Direct Electronic Identification of Oligonucleotides with Inelastic Electron








- Slides: 36

Direct Electronic Identification of Oligonucleotides with Inelastic Electron Tunneling Spectroscopy John Lund, Declan Ryan, Ranjana Mehta, Maryam Rahimi and Babak A. Parviz Center of Excellence in Genomic Sciences Microscale Life Sciences Center University of Washington USA

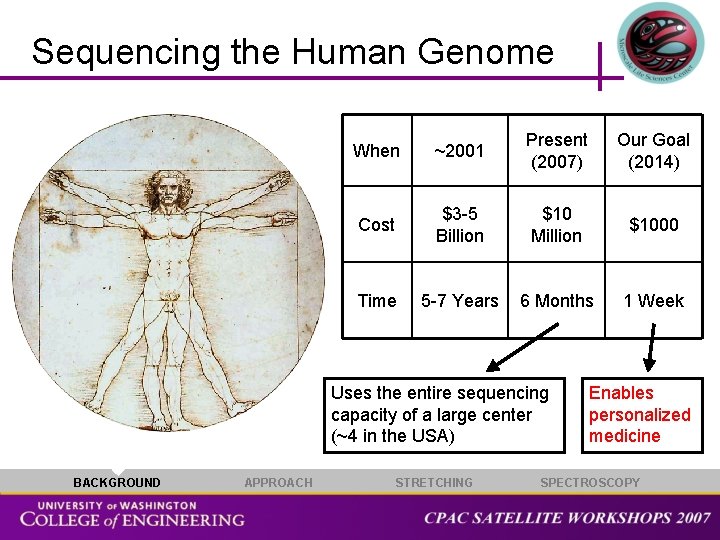
Sequencing the Human Genome When ~2001 Present (2007) Our Goal (2014) Cost $3 -5 Billion $10 Million $1000 Time 5 -7 Years 6 Months 1 Week Uses the entire sequencing capacity of a large center (~4 in the USA) BACKGROUND APPROACH STRETCHING Enables personalized medicine SPECTROSCOPY

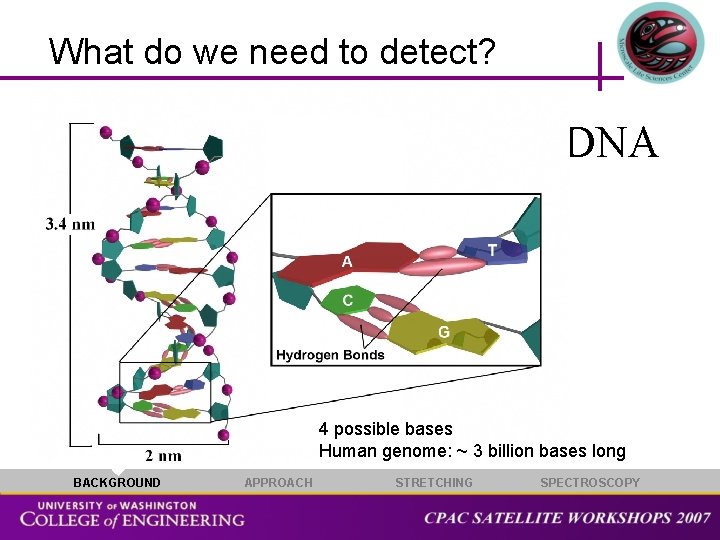
What do we need to detect? DNA 4 possible bases Human genome: ~ 3 billion bases long BACKGROUND APPROACH STRETCHING SPECTROSCOPY

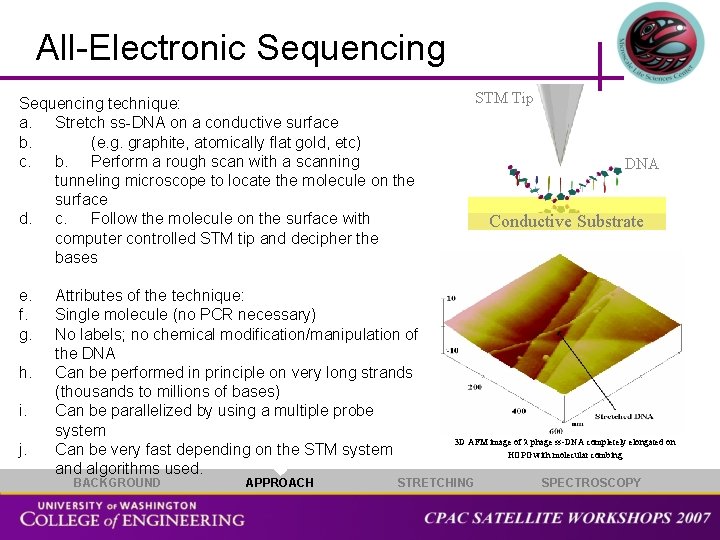
All-Electronic Sequencing technique: a. Stretch ss-DNA on a conductive surface b. (e. g. graphite, atomically flat gold, etc) c. b. Perform a rough scan with a scanning tunneling microscope to locate the molecule on the surface d. c. Follow the molecule on the surface with computer controlled STM tip and decipher the bases e. f. g. h. i. j. Attributes of the technique: Single molecule (no PCR necessary) No labels; no chemical modification/manipulation of the DNA Can be performed in principle on very long strands (thousands to millions of bases) Can be parallelized by using a multiple probe system Can be very fast depending on the STM system and algorithms used. BACKGROUND APPROACH STM Tip DNA Conductive Substrate 3 D AFM image of phage ss-DNA completely elongated on HOPG with molecular combing STRETCHING SPECTROSCOPY

