Disease Gene Finding Table of contents Background Why

Disease Gene Finding. Table of contents: Background Why do we want to find disease genes, how has it been done until now? Networks – deducing functional relationships from network theory Networks Biological networks Functional modules / network clusters Phenotype association Grouping disorders based on their phenotype. Biological implications of phenotype clusters. Method and examples Combining network theory and phenotype associations in an automated large scale disease gene finding platform Proof of concept.

Abstract Aim Find new disease genes. Means Use protein interaction networks and phenotype association networks for inferring phenotype gneotype relationships. Proof Interesting candidates are reported to experimental collaborators who perform mutational analysis in patient material.

Background
Background Aim Finding genes responsible for major genetic disorders can lead to diagnostics, potential drug targets, treatments and large amounts of information about molecular cell biology in general.
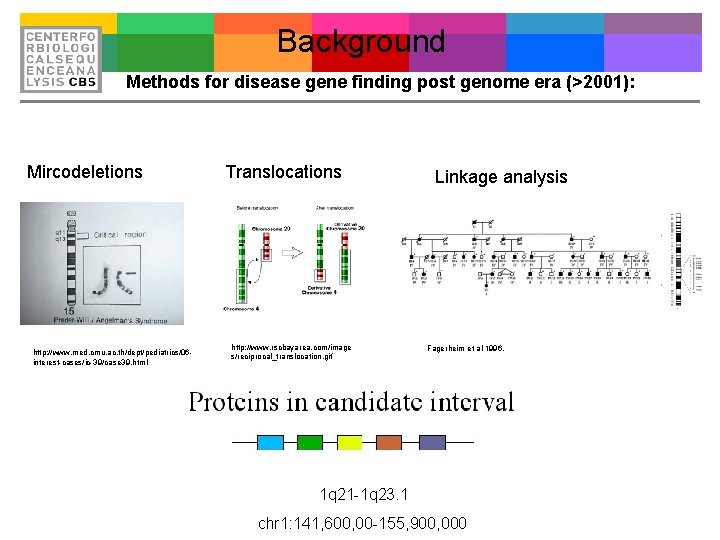
Background Methods for disease gene finding post genome era (>2001): Mircodeletions http: //www. med. cmu. ac. th/dept/pediatrics/06 interest-cases/ic-39/case 39. html Translocations http: //www. rscbayarea. com/image s/reciprocal_translocation. gif Linkage analysis Fagerheim et al 1996. 1 q 21 -1 q 23. 1 chr 1: 141, 600, 00 -155, 900, 000
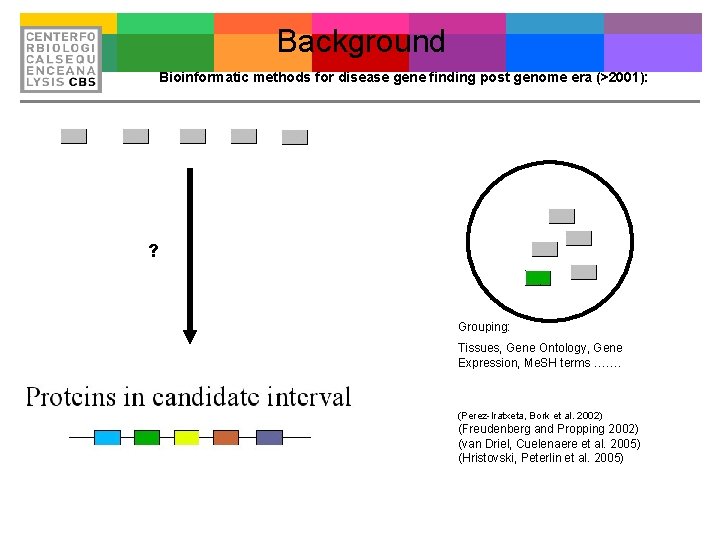
Background Bioinformatic methods for disease gene finding post genome era (>2001): ? Grouping: Tissues, Gene Ontology, Gene Expression, Me. SH terms ……. (Perez-Iratxeta, Bork et al. 2002) (Freudenberg and Propping 2002) (van Driel, Cuelenaere et al. 2005) (Hristovski, Peterlin et al. 2005)

Disease Gene Finding. Table of contents: Background Why do we want to find disease genes, how has it been done until now? Networks – deducing functional relationships from network theory Networks Biological networks Functional modules / network clusters Phenotype association Grouping disorders based on their phenotype. Biological implications of phenotype clusters. Method and examples Combining network theory and phenotype associations in an automated large scale disease gene finding platform Proof of concept.

Networks and functional modules Deducing functional relationships from network theory

Networks and functional modules Deducing functional relationships from network theory Network theory is boooooring

Networks Text mining of full text corpora e. g Pub. Med Central http: //www. biosolveit. de/To. PNet/screenshots/fig 1. html

