Differential expression analysis with RNASeq GOn Ramp Beta

Differential expression analysis with RNA-Seq G-On. Ramp Beta Users Workshop Wilson Leung 06/2017

Outline Design considerations for RNA-Seq experiments Interpret Fast. QC results Optimize alignment parameters for HISAT Assess alignment statistics with Collect. Rna. Seq. Metrics Assemble transcripts with String. Tie Differential expression analysis with DESeq 2
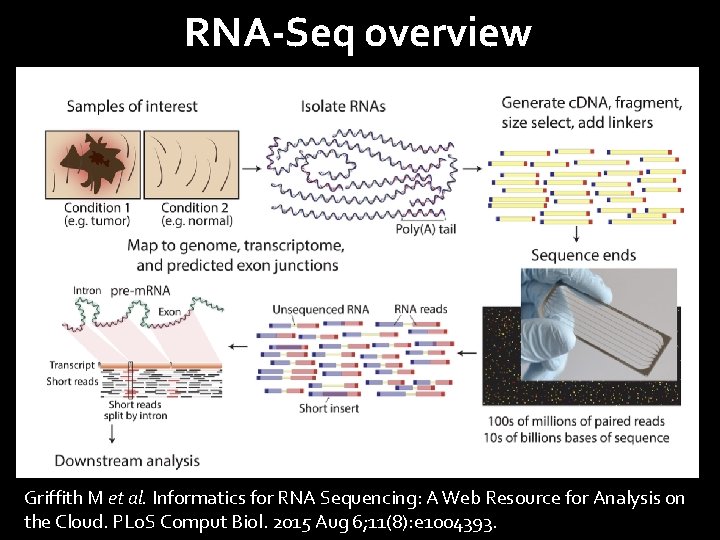
RNA-Seq overview Griffith M et al. Informatics for RNA Sequencing: A Web Resource for Analysis on the Cloud. PLo. S Comput Biol. 2015 Aug 6; 11(8): e 1004393.
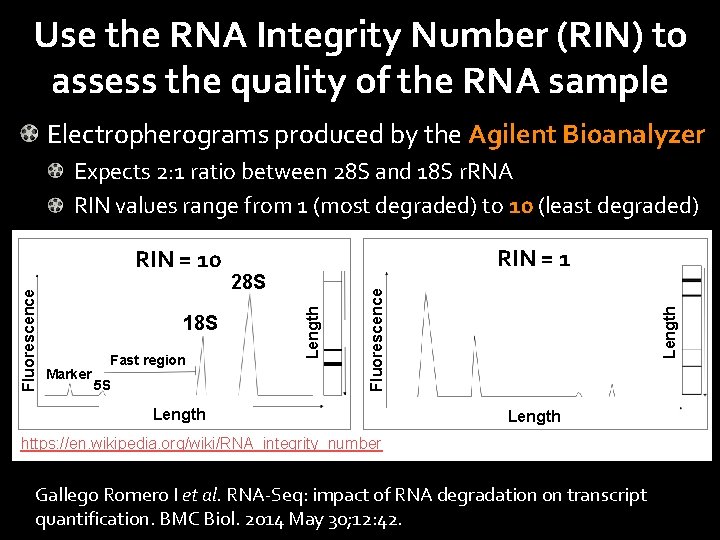
Use the RNA Integrity Number (RIN) to assess the quality of the RNA sample Electropherograms produced by the Agilent Bioanalyzer Expects 2: 1 ratio between 28 S and 18 S r. RNA RIN values range from 1 (most degraded) to 10 (least degraded) Marker 5 S Length Fast region 28 S Fluorescence 18 S RIN = 1 Length Fluorescence RIN = 10 Length https: //en. wikipedia. org/wiki/RNA_integrity_number Gallego Romero I et al. RNA-Seq: impact of RNA degradation on transcript quantification. BMC Biol. 2014 May 30; 12: 42.

Common applications of RNA-Seq Transcriptome profiling Identify novel transcripts (e. g. , gene annotations) Quantify expression levels Differential expression Different developmental stages; treatment versus control Alternative splicing Visualization and integration with other datasets Correlate with epigenomic landscape Histone modifications, DNA methylation, etc. Conesa A et al. A survey of best practices for RNA-Seq data analysis. Genome Biol. 2016 Jan 26; 17: 13.
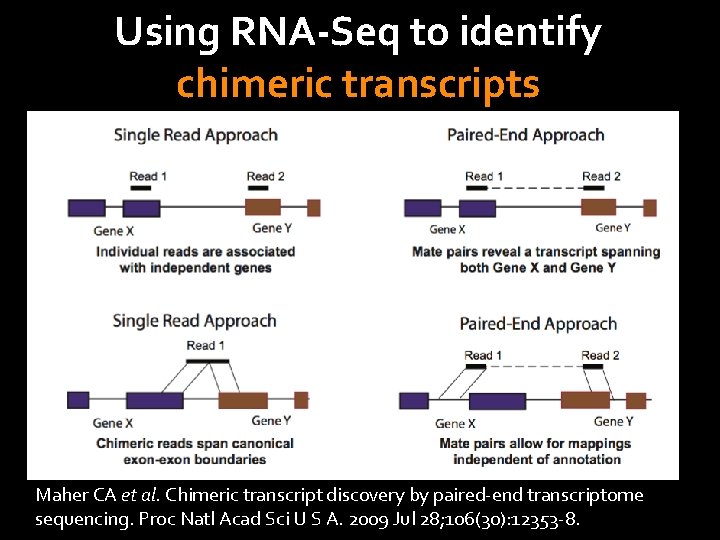
Using RNA-Seq to identify chimeric transcripts Maher CA et al. Chimeric transcript discovery by paired-end transcriptome sequencing. Proc Natl Acad Sci U S A. 2009 Jul 28; 106(30): 12353 -8.