How Fast Can STMs Work? Carbon atoms on the surface of HOPG imaged at tip speed of 40000 nm/s BACKGROUND This is equivalent to reading a whole bacterial genome in 10 seconds. APPROACH STRETCHING SPECTROSCOPY

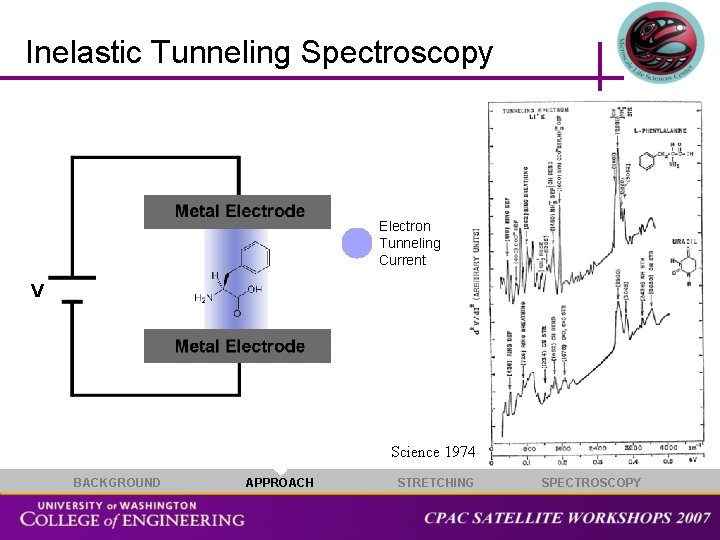
Inelastic Tunneling Spectroscopy Electron Tunneling Current V Science 1974 BACKGROUND APPROACH STRETCHING SPECTROSCOPY

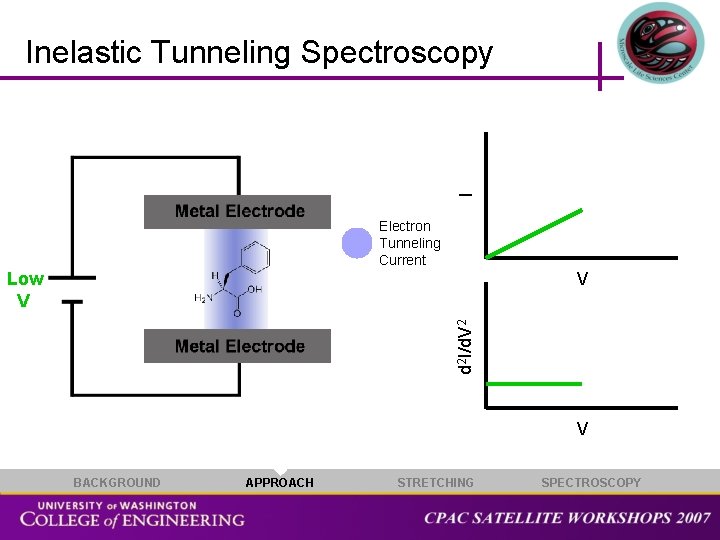
I Inelastic Tunneling Spectroscopy Electron Tunneling Current V d 2 I/d. V 2 Low V V BACKGROUND APPROACH STRETCHING SPECTROSCOPY

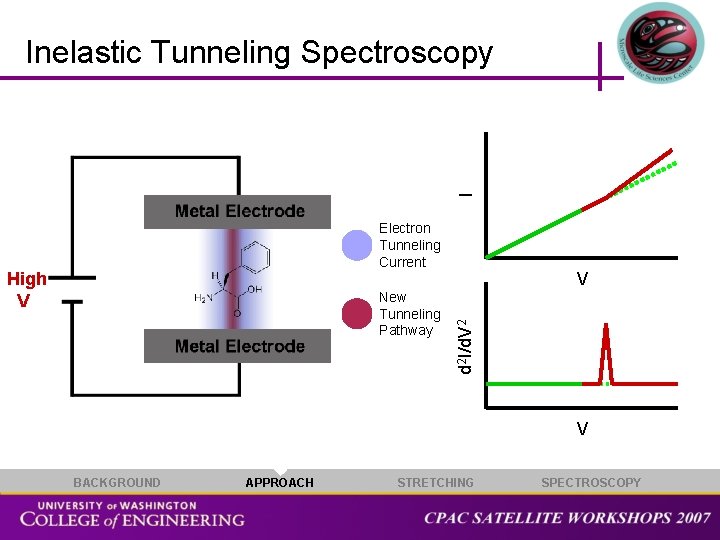
I Inelastic Tunneling Spectroscopy Electron Tunneling Current New Tunneling Pathway V d 2 I/d. V 2 High V V BACKGROUND APPROACH STRETCHING SPECTROSCOPY