Networks Protein interaction networks of physical interactions. (Barabasi and Oltvai 2004).
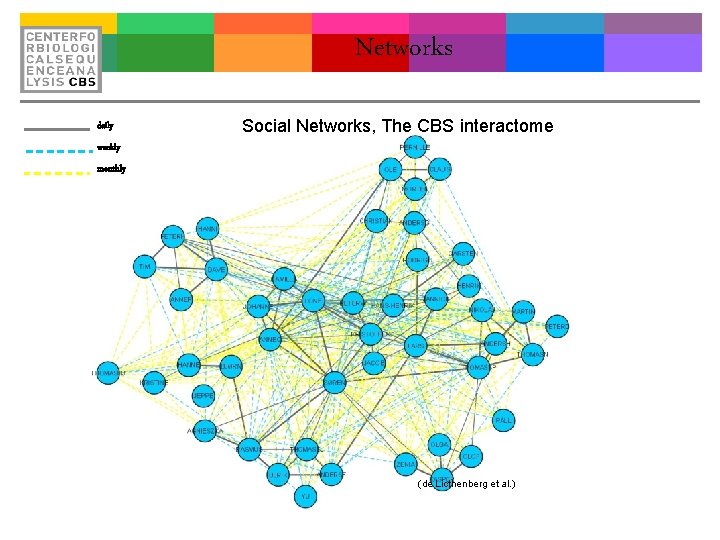
Networks daily Social Networks, The CBS interactome weekly monthly (de Licthenberg et al. )
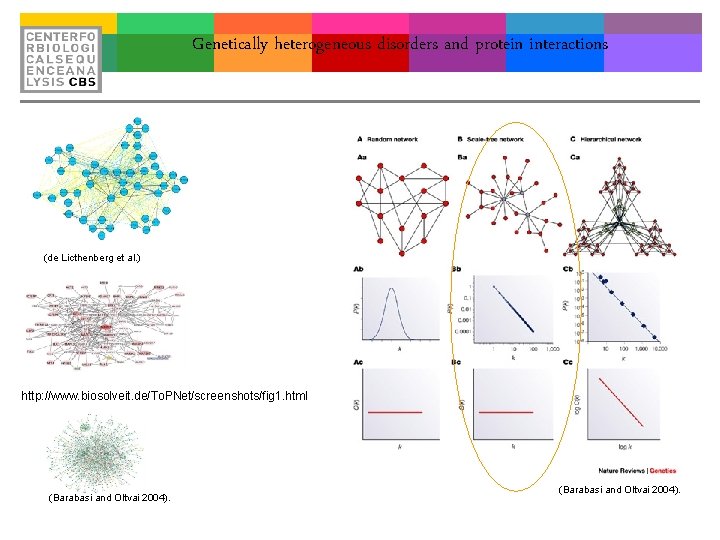
Genetically heterogeneous disorders and protein interactions (de Licthenberg et al. ) http: //www. biosolveit. de/To. PNet/screenshots/fig 1. html (Barabasi and Oltvai 2004).
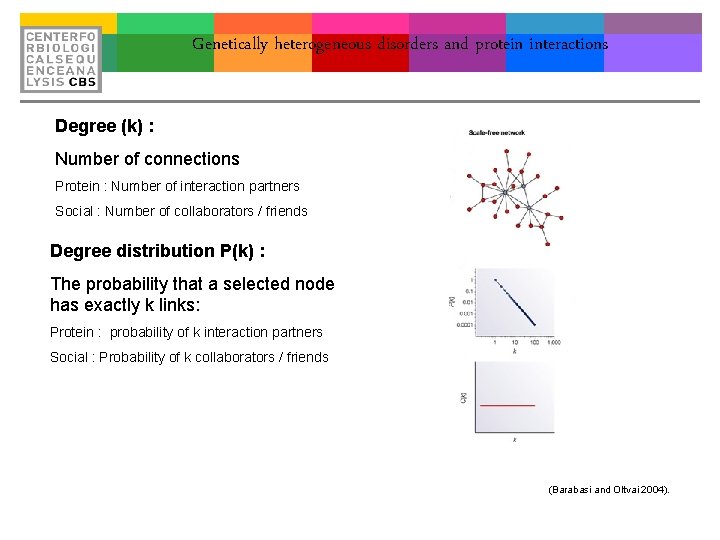
Genetically heterogeneous disorders and protein interactions Degree (k) : Number of connections Protein : Number of interaction partners Social : Number of collaborators / friends Degree distribution P(k) : The probability that a selected node has exactly k links: Protein : probability of k interaction partners Social : Probability of k collaborators / friends (Barabasi and Oltvai 2004).
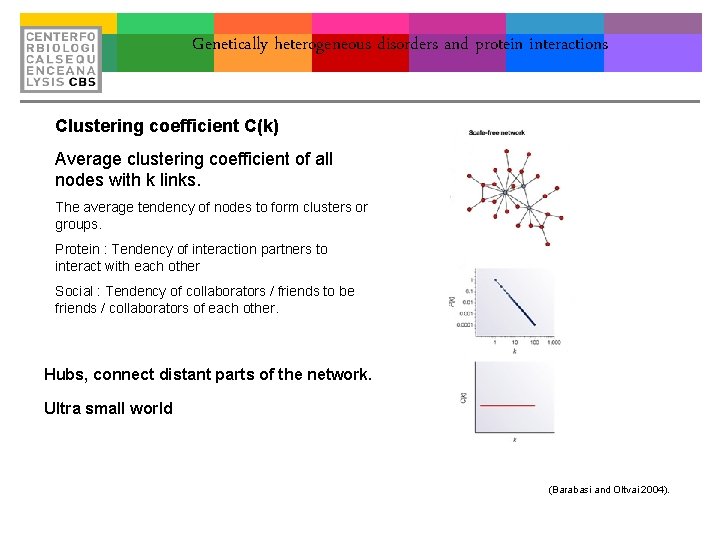
Genetically heterogeneous disorders and protein interactions Clustering coefficient C(k) Average clustering coefficient of all nodes with k links. The average tendency of nodes to form clusters or groups. Protein : Tendency of interaction partners to interact with each other Social : Tendency of collaborators / friends to be friends / collaborators of each other. Hubs, connect distant parts of the network. Ultra small world (Barabasi and Oltvai 2004).
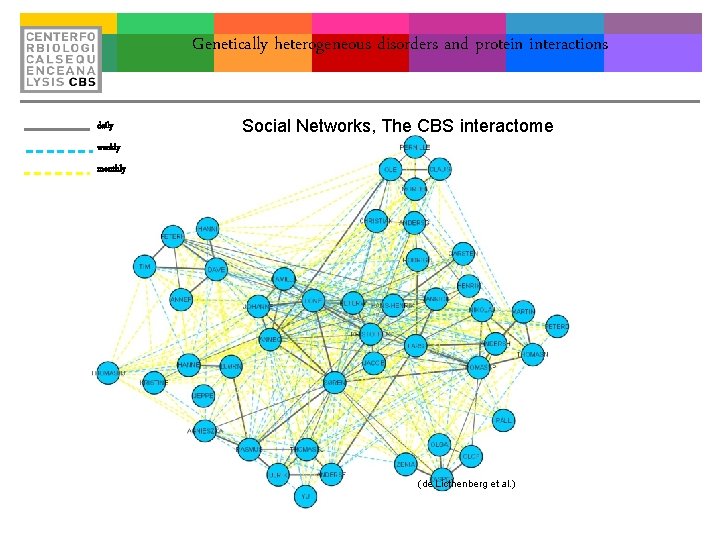
Genetically heterogeneous disorders and protein interactions daily Social Networks, The CBS interactome weekly monthly (de Licthenberg et al. )
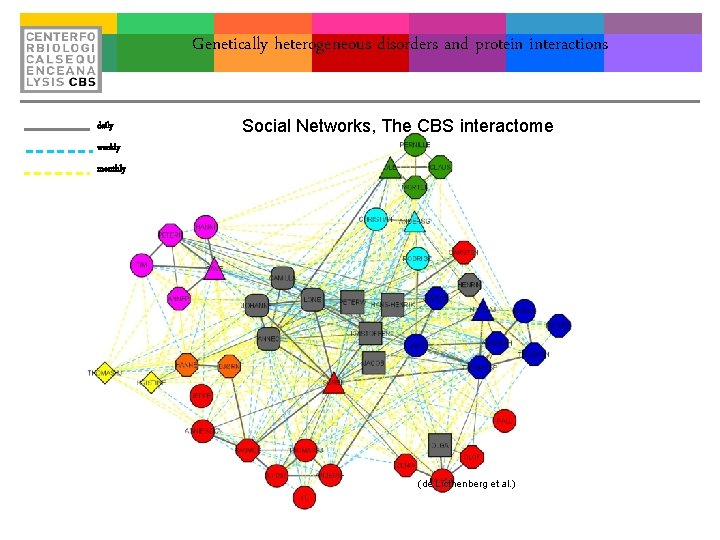