The optimal RNA-Seq sequencing and analysis protocols depend on the goals of the study
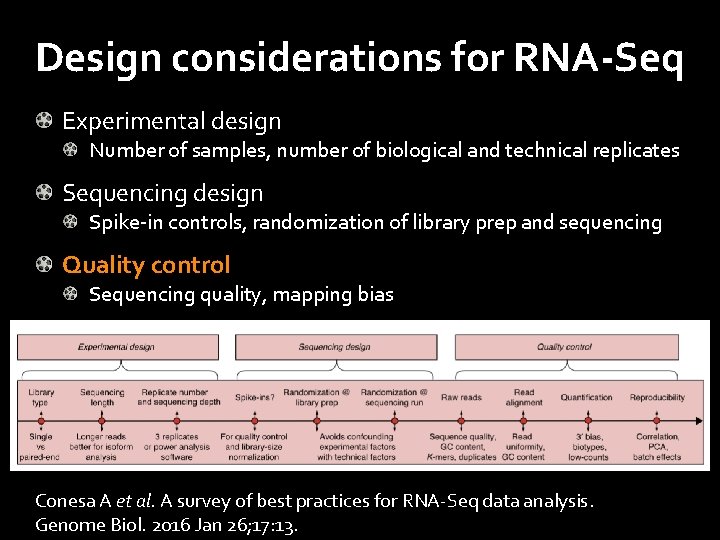
Design considerations for RNA-Seq Experimental design Number of samples, number of biological and technical replicates Sequencing design Spike-in controls, randomization of library prep and sequencing Quality control Sequencing quality, mapping bias Conesa A et al. A survey of best practices for RNA-Seq data analysis. Genome Biol. 2016 Jan 26; 17: 13.

Biological and technical replicates Biological replicates RNA from independent growth of cells and tissues Account for biological variations Technical replicates Different library preparations of the same RNA-Seq sample Account for batch effects from library preparations Sample loading, cluster amplifications, etc. ENCODE long RNA-Seq standards: https: //www. encodeproject. org/data-standards/rna-seq/long-rnas/ Blainey P et al. Points of significance: replication. Nat Methods. 2014 Sep; 11(9): 879 -80.

Recommended RNA-Seq sequencing depth based on genome size Genome Size Small (bacteria / fungi) Differential expression analysis (# reads) 5 M Detect rare transcripts / de novo assembly (# reads) Read length 30 – 65 M 50 bp Intermediate 10 M (D. melanogaster) 70 – 130 M 50 – 100 bp Large (Human) 100 – 200 M > 100 bp 15 – 25 M https: //genohub. com/next-generation-sequencing-guide/#depth 2

How many biological replicates? As many as possible… Analysis of 48 biological replicates in two conditions Requires 20 biological replicates to detect > 85% of all differentially expressed genes Recommend at least six biological replicates per condition Twelve biological replicates needed to detect smaller fold changes (≥ 0. 3 -fold difference in expression) Three biological replicates per condition can usually detect genes with ≥ 2 -fold difference in expression Schurch NJ et al. How many biological replicates are needed in an RNA-Seq experiment and which differential expression tool should you use? RNA. 2016 Jun; 22(6): 839 -51.
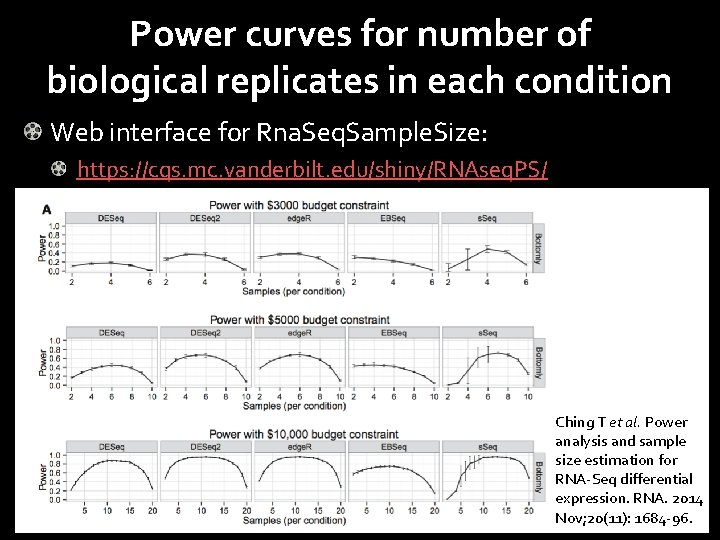
Power curves for number of biological replicates in each condition Web interface for Rna. Seq. Sample. Size: https: //cqs. mc. vanderbilt. edu/shiny/RNAseq. PS/ Power 0. 8 0. 6 FDR = 0. 05 FDR = 0. 01 0. 4 0. 2 0. 0 0 10 20 30 40 50 60 # Samples in each condition Ching T et al. Power analysis and sample size estimation for RNA-Seq differential expression. RNA. 2014 Nov; 20(11): 1684 -96.