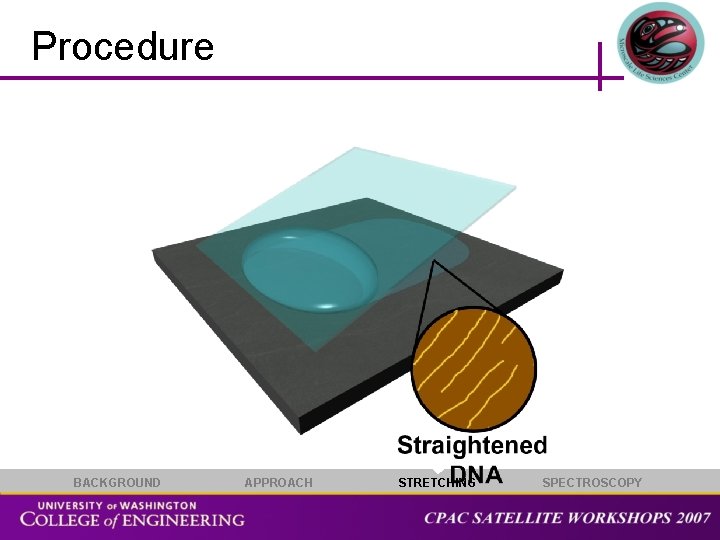
Molecular Extension • Our stretching approach employs molecular combing to orient DNA molecules on atomically flat surfaces • The interaction of the DNA and surface is tuned using coordinating ions or self-assembled monolayers • DNA molecules are stretched by a receding meniscus between a substrate DNA Molecule Droplet Meniscus Substrate BACKGROUND APPROACH STRETCHING SPECTROSCOPY

Experimental Details • We verified our technique with two phage genome systems • The Hind III digest of phage DNA, which yields 8 fragments with effective size range of 125 bp to 23 kb • Virion X 174 DNA is ss, covalently closed, circular, and 5, 386 bases in length • The ds- phage DNA was disrupted to ss-DNA by heating for 5 minutes and immediately cooling on ice • Mg. Cl 2 is used to mediate adhesion between the DNA and freshly-cleaved graphite surface BACKGROUND APPROACH STRETCHING SPECTROSCOPY

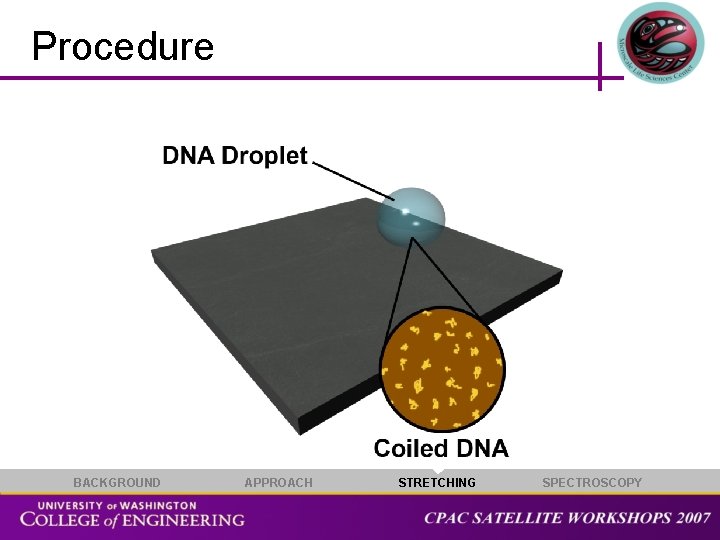
Procedure BACKGROUND APPROACH STRETCHING SPECTROSCOPY

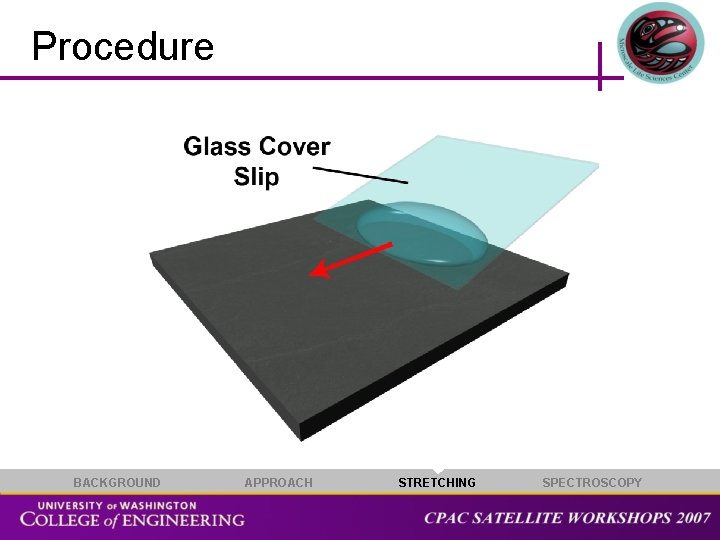
Procedure BACKGROUND APPROACH STRETCHING SPECTROSCOPY

Procedure BACKGROUND APPROACH STRETCHING SPECTROSCOPY

Procedure BACKGROUND APPROACH STRETCHING SPECTROSCOPY

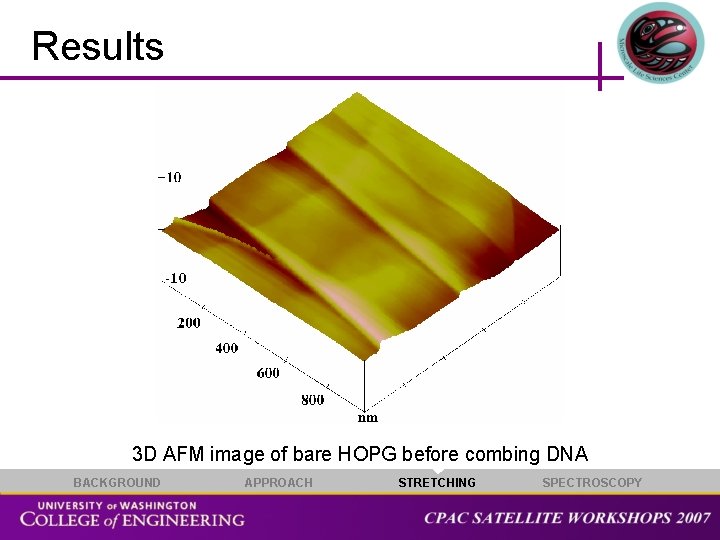
Results 3 D AFM image of bare HOPG before combing DNA BACKGROUND APPROACH STRETCHING SPECTROSCOPY

