesive apparatus : a sort of sucking disk as an adaptive structure. G. dakpathari G. brevipinnis G. ngapang § several new species have been reported from India in the past few years (Vishwanath & Linthiongambi 2006, 2007). § The debate over the criteria to recognize the species and subspecies boundaries has received considerable attention in recent years — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org
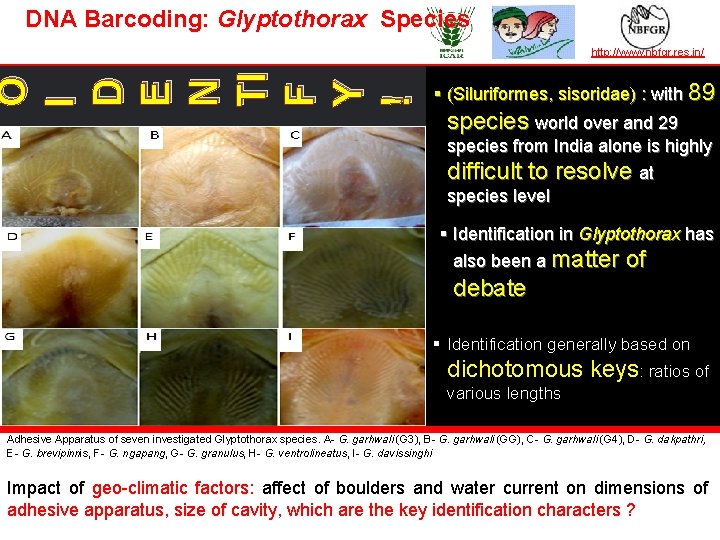
DNA Barcoding: Glyptothorax Species O I D E N TI F Y ! TAXONOMY IN ACTION http: //www. nbfgr. res. in/ § (Siluriformes, sisoridae) : with 89 species world over and 29 species from India alone is highly difficult to resolve at species level § Identification in Glyptothorax has also been a matter of debate § Identification generally based on dichotomous keys: ratios of © Author of image | affiliation various lengths Adhesive Apparatus of seven investigated Glyptothorax species. A- G. garhwali (G 3), B- G. garhwali (GG), C- G. garhwali (G 4), D- G. dakpathri, E- G. brevipinnis, F- G. ngapang, G- G. granulus, H- G. ventrolineatus, I- G. davissinghi Impact of geo-climatic factors: affect of boulders and water current on dimensions of adhesive apparatus, size of cavity, which are the key identification characters ? — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org

DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION http: //www. nbfgr. res. in/ recognition based on small DNA barcoding: for taxon § Taxon sequence fragment (655 base pairs) recognition and classification near the 5’end of mitochondrial gene cytochrome c oxidase I (COI) with universal primers is proved to be effective (Hebert et al. , 2003). § This barcode is also suitable marker for discriminating between closely related species of marine and freshwater fishes © Arthur D. Chapman | Australian Biodiversity Information Services DNA barcoding is also being used for detection of species substitution in processed food products, products illegal products from regulated species control of invasive species, pest management, water quality control etc. — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org

DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION Present Study: in Glyptothorax http: //www. nbfgr. res. in/ Resolving ambiguities (i) Use of well recognized DNA Species barcoding methodology vis-à-vis morphological character-based criteria to fix the molecular signature for species ii) To determine if COI based barcoding is suitable for species identification as well as molecular phylogeny in Glyptothorax catfishes with overlapping range of morphometric characters and Voucher specimens of Glyptothorax Antonio G. Valdecasas |MNCN, CSIC | Spain iii)© To prove whether Glyptothorax genus of catfish is monophyletic based on currently analyzed COI dataset as well as concatamer with earlier dataset of Cyt b. — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org

DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION http: //www. nbfgr. res. in/ Glyptothorax Sampling Sites [ ] in India § Khanda Gad Near Srinagar in Garhwal, Uttarakhand § Maldevta, Dehradun, Uttarakhand § Yamuna Barrage, Dakpathar, Dehradun, Uttarakhand § Tatapani, Shimla, Himachal Pradesh § Downstream of Poringalkuthu Dam, Trishur, Kerala § Serou, Manipur — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org
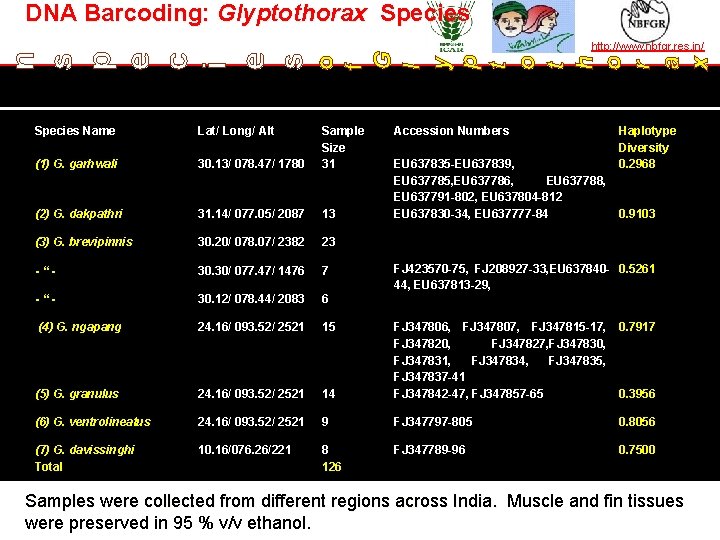
DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ o f G l y p t o t h o r a x n s p e c i e s TAXONOMY IN ACTION Species Name Lat/ Long/ Alt (1) G. garhwali 30. 13/ 078. 47/ 1780 Sample Size 31 (2) G. dakpathri 31. 14/ 077. 05/ 2087 13 (3) G. brevipinnis 30. 20/ 078. 07/ 2382 23 -“- 30. 30/ 077. 47/ 1476 7 -“- 30. 12/ 078. 44/ 2083 6 (4) G. ngapang 24. 16/ 093. 52/ 2521 15 (5) G. granulus 24. 16/ 093. 52/ 2521 (6) G. ventrolineatus (7) G. davissinghi Total Accession Numbers EU 637835 -EU 637839, EU 637785, EU 637786, EU 637788, EU 637791 -802, EU 637804 -812 EU 637830 -34, EU 637777 -84 Haplotype Diversity 0. 2968 0. 9103 FJ 423570 -75, FJ 208927 -33, EU 637840 - 0. 5261 44, EU 637813 -29, 0. 7917 14 FJ 347806, FJ 347807, FJ 347815 -17, FJ 347820, FJ 347827, FJ 347830, FJ 347831, FJ 347834, FJ 347835, FJ 347837 -41 FJ 347842 -47, FJ 347857 -65 24. 16/ 093. 52/ 2521 9 FJ 347797 -805 0. 8056 10. 16/076. 26/221 8 126 FJ 347789 -96 0. 7500 0. 3956 Samples were collected from different regions across India. Muscle and fin tissues were preserved in 95 % v/v ethanol. — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org

DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ Me t h o d o l o g i e s f o l l o w e d : TAXONOMY IN ACTION § Approximately 50 mg of caudal or anal fin or muscular tissue was used for DNA isolation following standard phenol/chloroform method (Sambrook et al. , 1989) with partial modifications. § The partial COI gene was amplified using the universal primers Fish F 1 5’TCAACCACAAAGACATTGGCAC 3 Fish. R 1 -5’TAGACTTCTGGGTGGCCAAA GAATCA 3 (Ward et al. 2005). & § DNA sequencing was performed following the dideoxynucleotide chain termination method (Sanger et al. , 1977), 1977 using an automated ABI 3730 sequencer § The COI sequences were aligned and primer sequences were trimmed, to get uniform length sequences of 655 base. § Sequences were aligned using CLUSTALW (Thompson et al. 1994), refereed against electropherogram and submitted to Gen. Bank. The phylogenetic trees were visualized and edited where necessary with Fig. Tree (http: //tree. bio. ed. ac. uk/software/figtree/) and Dendroscope (http: //ab. inf. uni— the leader in establishing and operating partnerships for taxonomy developing 2010) countries software | www. bionet-intl. org | bionet@bionet-intl. org tuebingen. de/software/dendroscope/) (Huson et al. , in 2007;
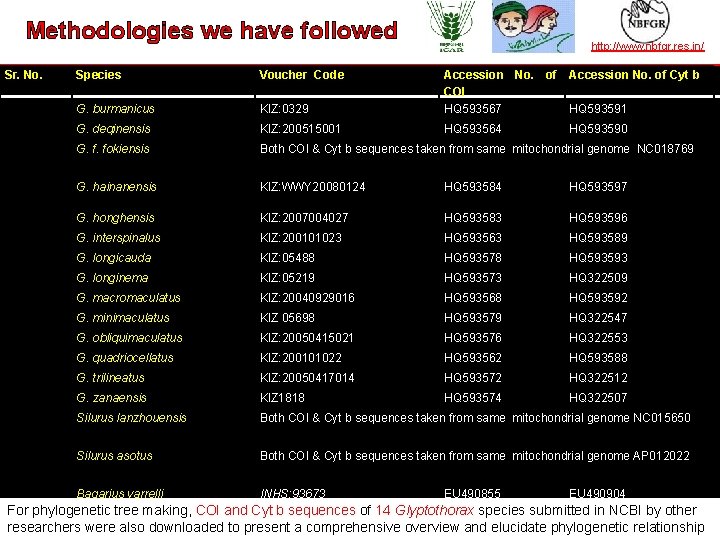
Methodologies followed TAXONOMY we INhave ACTION Sr. No. Species Voucher Code G. burmanicus http: //www. nbfgr. res. in/ Accession No. of Cyt b KIZ: 0329 Accession No. of COI HQ 593567 G. deqinensis KIZ: 200515001 HQ 593564 HQ 593590 G. f. fokiensis Both COI & Cyt b sequences taken from same mitochondrial genome NC 018769 G. hainanensis KIZ: WWY 20080124 HQ 593584 HQ 593597 G. honghensis KIZ: 2007004027 HQ 593583 HQ 593596 G. interspinalus KIZ: 200101023 HQ 593563 HQ 593589 G. longicauda KIZ: 05488 HQ 593578 HQ 593593 G. longinema KIZ: 05219 HQ 593573 HQ 322509 G. macromaculatus KIZ: 20040929016 HQ 593568 HQ 593592 G. minimaculatus KIZ 05698 HQ 593579 HQ 322547 G. obliquimaculatus KIZ: 20050415021 HQ 593576 HQ 322553 G. quadriocellatus KIZ: 200101022 HQ 593562 HQ 593588 G. trilineatus KIZ: 