Genetically heterogeneous disorders and protein interactions daily Social Networks, The CBS interactome weekly monthly (de Licthenberg et al. )
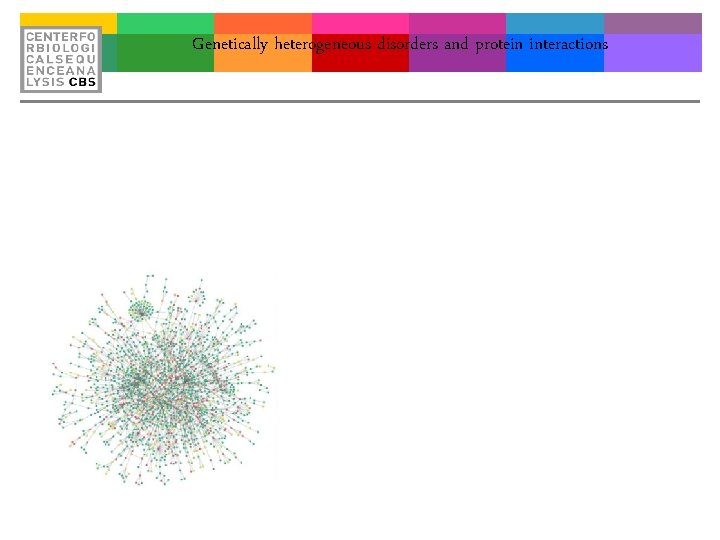
Genetically heterogeneous disorders and protein interactions
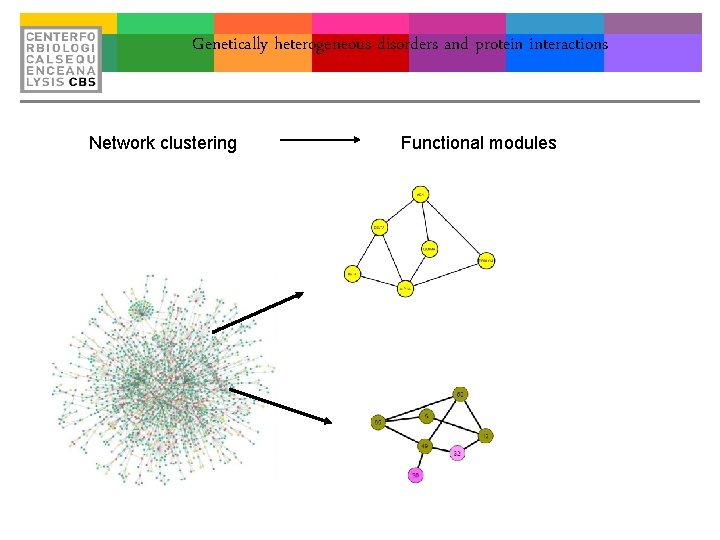
Genetically heterogeneous disorders and protein interactions Network clustering Functional modules
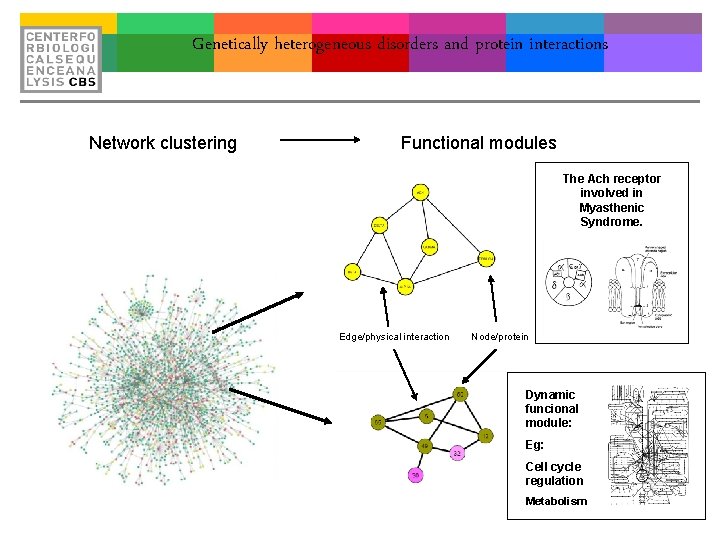
Genetically heterogeneous disorders and protein interactions Network clustering Functional modules The Ach receptor involved in Myasthenic Syndrome. Edge/physical interaction Node/protein Dynamic funcional module: Eg: Cell cycle regulation Metabolism
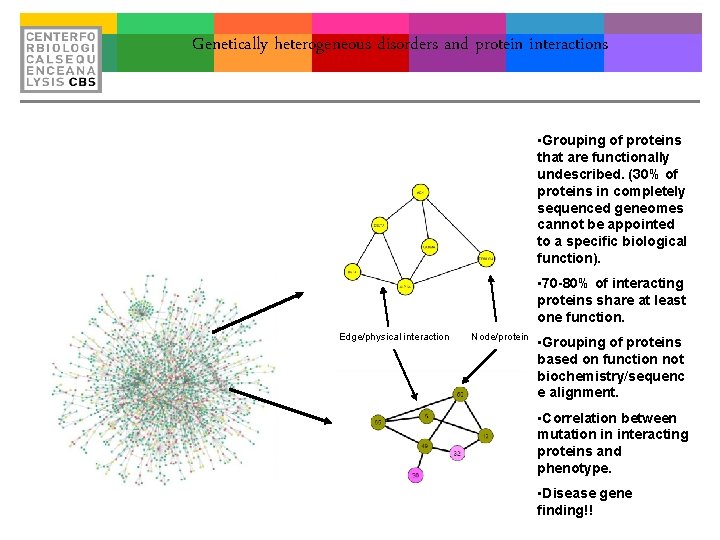
Genetically heterogeneous disorders and protein interactions • Grouping of proteins that are functionally undescribed. (30% of proteins in completely sequenced geneomes cannot be appointed to a specific biological function). • 70 -80% of interacting proteins share at least one function. Edge/physical interaction Node/protein • Grouping of proteins based on function not biochemistry/sequenc e alignment. • Correlation between mutation in interacting proteins and phenotype. • Disease gene finding!!
Disease Gene Finding. Table of contents: Background Why do we want to find disease genes, how has it been done until now? Networks – deducing functional relationships from network theory Networks Biological networks Functional modules / network clusters Phenotype association Grouping disorders based on their phenotype. Biological implications of phenotype clusters. Method and examples Combining network theory and phenotype associations in an automated large scale disease gene finding platform Proof of concept.