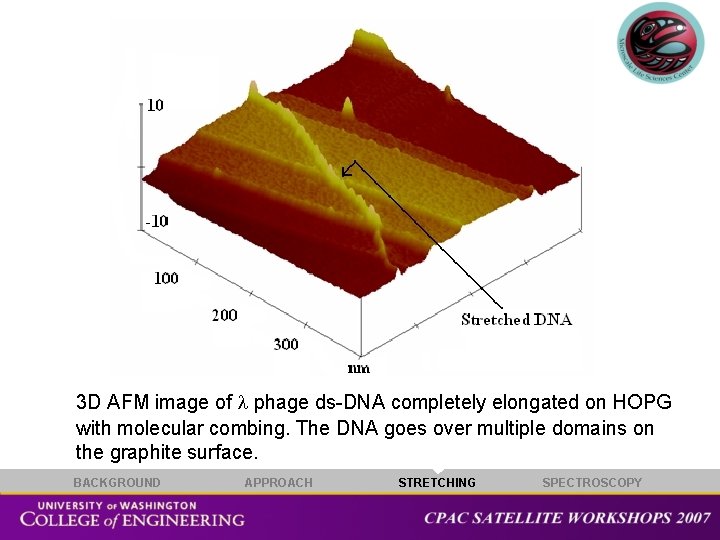
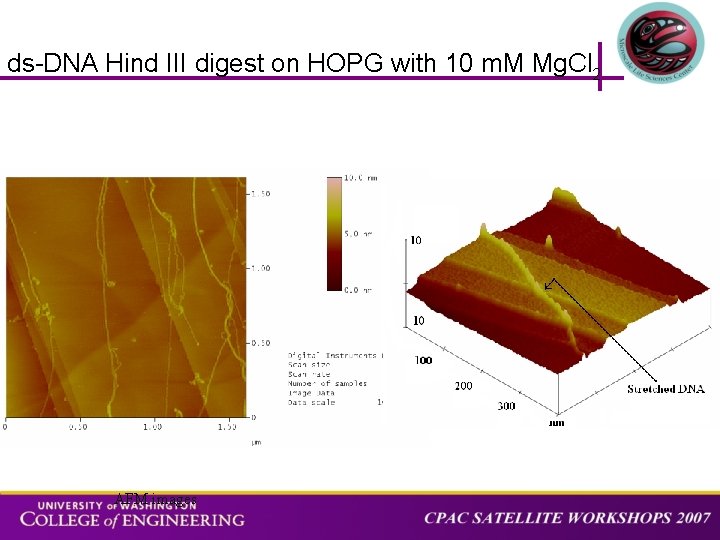
3 D AFM image of phage ds-DNA completely elongated on HOPG with molecular combing. The DNA goes over multiple domains on the graphite surface. BACKGROUND APPROACH STRETCHING SPECTROSCOPY

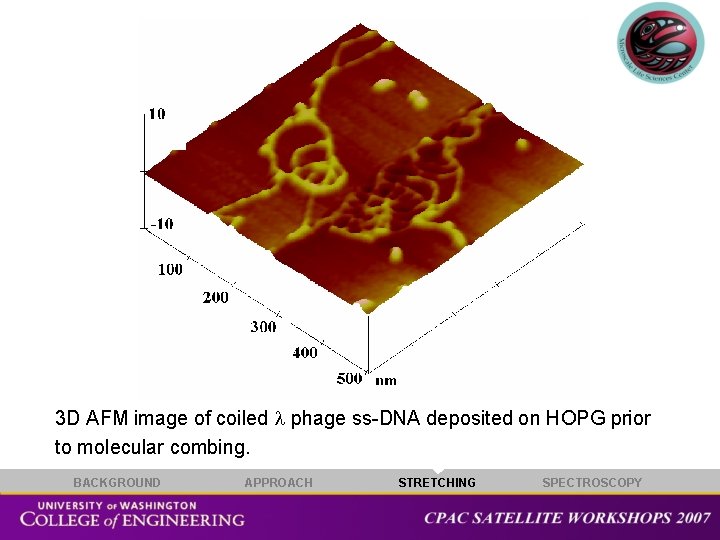
3 D AFM image of coiled phage ss-DNA deposited on HOPG prior to molecular combing. BACKGROUND APPROACH STRETCHING SPECTROSCOPY