Using Galaxy to perform RNA-Seq analysis Quality control with Fast. QC Trim low quality bases with Trimmomatic Read mapping with HISAT Transcriptome assembly with String. Tie Differential expression analysis with DESeq 2 Based on RNA-Seq tutorial developed by Mo Heydarian and Mallory Freeberg at Johns Hopkins University: https: //galaxyproject. org/tutorials/nt_rnaseq/
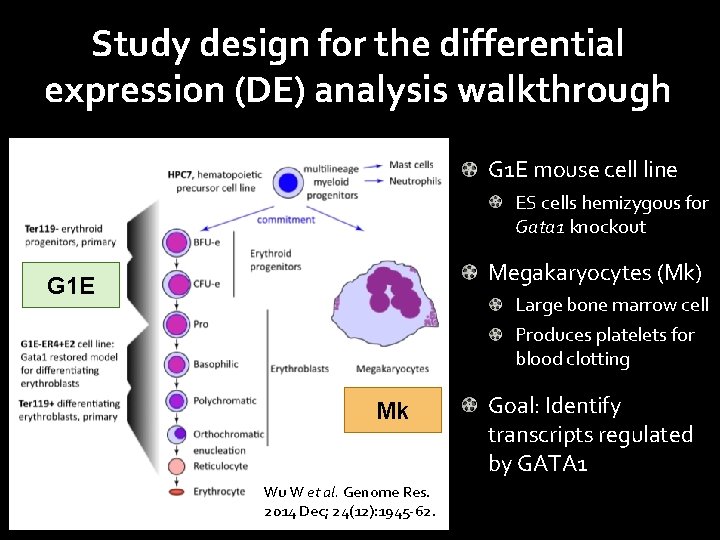
Study design for the differential expression (DE) analysis walkthrough G 1 E mouse cell line ES cells hemizygous for Gata 1 knockout Megakaryocytes (Mk) G 1 E Large bone marrow cell Produces platelets for blood clotting Mk Wu W et al. Genome Res. 2014 Dec; 24(12): 1945 -62. Goal: Identify transcripts regulated by GATA 1
Quality control with Fast. QC Determine quality encoding of fastq files Assess quality of sequencing sample Identify overrepresented sequences Adapters, potential contamination, etc.
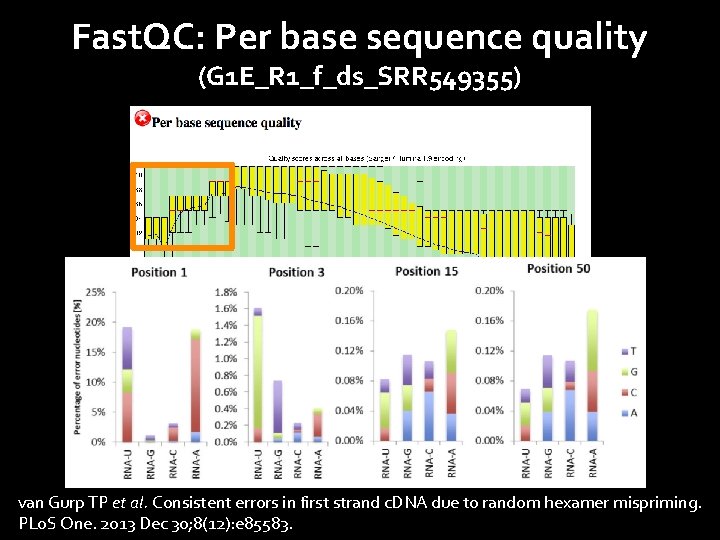
Fast. QC: Per base sequence quality (G 1 E_R 1_f_ds_SRR 549355) van Gurp TP et al. Consistent errors in first strand c. DNA due to random hexamer mispriming. PLo. S One. 2013 Dec 30; 8(12): e 85583.
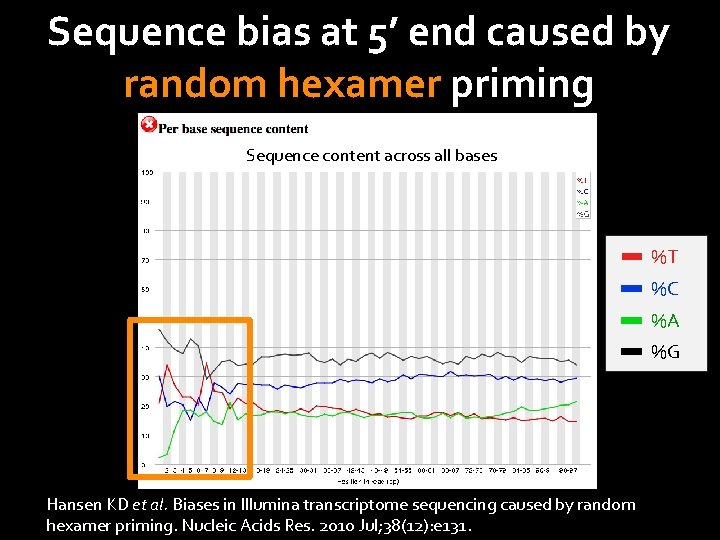
Sequence bias at 5’ end caused by random hexamer priming Sequence content across all bases %T %C %A %G Hansen KD et al. Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res. 2010 Jul; 38(12): e 131.
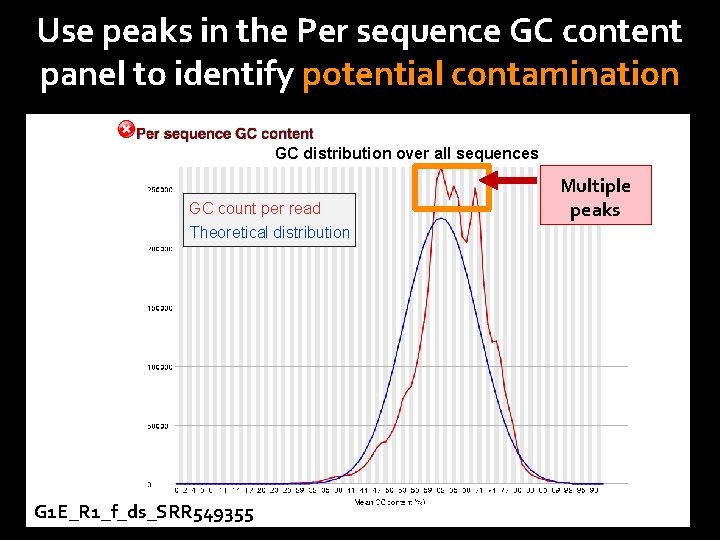
Use peaks in the Per sequence GC content panel to identify potential contamination GC distribution over all sequences GC count per read Theoretical distribution G 1 E_R 1_f_ds_SRR 549355 Multiple peaks