Phenotype association
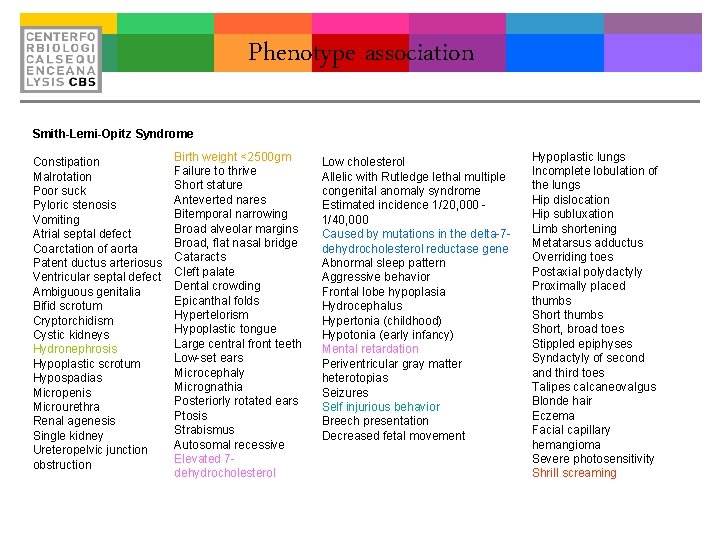
Phenotype association Smith-Lemi-Opitz Syndrome Constipation Malrotation Poor suck Pyloric stenosis Vomiting Atrial septal defect Coarctation of aorta Patent ductus arteriosus Ventricular septal defect Ambiguous genitalia Bifid scrotum Cryptorchidism Cystic kidneys Hydronephrosis Hypoplastic scrotum Hypospadias Micropenis Microurethra Renal agenesis Single kidney Ureteropelvic junction obstruction Birth weight <2500 gm Failure to thrive Short stature Anteverted nares Bitemporal narrowing Broad alveolar margins Broad, flat nasal bridge Cataracts Cleft palate Dental crowding Epicanthal folds Hypertelorism Hypoplastic tongue Large central front teeth Low-set ears Microcephaly Micrognathia Posteriorly rotated ears Ptosis Strabismus Autosomal recessive Elevated 7 dehydrocholesterol Low cholesterol Allelic with Rutledge lethal multiple congenital anomaly syndrome Estimated incidence 1/20, 000 - 1/40, 000 Caused by mutations in the delta-7 dehydrocholesterol reductase gene Abnormal sleep pattern Aggressive behavior Frontal lobe hypoplasia Hydrocephalus Hypertonia (childhood) Hypotonia (early infancy) Mental retardation Periventricular gray matter heterotopias Seizures Self injurious behavior Breech presentation Decreased fetal movement Hypoplastic lungs Incomplete lobulation of the lungs Hip dislocation Hip subluxation Limb shortening Metatarsus adductus Overriding toes Postaxial polydactyly Proximally placed thumbs Short, broad toes Stippled epiphyses Syndactyly of second and third toes Talipes calcaneovalgus Blonde hair Eczema Facial capillary hemangioma Severe photosensitivity Shrill screaming
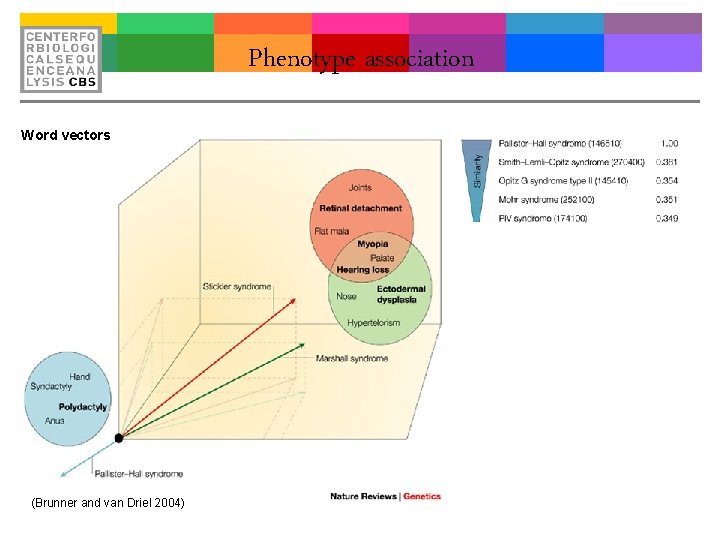
Phenotype association Word vectors (Brunner and van Driel 2004)
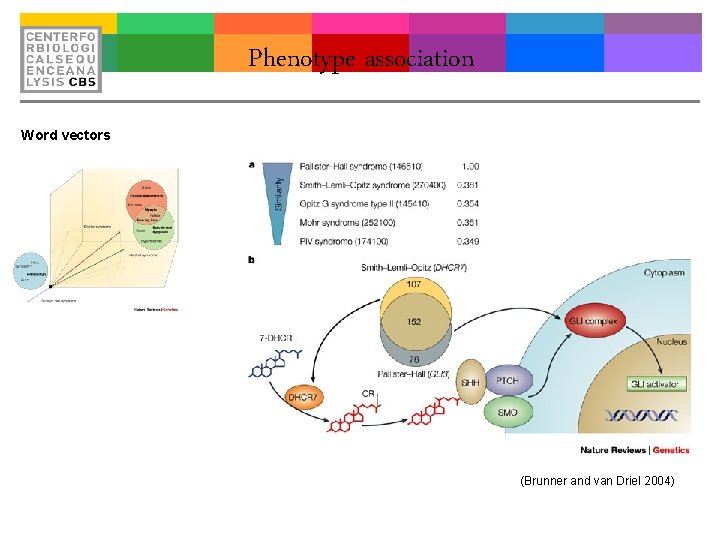
Phenotype association Word vectors (Brunner and van Driel 2004)
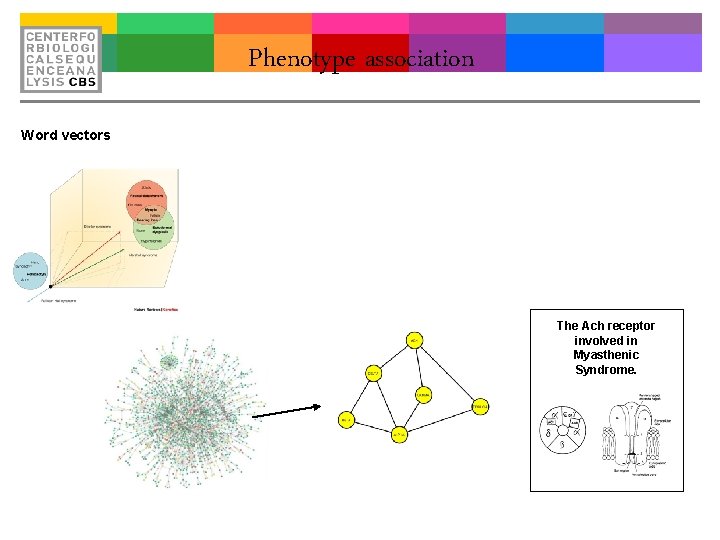
Phenotype association Word vectors The Ach receptor involved in Myasthenic Syndrome.
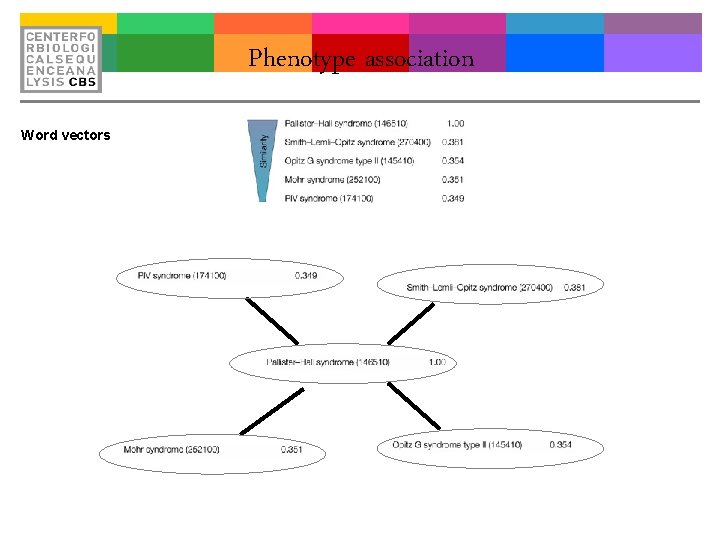
Phenotype association Word vectors