20050417014 HQ 593572 HQ 322512 G. zanaensis KIZ 1818 HQ 593574 HQ 322507 Silurus lanzhouensis Both COI & Cyt b sequences taken from same mitochondrial genome NC 015650 Silurus asotus Both COI & Cyt b sequences taken from same mitochondrial genome AP 012022 Bagarius yarrelli INHS: 93673 EU 490855 HQ 593591 EU 490904 For phylogenetic tree making, COI and Cyt b sequences of 14 Glyptothorax species submitted in NCBI by other researchers —were alsoin downloaded to present a comprehensive overview elucidate phylogenetic relationship the leader establishing and operating partnerships for taxonomy in developing countriesand | www. bionet-intl. org | bionet@bionet-intl. org

DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION Result highlights: http: //www. nbfgr. res. in/ Analysis of COI : Out of 655 positions, § § G. garhwali 514 were conserved (82. 6%) 114 (17. 4%) were variable 10 were singleton (1. 5%) 104 (15. 9%) were parsimoniously informative (at least two of nucleotides occurring with a minimum frequency of two). As the adhesive apparatus is key character, it was photographed in mature specimens of all seven species. By adhesive apparatus and morphological data, two populations of G. garhwali considered as different species — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org
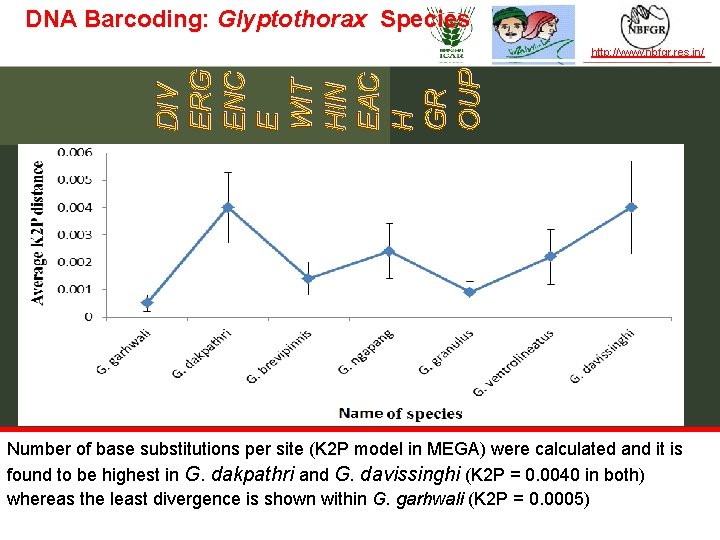
DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ DIV ERG ENC E WIT HIN EAC H GR OUP TAXONOMY IN ACTION Number of base substitutions per site (K 2 P model in MEGA) were calculated and it is found to be highest in G. dakpathri and G. davissinghi (K 2 P = 0. 0040 in both) whereas the least divergence is shown within G. garhwali (K 2 P = 0. 0005) — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org
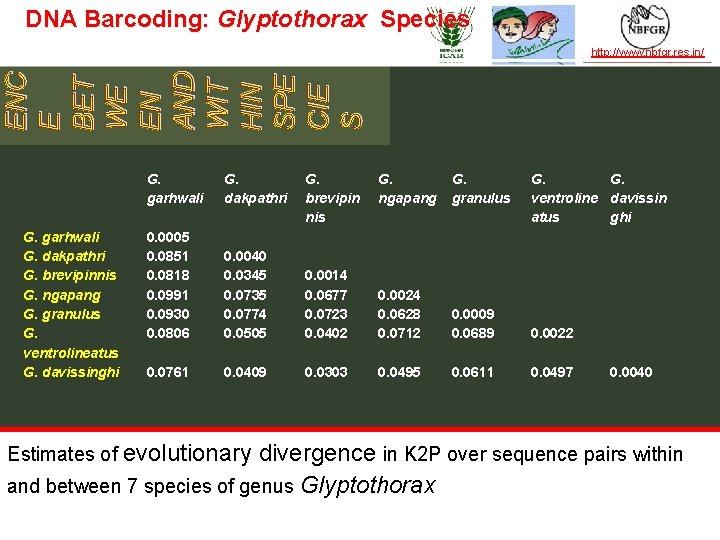
DNA Barcoding: Glyptothorax Species TAXONOMY IN ACTION ENC E BET WE EN AND WIT HIN SPE CIE S http: //www. nbfgr. res. in/ G. garhwali G. dakpathri G. brevipinnis G. ngapang G. granulus G. ventrolineatus G. davissinghi G. garhwali G. dakpathri 0. 0005 0. 0851 0. 0818 0. 0991 0. 0930 0. 0806 0. 0040 0. 0345 0. 0735 0. 0774 0. 0505 0. 0761 0. 0409 G. brevipin nis G. ngapang G. granulus G. G. ventroline davissin atus ghi 0. 