Investigate overrepresented sequences G 1 E Mk Fth 1
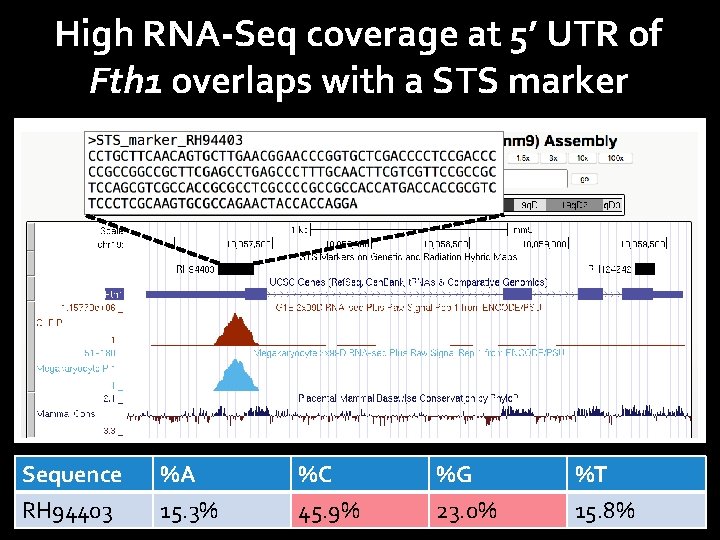
High RNA-Seq coverage at 5’ UTR of Fth 1 overlaps with a STS marker Sequence RH 94403 %A 15. 3% %C 45. 9% %G 23. 0% %T 15. 8%

RNA-Seq read mapping with HISAT Many alignment parameters available… Which parameters should be changed?

Strand-specific RNA-Seq libraries Standard libraries do not preserve strand information Prefer strand-specific RNA-Seq Transcript reconstruction and quantification Detect antisense transcripts Library prep kits available from Illumina: Tru. Seq Stranded Total RNA Sample Prep Kit Tru. Seq Stranded m. RNA Sample Prep Kit Zhao S et al. Comparison of stranded and non-stranded RNA-Seq transcriptome profiling and investigation of gene overlap. BMC Genomics. 2015 Sep 3; 16: 675.

Orientations of RNA-Seq reads depend on the paired-end protocol http: //sailfish. readthedocs. io/en/master/library_type. html Tru. Seq Strand-Specific Total RNA: First Strand (R/RF) fr-firststrand (F 2 R 1) Nu. GEN Encore: Second Strand (F/FR) fr-secondstrand (F 1 R 2) Nu. GEN Ovation V 2: FR Unstranded fr-unstranded See Supplemental Table S 5 in Griffith M et al. PLo. S Comput Biol. 2015 Aug 6; 11(8): e 1004393.

Use infer_experiment. py to infer the design of the RNA-Seq experiment Part of the RSe. QC package: http: //rseqc. sourceforge. net/ Map RNA-Seq reads using default parameters Run infer_experiment. py to infer experimental design Map RNA-Seq reads using the inferred parameters Available through the Galaxy Tool Shed (rseqc):

Common changes to HISAT alignment parameters Minimum and maximum intron lengths Specify strand-specific information GTF file with known splice sites Use known gene annotations to guide read mapping if available Transcriptome assembly reporting
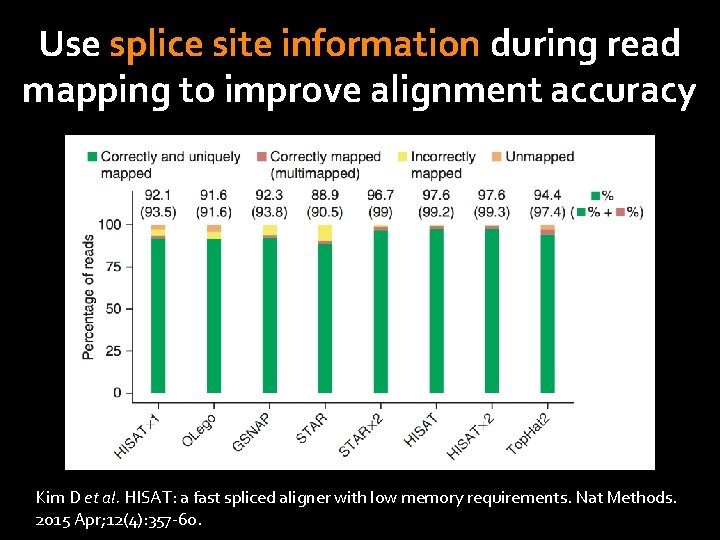
Use splice site information during read mapping to improve alignment accuracy Kim D et al. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015 Apr; 12(4): 357 -60.