Disease Gene Finding. Table of contents: Background Why do we want to find disease genes, how has it been done until now? Networks – deducing functional relationships from network theory Networks Biological networks Functional modules / network clusters Phenotype association Grouping disorders based on their phenotype. Biological implications of phenotype clusters. Method and examples Combining network theory and phenotype associations in an automated large scale disease gene finding platform Proof of concept.

Method – Proof of concept
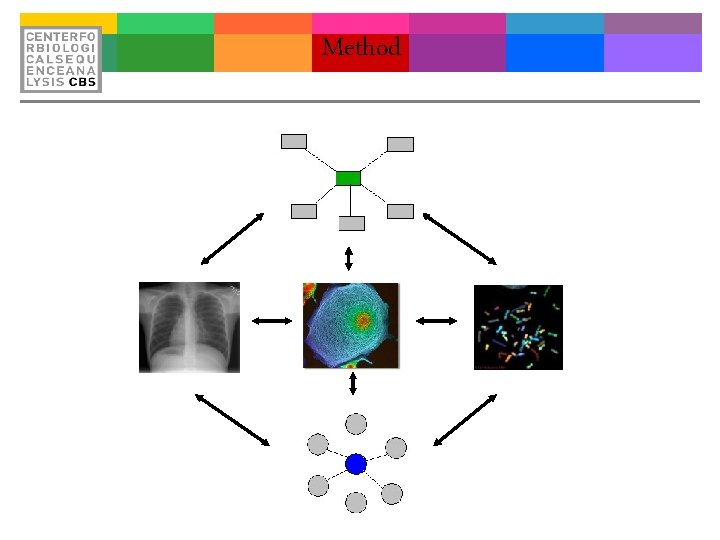
Method
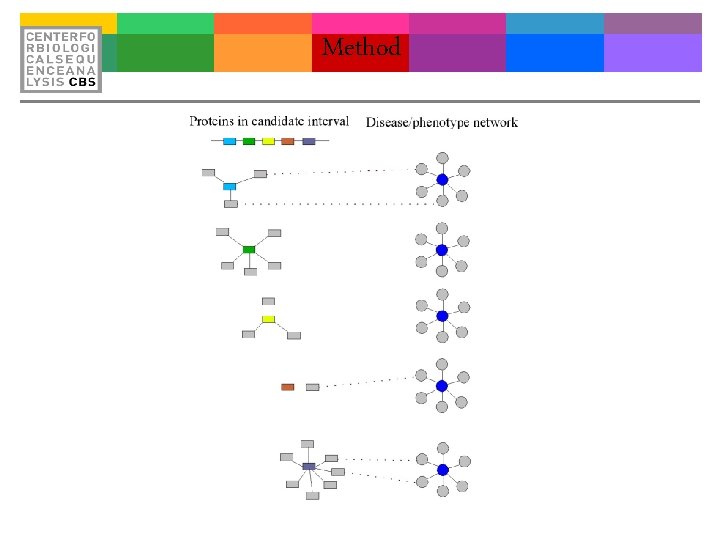
Method