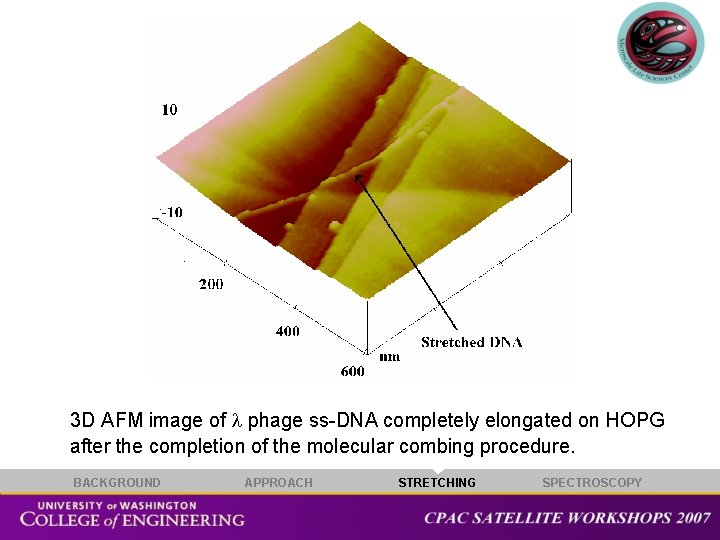
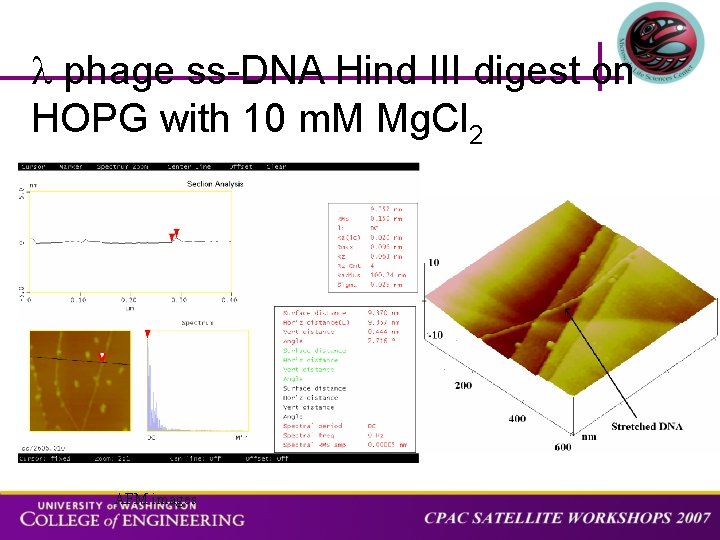
3 D AFM image of phage ss-DNA completely elongated on HOPG after the completion of the molecular combing procedure. BACKGROUND APPROACH STRETCHING SPECTROSCOPY

STM results BACKGROUND APPROACH STRETCHING SPECTROSCOPY

BACKGROUND APPROACH STRETCHING SPECTROSCOPY

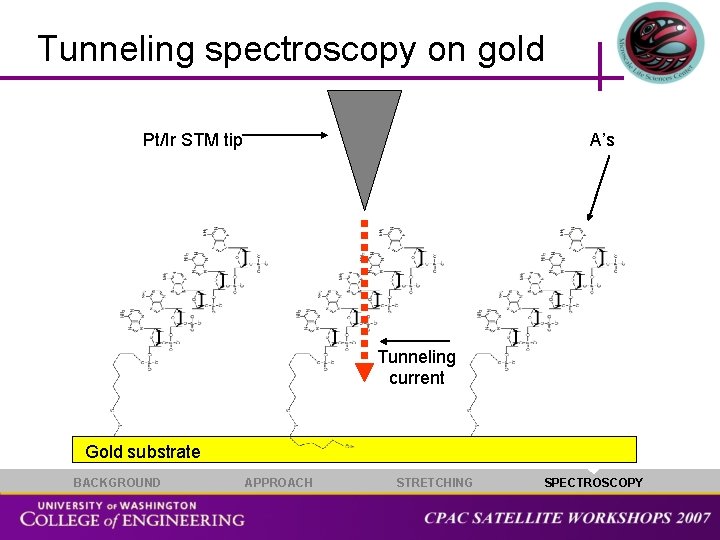
Tunneling spectroscopy on gold Pt/Ir STM tip A’s Tunneling current Gold substrate BACKGROUND APPROACH STRETCHING SPECTROSCOPY

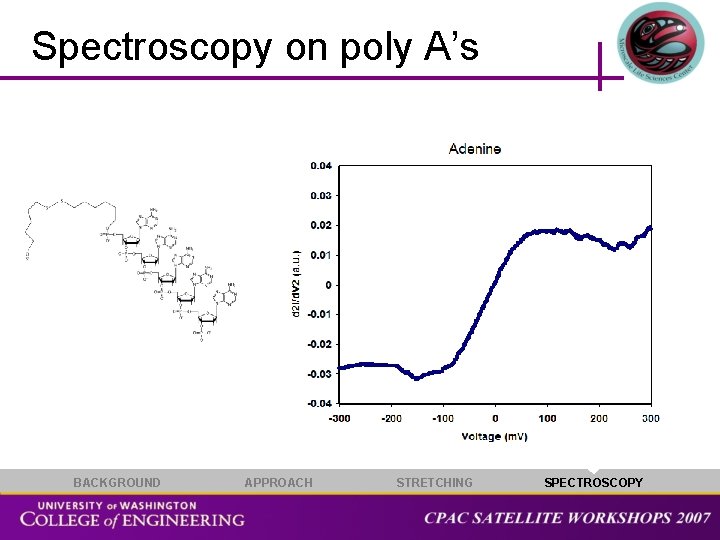
Spectroscopy on poly A’s BACKGROUND APPROACH STRETCHING SPECTROSCOPY