RNA-Seq alignment strategy for multiple samples Map reads from each RNA-Seq sample separately Use the --novel-splicesite-outfile option to report splice sites identified in each sample Combine splice junctions from all samples under all conditions into a single splice junction file Filter splice junctions by score Retain junctions that appear in multiple biological replicates Map reads from each RNA-Seq sample using the combined splice junctions file Use the --novel-splicesite-infile option
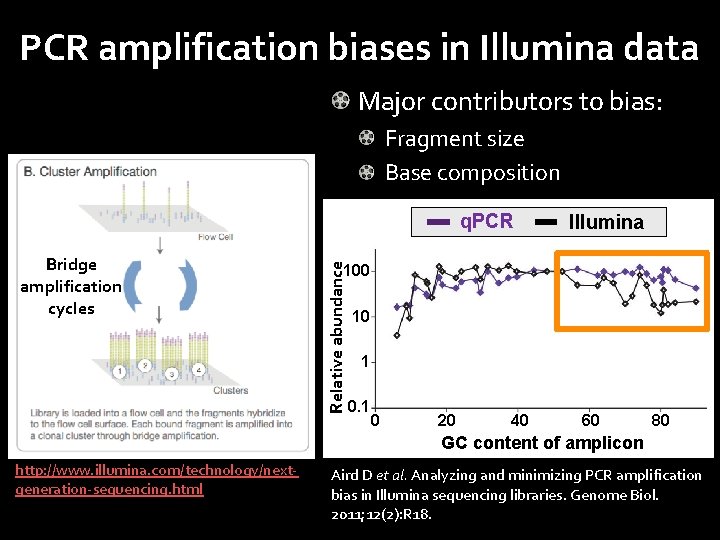
PCR amplification biases in Illumina data Major contributors to bias: Fragment size Base composition Bridge amplification cycles Relative abundance q. PCR Illumina 100 10 1 0 20 40 60 80 GC content of amplicon http: //www. illumina. com/technology/nextgeneration-sequencing. html Aird D et al. Analyzing and minimizing PCR amplification bias in Illumina sequencing libraries. Genome Biol. 2011; 12(2): R 18.

Identify duplicate reads based on sequence alignments Assumption: Rare for different sequenced fragments to have the same start and end positions The Picard tool Mark. Duplicates only considers the start position RNA-Seq data violates this assumption: Reads map to a smaller portion of the genome Probability of reads with the same start position depends on gene length and expression levels Recommendation: Mark potential duplicates to verify that all samples have similar levels of “duplication” Retain duplicate reads in differential expression analyses Williams AG et al. RNA-Seq Data: Challenges in and Recommendations for Experimental Design and Analysis. Curr Protoc Hum Genet. 2014 Oct 1; 83: 11. 13. 1 -20.
DEMO: Mark optical and PCR duplicates using the Picard tool Mark. Duplicates

Assess RNA-Seq read alignments with Collect. Rna. Seq. Metrics Requires gene annotations: Gene annotations in GTF / GFF format UCSC Genes in ref. Flat format (from the UCSC Table Browser) BAM dataset collection Reference genome (mm 10) Gene annotations in ref. Flat format r. RNAs in interval list format Second read transcription strand
DEMO: Use Collect. Rna. Seq. Metrics to assess RNA-Seq alignments
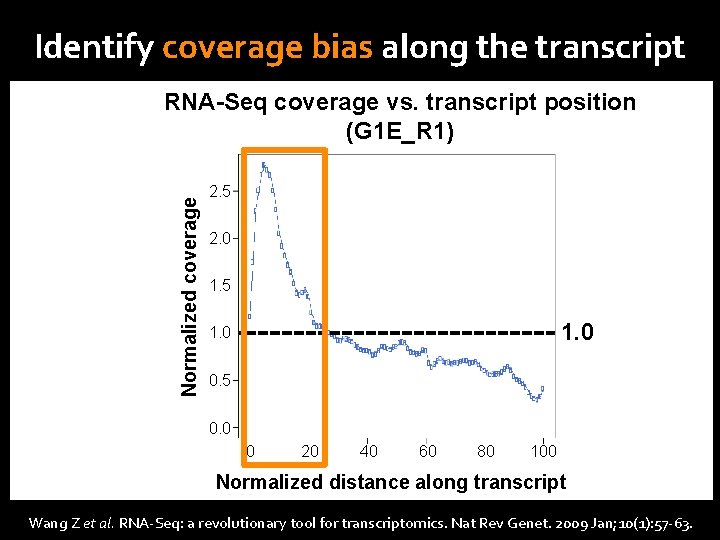
Identify coverage bias along the transcript Normalized coverage RNA-Seq Read Count RNA-Seq coverage vs. transcript position (G 1 E_R 1) 5’ 2. 5 c. DNA fragmentation DNase I treatment or sonication 2. 0 1. 5 RNA fragmentation 1. 0 RNA hydrolysis or nebulization 0. 5 Gene Span 0. 0 (5, 099 genes) 0 20 3’ 40 60 80 100 Normalized distance along transcript Wang Z et al. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009 Jan; 10(1): 57 -63.