Proof of Concept Input all critical intervals in OMIM (Approx 900) %125480 MAJOR AFFECTIVE DISORDER 1 %132800 MULTIPLE SELF-HEALING SQUAMOUS EPITHELIOMA %137100 IMMUNOGLOBULIN A DEFICIENCY SUSCEPTIBILITY 1 %137580 GILLES DE LA TOURETTE SYNDROME %143850 ORTHOSTATIC HYPOTENSIVE DISORDER %156240 MESOTHELIOMA, MALIGNANT %157900 MOEBIUS SYNDROME 1 %177900 PSORIASIS SUSCEPTIBILITY 1 %209850 AUTISM %252350 MOYA DISEASE 1 %608631 ASPERGER SYNDROME, SUSCEPTIBILITY TO, 2 ; ; ASPG 2 %301845 BAZEX SYNDROME; BZX %608389 BRANCHIOOTIC SYNDROME 3 %600175 SPINAL MUSCULAR ATROPHY %600318 DIABETES MELLITUS, INSULIN-DEPENDENT, 3; IDDM 3 ; ; INSULIN-DEPENDENT DIABETES MELLITUS 3 %601042 CHOREOATHETOSIS/SPASTICITY %601388 DIABETES MELLITUS, INSULIN-DEPENDENT, 12; IDDM 12 ; ; INSULIN-DEPENDENT DIABETES MELLITUS 12 %601493 CARDIOMYOPATHY, DILATED, 1 C; CMD 1 C %603694 DIABETES MELLITUS, NONINSULIN-DEPENDENT, 3 ; ; NIDDM 3; ; NONINSULIN-DEPENDENT DIABETES US 3 %604288 CARDIOMYOPATHY, DILATED, 1 H; CMD 1 H