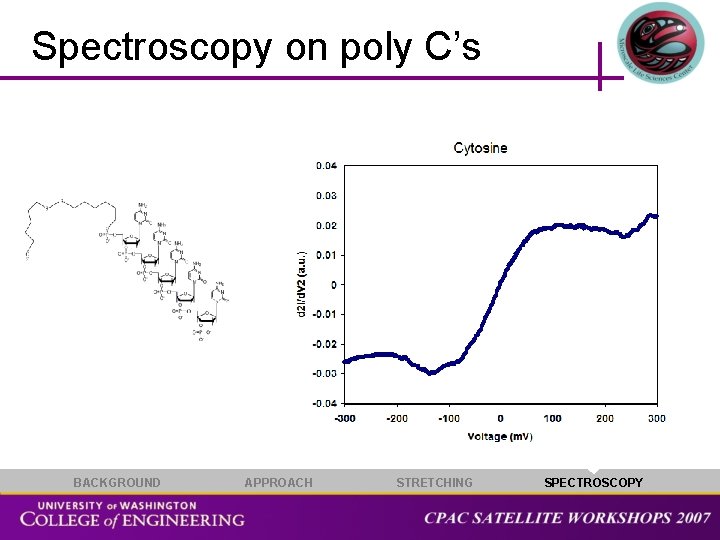
Spectroscopy on poly C’s BACKGROUND APPROACH STRETCHING SPECTROSCOPY

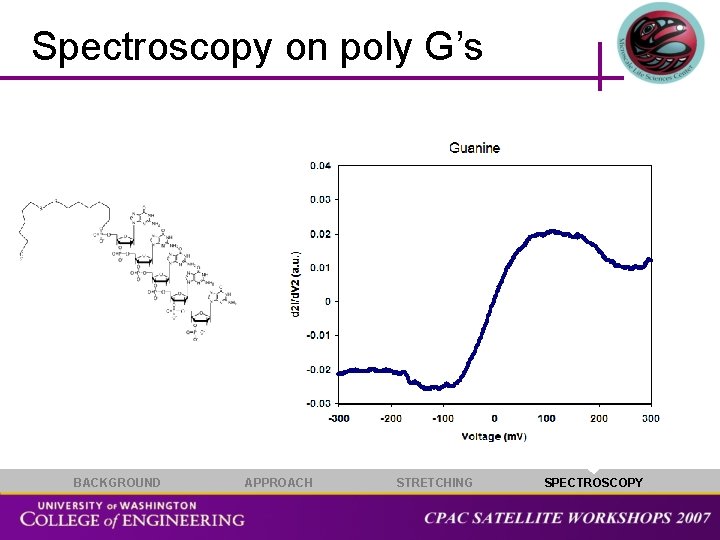
Spectroscopy on poly G’s BACKGROUND APPROACH STRETCHING SPECTROSCOPY

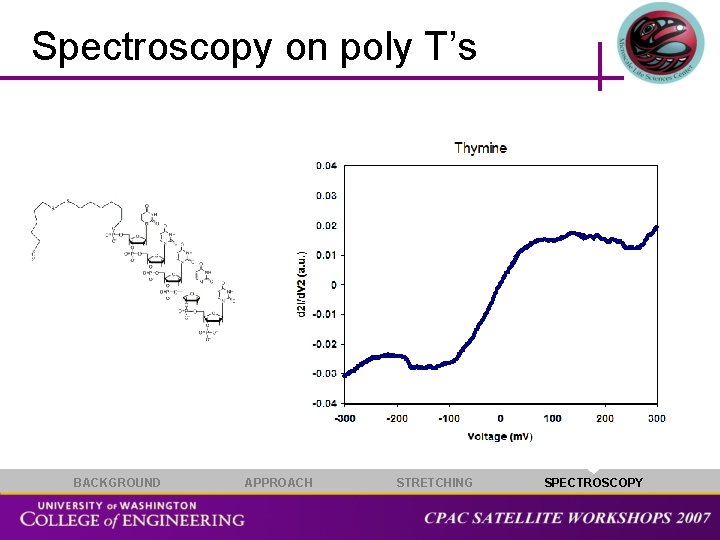
Spectroscopy on poly T’s BACKGROUND APPROACH STRETCHING SPECTROSCOPY

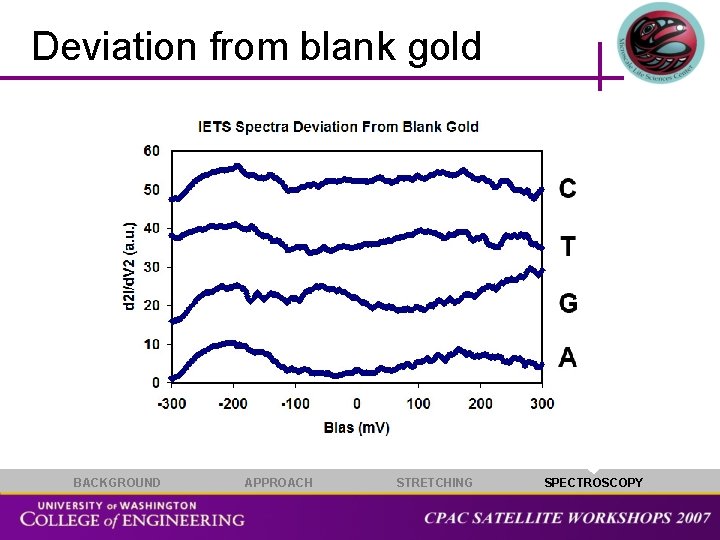
Deviation from blank gold BACKGROUND APPROACH STRETCHING SPECTROSCOPY