Two common approaches to RNA-Seq assembly Reference-based assembly Map RNA-Seq reads against a reference genome Examples: Top. Hat 2, HISAT Assemble transcripts from mapped RNA-Seq reads Examples: Cufflinks, String. Tie De novo transcriptome assembly Assemble transcripts from RNA-Seq reads Examples: Oases, Trinity More computationally expensive Merge assemblies produced by different parameters
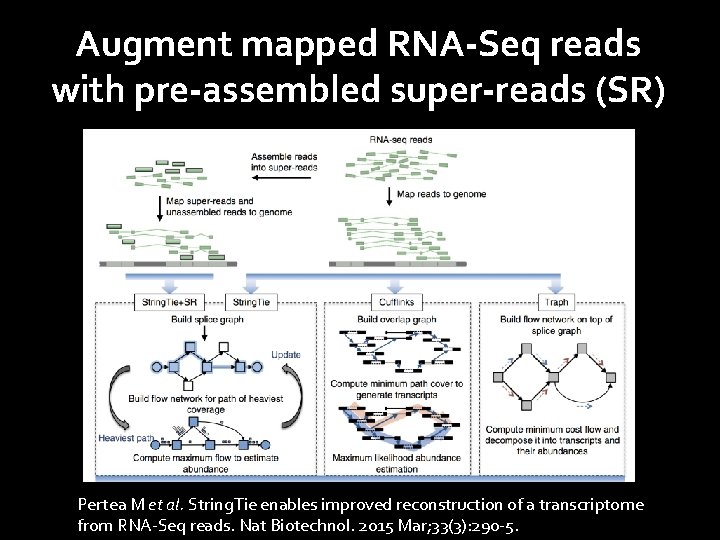
Augment mapped RNA-Seq reads with pre-assembled super-reads (SR) Pertea M et al. String. Tie enables improved reconstruction of a transcriptome from RNA-Seq reads. Nat Biotechnol. 2015 Mar; 33(3): 290 -5.
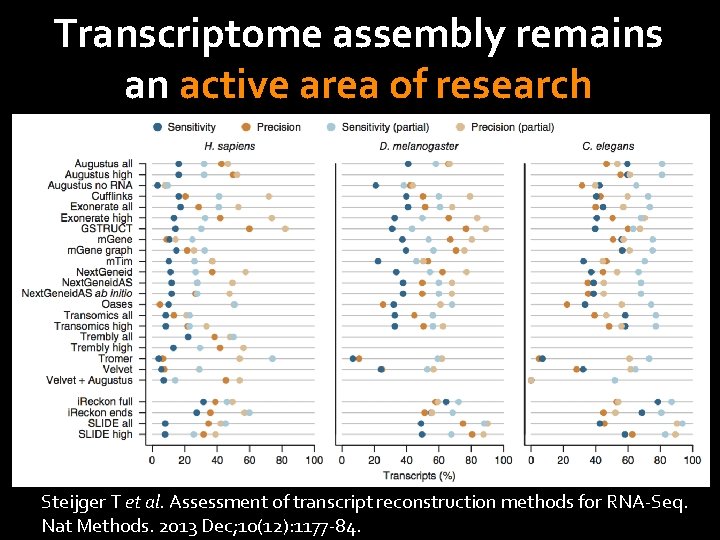
Transcriptome assembly remains an active area of research Korf I. Genomics: the state of the art in RNA-Seq analysis. Nat Methods. 2013 Dec; 10(12): 1165 -6. Steijger T et al. Assessment of transcript reconstruction methods for RNA-Seq. Nat Methods. 2013 Dec; 10(12): 1177 -84.
DEMO: Assemble transcripts from mapped RNA-Seq reads using String. Tie

Metrics for quantifying gene expression levels RPKM Reads Per Kilobase per Million mapped reads Normalize relative to sequencing depth and gene length FPKM Similar to RPKM but count DNA fragments instead of reads Used in paired end RNA-Seq experiments to avoid bias TPM Transcripts Per Million Normalize for gene length, then normalize by sequencing depth Wagner GP et al. Measurement of m. RNA abundance using RNA-Seq data: RPKM measure is inconsistent among samples. Theory Biosci. 2012 Dec; 131(4): 281 -5.

RPKM, FPKM, and TPM compare relative gene expression levels within a sample Most differential expression analysis tools use read counts as input
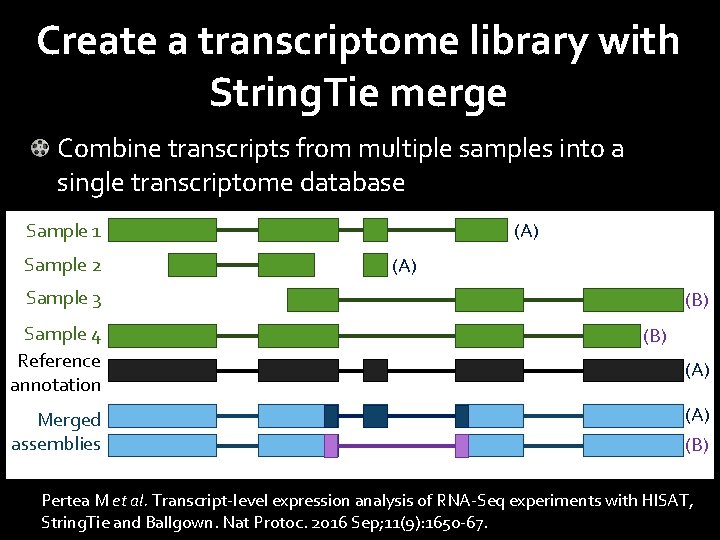
Create a transcriptome library with String. Tie merge Combine transcripts from multiple samples into a single transcriptome database Sample 1 Sample 2 (A) Sample 3 Sample 4 Reference annotation Merged assemblies (B) (A) (B) Pertea M et al. Transcript-level expression analysis of RNA-Seq experiments with HISAT, String. Tie and Ballgown. Nat Protoc. 2016 Sep; 11(9): 1650 -67.
DEMO: Use String. Tie merge to construct a transcriptome database for G 1 E and Mk