Proof of Concept Input all critical intervals in OMIM (Approx 900) %125480 MAJOR AFFECTIVE DISORDER 1 %132800 MULTIPLE SELF-HEALING SQUAMOUS EPITHELIOMA %137100 IMMUNOGLOBULIN A DEFICIENCY SUSCEPTIBILITY 1 %137580 GILLES DE LA TOURETTE SYNDROME %143850 ORTHOSTATIC HYPOTENSIVE DISORDER %156240 MESOTHELIOMA, MALIGNANT %157900 MOEBIUS SYNDROME 1 %177900 PSORIASIS SUSCEPTIBILITY 1 %209850 AUTISM %252350 MOYA DISEASE 1 %608631 ASPERGER SYNDROME, SUSCEPTIBILITY TO, 2 ; ; ASPG 2 %301845 BAZEX SYNDROME; BZX %608389 BRANCHIOOTIC SYNDROME 3 14 q 23. 1 SIX 1 %600175 SPINAL MUSCULAR ATROPHY %600318 DIABETES MELLITUS, INSULIN-DEPENDENT, 3; IDDM 3 ; ; INSULIN-DEPENDENT DIABETES MELLITUS 3 %601042 CHOREOATHETOSIS/SPASTICITY %601388 DIABETES MELLITUS, INSULIN-DEPENDENT, 12; IDDM 12 ; ; INSULIN-DEPENDENT DIABETES MELLITUS 12 %601493 CARDIOMYOPATHY, DILATED, 1 C; CMD 1 C 10 q 21 -q 23 VINC_HUMAN %603694 DIABETES MELLITUS, NONINSULIN-DEPENDENT, 3 ; ; NIDDM 3; ; NONINSULIN-DEPENDENT DIABETES US 3 %604288 CARDIOMYOPATHY, DILATED, 1 H; CMD 1 H
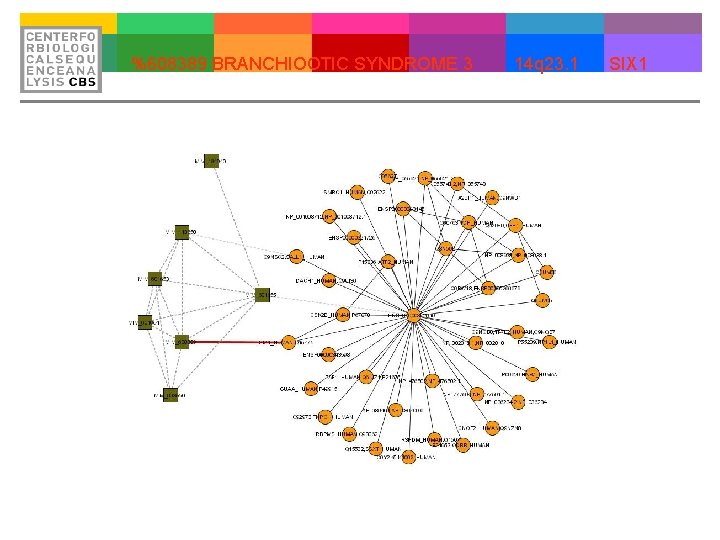
%608389 BRANCHIOOTIC SYNDROME 3 14 q 23. 1 SIX 1