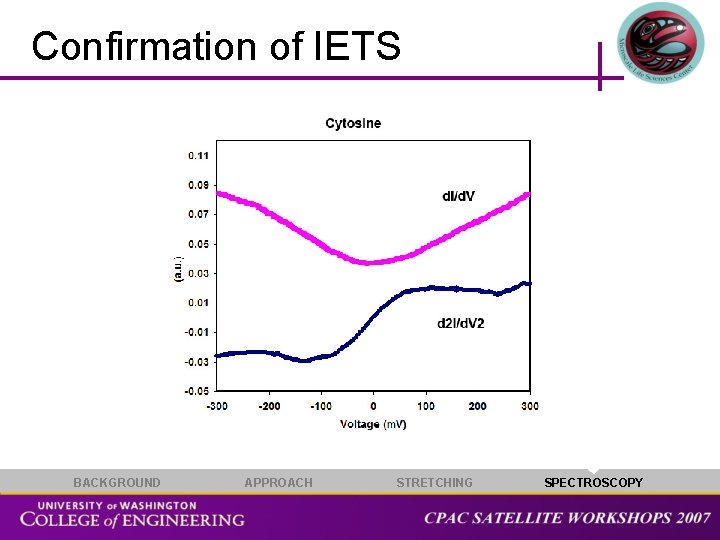
Confirmation of IETS BACKGROUND APPROACH STRETCHING SPECTROSCOPY

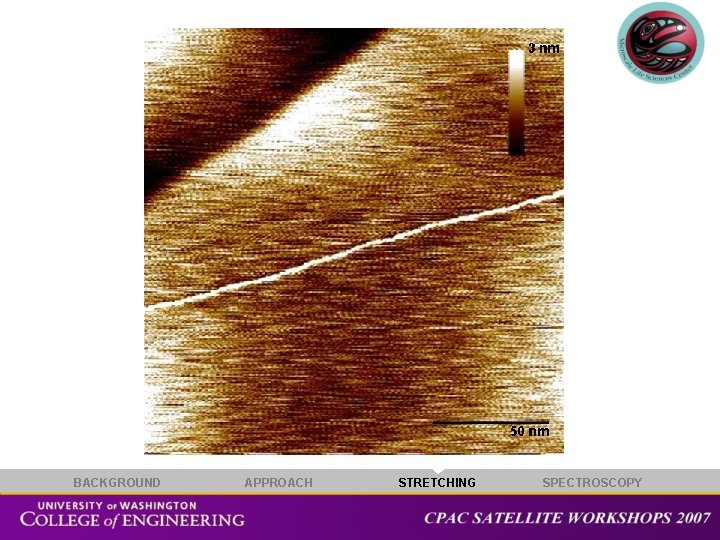
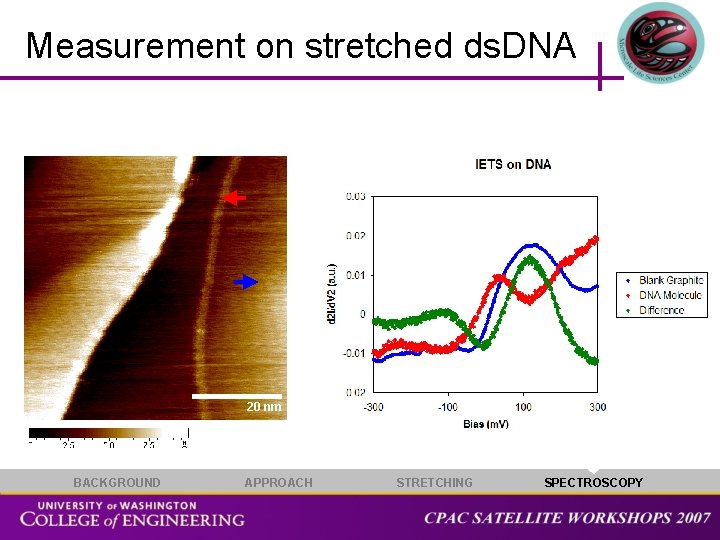
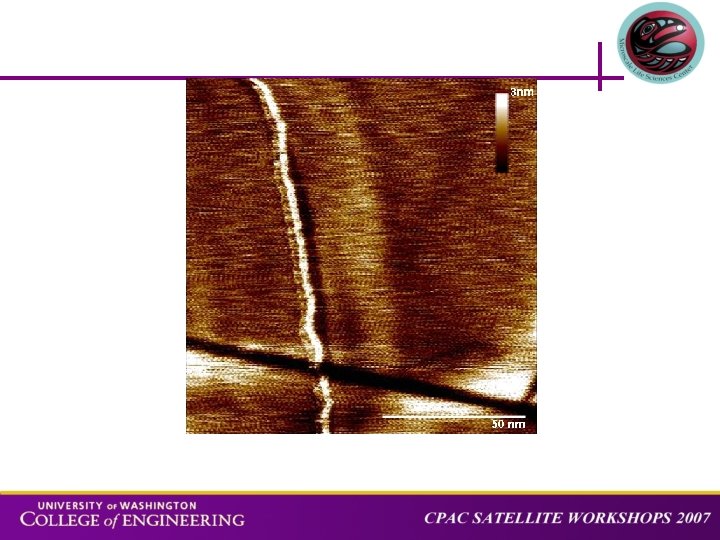
Measurement on stretched ds. DNA 20 nm BACKGROUND APPROACH STRETCHING SPECTROSCOPY