Differential expression analysis tools Compare genes expression levels: DESeq 2 (http: //bioconductor. org/packages/release/bioc/html/DESeq 2. html) edge. R (http: //bioconductor. org/packages/release/bioc/html/edge. R. html) Compare transcripts expression levels: Cuffdiff (http: //cole-trapnell-lab. github. io/cufflinks/cuffdiff/) Ballgown (http: //bioconductor. org/packages/release/bioc/html/ballgown. html) MISO (http: //genes. mit. edu/burgelab/miso/index. html) Salmon (https: //combine-lab. github. io/salmon/) Compare both genes and transcripts expression levels: RSEM + EBSeq (http: //deweylab. biostat. wisc. edu/rsem/README. html)

Count the number of reads that overlap with each gene using htseq-count Three modes of overlap resolution: union intersection_strict intersection_nonempty htseq-count ignores multi-mapped reads http: //www-huber. embl. de/HTSeq/doc/overview. html

feature. Counts is a faster alternative to htseq-count Liao Y et al. feature. Counts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014 Apr 1; 30(7): 923 -30.

Issues with the GTF files produced by the UCSC Table Browser The gene_id and transcript_id fields in the GTF file have the same values GTF format stipulates that different isoforms derived from the same gene should have the same gene_id Two potential solutions: Download gene. Pred file from the UCSC Genome Browser web site, then use gene. Pred. To. Gtf to create the GTF file http: //genomewiki. ucsc. edu/index. php/Genes_in_gtf_or_gff_format Use the GTF files from Ensembl http: //www. ensembl. org/info/data/ftp/index. html
DEMO: Calculate the read count for each transcript using feature. Counts
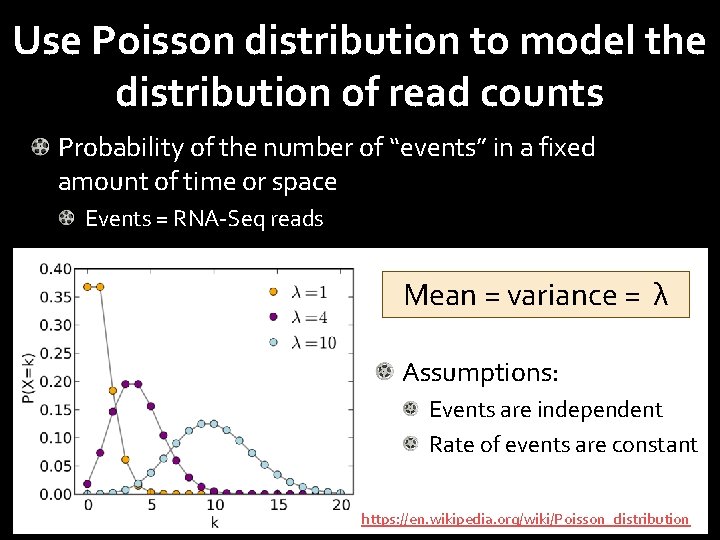
Use Poisson distribution to model the distribution of read counts Probability of the number of “events” in a fixed amount of time or space Events = RNA-Seq reads Mean = variance = λ Assumptions: Events are independent Rate of events are constant https: //en. wikipedia. org/wiki/Poisson_distribution
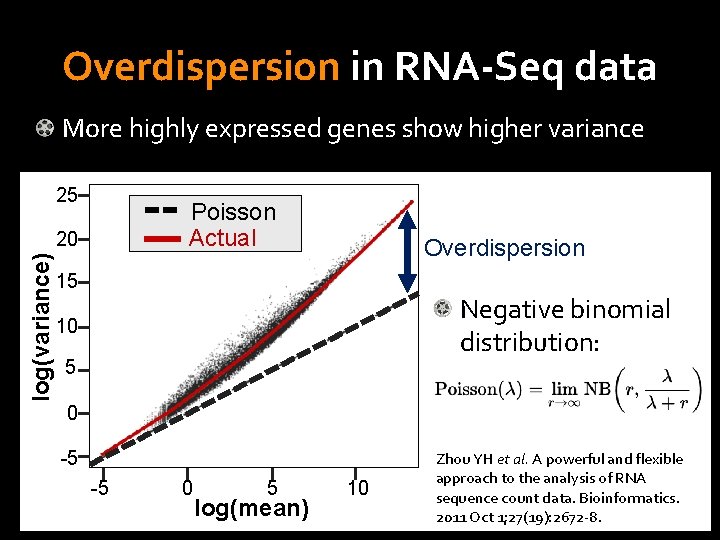
Overdispersion in RNA-Seq data More highly expressed genes show higher variance 25 Poisson Actual log(variance) 20 Overdispersion 15 Negative binomial distribution: 10 5 0 -5 -5 0 5 log(mean) 10 Zhou YH et al. A powerful and flexible approach to the analysis of RNA sequence count data. Bioinformatics. 2011 Oct 1; 27(19): 2672 -8.
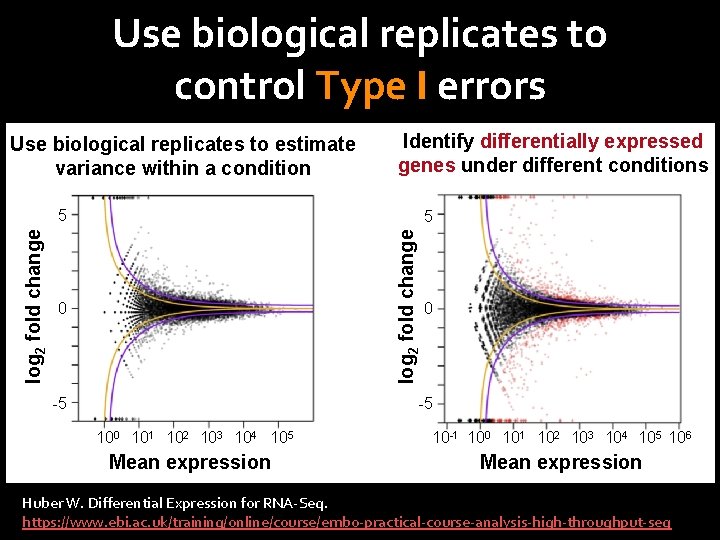