0014 0. 0677 0. 0723 0. 0402 0. 0024 0. 0628 0. 0712 0. 0009 0. 0689 0. 0022 0. 0303 0. 0495 0. 0611 0. 0497 0. 0040 Estimates of evolutionary divergence in K 2 P over sequence pairs within and between 7 species of genus Glyptothorax — the leader in establishing and operating partnerships for taxonomy in developing countries | www. bionet-intl. org | bionet@bionet-intl. org
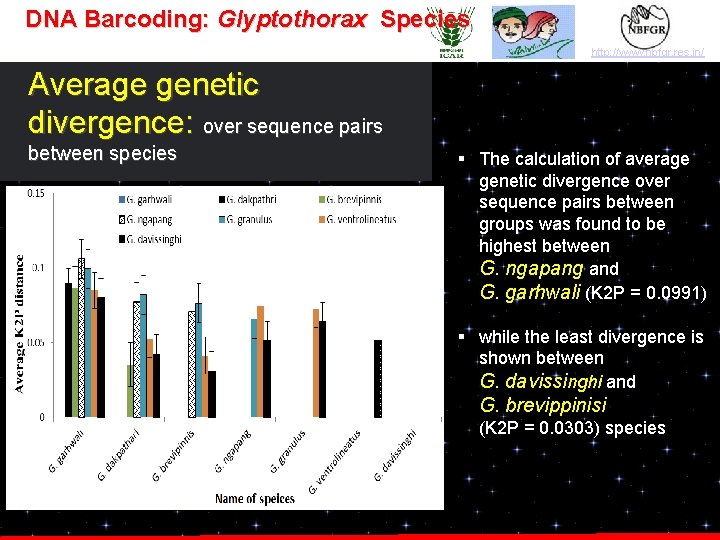
DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ Average genetic divergence: over sequence pairs between species § The calculation of average genetic divergence over sequence pairs between groups was found to be highest between G. ngapang and G. garhwali (K 2 P = 0. 0991) § while the least divergence is shown between G. davissinghi and G. brevippinisi (K 2 P = 0. 0303) species
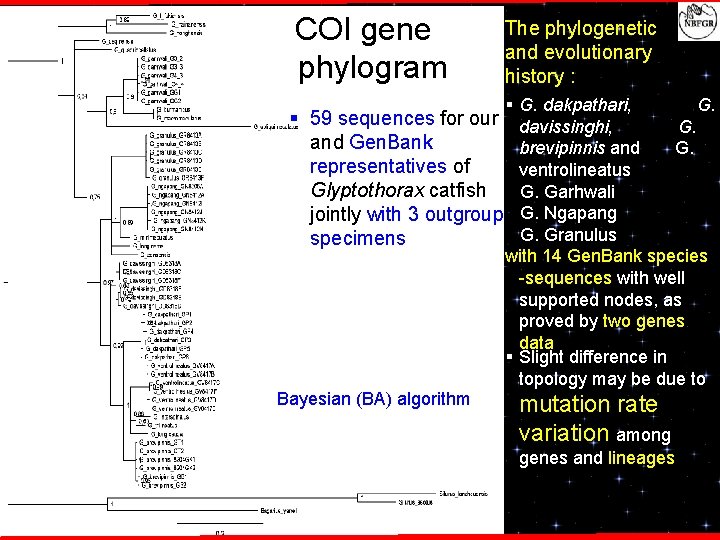
COI gene phylogram The phylogenetic and evolutionary history : § G. dakpathari, G. § 59 sequences for our davissinghi, G. and Gen. Bank brevipinnis and G. representatives of ventrolineatus Glyptothorax catfish G. Garhwali jointly with 3 outgroup G. Ngapang G. Granulus specimens with 14 Gen. Bank species -sequences with well supported nodes, as proved by two genes data § Slight difference in topology may be due to Bayesian (BA) algorithm mutation rate variation among genes and lineages

Combined phylogram of COI and Cyt-b genes. Results revealed § 59 sequences for our and Gen. Bank representatives of Glyptothorax catfishes jointly with 3 outgroup specimens that COI, as well as § In the nodes the support levels (%) are: BA, ML, MP, and NJ. Glyptothorax from India § Abbreviation 100 x 4 denotes: 100, 100 and 100 x 3 denotes: 100, 100 Bayesian (BA) algorithm Cyt-b (Singh et al. , 2012 and this study), are the effective markers for identifications of seven species of
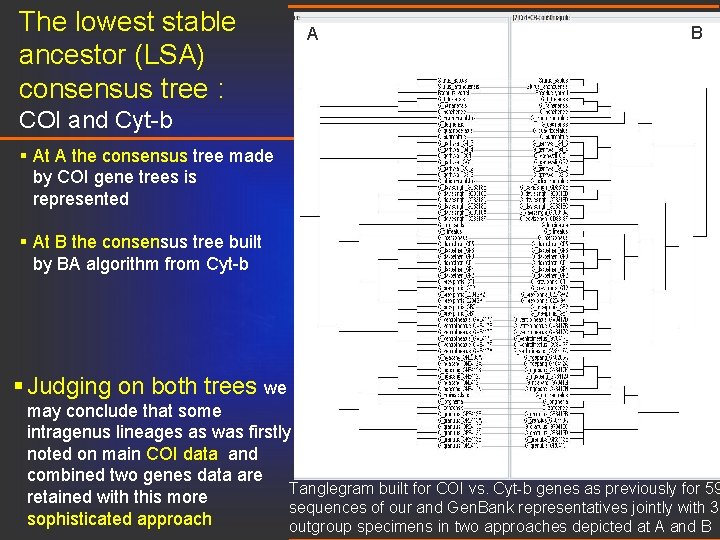
The lowest stable ancestor (LSA) consensus tree : A B COI and Cyt-b § At A the consensus tree made by COI gene trees is represented § At B the consensus tree built by BA algorithm from Cyt-b § Judging on both trees we may conclude that some intragenus lineages as was firstly noted on main COI data and combined two genes data are Tanglegram built for COI vs. Cyt-b genes as previously for 59 retained with this more sequences of our and Gen. Bank representatives jointly with 3 sophisticated approach outgroup specimens in two approaches depicted at A and B

DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ Findings: § These seven species of Sisoridae from the Indian rivers were found genetically distinct from each other. § The adhesive apparatus, considered as the key character for species identification, is not supported by barcode data as G. garhwali has two types of adhesive apparatus § Pairwise comparisons, sequence alignment, intra-specific and inter-specific sequence divergence the pattern of nucleotide substitution found in COI gene of Glyptothorax species, is quite expected for a functionally conserved protein coding region, which has evolved over recent evolutionary time. § Glyptothorax exhibited more nucleotide change at third codon position, with most mutations being synonymous. We also observed anti G bias in this position in the seven species, and the G is mostly substituted with A (both purines with double ring nitrogenous base).

DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ Findings: § Estimate of genetic divergence with COI gene was sufficient enough to discriminate individuals of different Glyptothorx species within Sisoridae. § In all seven species, levels of intra-specific variation (K 2 P sequence divergence) were varied from G. garhwali (0. 05%), G. dakpathri (0. 40%), G. ngapang (0. 24%), G. davissinghi (0. 40% ), G. brevipinnis (0. 14%), G. granulus (0. 09%), G. ventrolineatus (0. 22%). § High inter-specific K 2 P sequence divergence is estimated between G. brevipinnis & G. garhwali (0. 129) despite small geographical distance between them and low inter-specific K 2 P sequence divergence is estimated between G. davissinghi and G. dakpathri (0. 002) despite a distance of over 2000 km.

DNA Barcoding: Glyptothorax Species http: //www. nbfgr. res. in/ Authors are thankful to Prof. W. Vishwanath for helping in identification of Glyptothorax species from North East India, National Bureau of Fish Genetic Resources of India for supporting this work.

THANK YOU!
- Slides: 20