Proof of Concept SIX 1 mutations cause branchio-oto-renal syndrome by disruption of EYA 1 -SIX 1 -DNA complexes. Ruf RG, Xu PX, Silvius D, Otto EA, Beekmann F, Muerb UT, Kumar S, Neuhaus TJ, Kemper MJ, Raymond RM Jr, Brophy PD, Berkman J, Gattas M, Hyland V, Ruf EM, Schwartz C, Chang EH, Smith RJ, Stratakis CA, Weil D, Petit C, Hildebrandt F. Department of Pediatrics, University of Michigan, Ann Arbor, MI 48109, USA. Urinary tract malformations constitute the most frequent cause of chronic renal failure in the first two decades of life. Branchio-otic (BO) syndrome is an autosomal dominant developmental disorder characterized by hearing loss. In branchio-oto-renal (BOR) syndrome, malformations of the kidney or urinary tract are associated. Haploinsufficiency for the human gene EYA 1, a homologue of the Drosophila gene eyes absent (eya), causes BOR and BO syndromes. We recently mapped a locus for BOR/BO syndrome (BOS 3) to human chromosome 14 q 23. 1. Within the 33 -megabase critical genetic interval, we located the SIX 1, SIX 4, and SIX 6 genes, which act within a genetic network of EYA and PAX genes to regulate organogenesis. These genes, therefore, represented excellent candidate genes for BOS 3. By direct sequencing of exons, we identified three different SIX 1 mutations in four BOR/BO kindreds, thus identifying SIX 1 as a gene causing BOR and BO syndromes. To elucidate how these mutations cause disease, we analyzed the functional role of these SIX 1 mutations with respect to protein-protein and protein-DNA interactions. We demonstrate that all three mutations are crucial for Eya 1 -Six 1 interaction, and the two mutations within the homeodomain region are essential for specific Six 1 -DNA binding. Identification of SIX 1 mutations as causing BOR/BO offers insights into the molecular basis of otic and renal developmental diseases in humans. PMID: 15141091 [Pub. Med - indexed for MEDLINE]
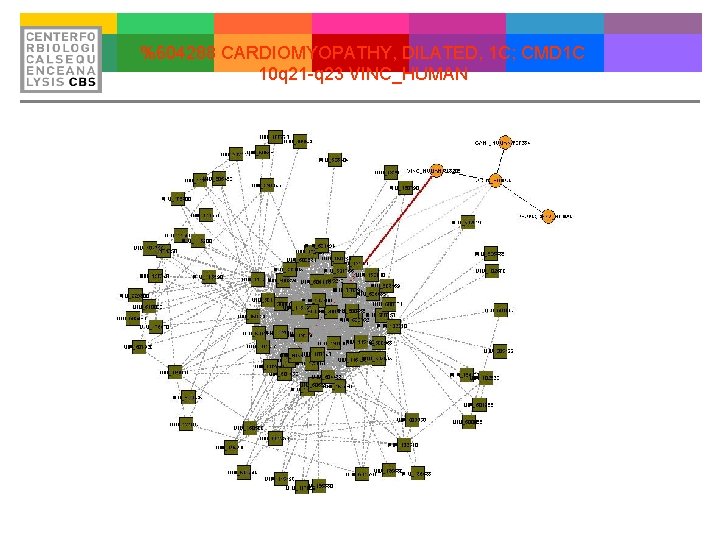
%604288 CARDIOMYOPATHY, DILATED, 1 C; CMD 1 C 10 q 21 -q 23 VINC_HUMAN

Proof of Concept Metavinculin mutations alter actin interaction in dilated cardiomyopathy. Olson TM, Illenberger S, Kishimoto NY, Huttelmaier S, Keating MT, Jockusch BM. Department of Pediatrics and the Division of Cardiology, University of Utah, Salt Lake City, Utah, USA. olson. timothy@mayo. edu BACKGROUND: Vinculin and its isoform metavinculin are protein components of intercalated discs, structures that anchor thin filaments and transmit contractile force between cardiac myocytes. We tested the hypothesis that heritable dysfunction of metavinculin may contribute to the pathogenesis of dilated cardiomyopathy (DCM). METHODS AND RESULTS: We performed mutational analyses of the metavinculin-specific exon of vinculin in 350 unrelated patients with DCM. One missense mutation (Arg 975 Trp) and one 3 -bp deletion (Leu 954 del) were identified. These mutations involved conserved amino acids, were absent in 500 control individuals, and significantly altered metavinculin-mediated cross-linking of actin filaments in an in vitro assay. Ultrastructural examination was performed in one patient (Arg 975 Trp), revealing grossly abnormal intercalated discs. A potential risk-conferring polymorphism (Ala 934 Val), identified in one DCM patient and one control individual, had a less pronounced effect on actin filament cross-linking. CONCLUSIONS: These data provide genetic and functional evidence for vinculin as a DCM gene and suggest that metavinculin plays a critical role in cardiac structure and function. Disruption of force transmission at the thin filament-intercalated disc interface is the likely mechanism by which mutations in metavinculin may lead to DCM.
Disease Gene Finding. Summery Background Why do we want to find disease genes, how has it been done until now? Networks – deducing functional relationships from network theory Networks Biological networks Functional modules / network clusters Phenotype association Grouping disorders based on their phenotype. Biological implications of phenotype clusters. Method and examples Combining network theory and phenotype associations in an automated large scale disease gene finding platform Proof of concept.
Hopes & Dreams for the future Disease Genes -Treasures in a genomic jungle. CBS - the Lara Croft of disease gene finding.

Acknowledgements Disease Gene Finding group at CBS: Olga Rigina : Database handling, Computer Scientist Olof Karlberg : Programmer, Pharmacologist Zenia M. Larsen : Expert in diabetes and related disorders, Engineer Páll Ísólfur Ólason : Engineer, data flow, text mining. Kasper Lage : Proteomics, genomics, diseases, Human Biologist Anders Hinsby : Proteomics, mass spec. expert, Human Biologist
- Slides: 41