Use biological replicates to control Type I errors Use biological replicates to estimate variance within a condition Identify differentially expressed genes under different conditions 5 log 2 fold change 5 0 -5 100 101 102 103 104 105 Mean expression 10 -1 100 101 102 103 104 105 106 Mean expression Huber W. Differential Expression for RNA-Seq. https: //www. ebi. ac. uk/training/online/course/embo-practical-course-analysis-high-throughput-seq
DEMO: Differential expression analysis using DESeq 2
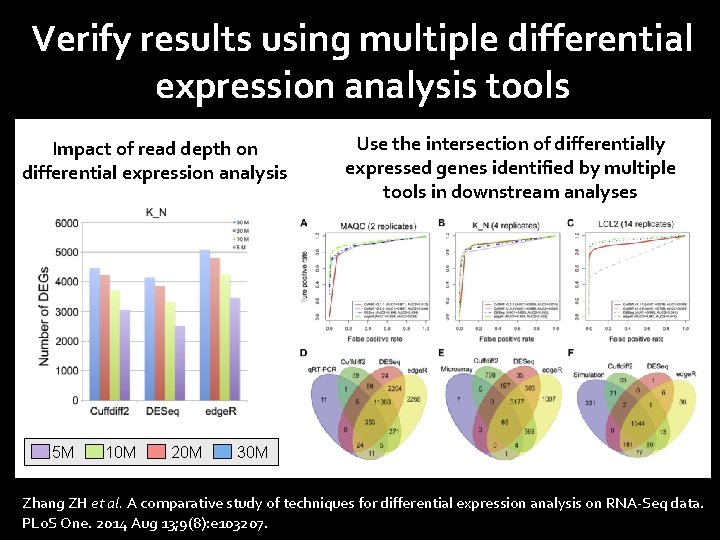
Verify results using multiple differential expression analysis tools Impact of read depth on differential expression analysis 5 M 10 M 20 M Use the intersection of differentially expressed genes identified by multiple tools in downstream analyses 30 M Zhang ZH et al. A comparative study of techniques for differential expression analysis on RNA-Seq data. PLo. S One. 2014 Aug 13; 9(8): e 103207.

Tools for analyzing differentially expressed genes Gene Ontology (GO) terms enrichment: top. GO (https: //bioconductor. org/packages/release/bioc/html/top. GO. html) go. STAG (https: //bioconductor. org/packages/release/bioc/html/go. STAG. html) DAVID (https: //david. ncifcrf. gov/) Pathway analysis: GAGE (http: //bioconductor. org/packages/release/bioc/html/gage. html) Reactome (http: //www. reactome. org/) Sample walkthrough: From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edge. R quasi-likelihood pipeline https: //www. bioconductor. org/help/workflows/Rna. Seq. Gene. Edge. RQL/

Additional resources Analysis of RNA-Seq data: gene-level exploratory analysis and differential expression http: //www-huber. embl. de/users/klaus/Teaching/DESeq 2 Predoc 2014. html Informatics for RNA-Seq: A web resource for analysis on the cloud https: //github. com/griffithlab/rnaseq_tutorial/wiki UC Davis Bioinformatics Core training course http: //bioinformatics. ucdavis. edu/training/documentation / So you want to do a: RNA-Seq experiment, Differential Gene Expression Analysis https: //github. com/msettles/Workshop_RNAseq

Summary Statistical power of RNA-Seq depends on the number of biological replicates and sequencing depth Assess quality of RNA-Seq reads with Fast. QC Assess RNA-Seq alignments with Picard tools Perform transcriptome assembly with String. Tie Perform differential expression analyses with DESeq 2

Questions? https: //flic. kr/p/bhy. T 8 B
- Slides: 55