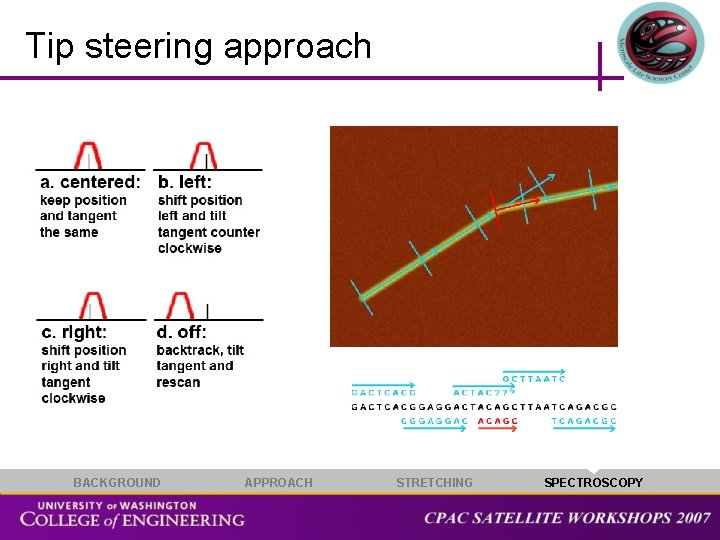
Tip steering approach BACKGROUND APPROACH STRETCHING SPECTROSCOPY

Conclusions • All-electronic genome sequencing requires costeffective and reproducible methods for extension of DNA on atomically flat surfaces • Molecular combing offers a simple and costeffective method for stretching DNA on surfaces • IETS is a promising method for identifying DNA bases on conductive substrates using STM • We have measured IETS spectra on 5 -mer DNA bases on gold and will apply our approach to sequencing strands of DNA in the future

Acknowledgments Our Research Group Members • Postdoctoral Research Fellows –Declan Ryan –Maryam Rahimi –Ranjana Mehta –Xiaorong Xiang (now at Intel) • Graduate Students –Jianchun Dong –Harvey Ho –Sam Kim (with D. Meldrum) –John Lund –Coretta Maremma –Chris Morris –Ehsan Saeedi –Angela Shum –Andrei Afanasiev –Jean Wang (with Lin) • Undergraduate Students –Lisa Oh –James Etzkorn Funding for our group: National Institutes of Health (NIH) Gordon and Betty Moore Foundation National Science Foundation (NSF) National Academies Keck Future Initiative (NAKFI) Defense Advanced Research Project Agency (DARPA) Office of Naval Research (ONR) University Initiative Fund (UIF) at UW UW Technology Gap Innovation Fund (TGIF)

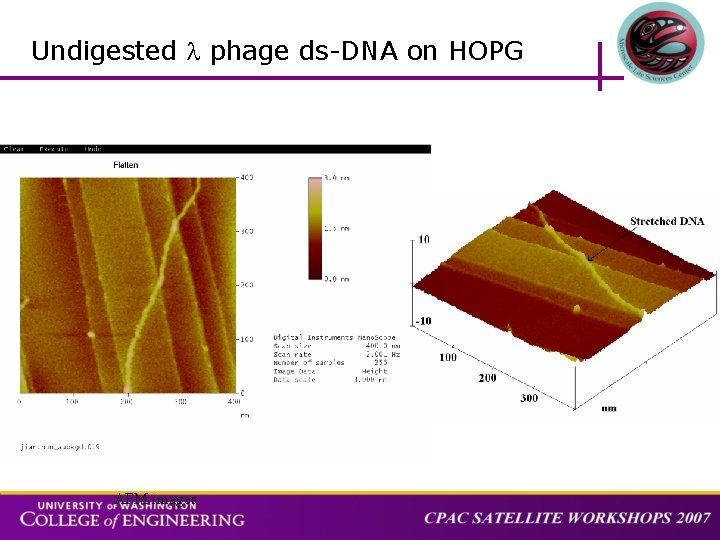
Undigested phage ds-DNA on HOPG AFM images

ds-DNA Hind III digest on HOPG with 10 m. M Mg. Cl 2 AFM images

phage ss-DNA Hind III digest on HOPG with 10 m. M Mg. Cl 2 AFM images

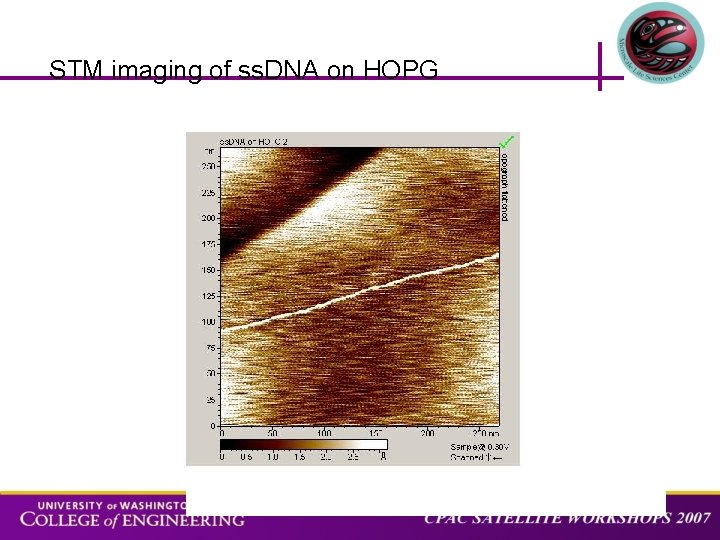
STM imaging of ss. DNA on HOPG
