DiamondBlackfan Anemia Gene Discovery Hanna T Gazda M

Diamond-Blackfan Anemia Gene Discovery Hanna T. Gazda, M. D. , Ph. D. Boston Children’s Hospital Harvard Medical School Boston, MA

Boston Children’s Hospital Boston, MA

Genetic DBA projects DBA gene discovery n Modifier genes n

DBA gene discovery project Boston Children’s Hospital (Genetics)– Hanna Gazda, Daniel Yuan, Shideh Kazerounian, Lindsay Swanson n Broad Institute, Cambridge, MA – Vijay Sankaran, Eric Lander n

Objectives of the presentation Ribosomal protein genes mutated in DBA n GATA 1 mutated in DBA n What does it mean for DBA families? n

Ribosomal components 60 S 5 S 5. 8 S r. RNA 28 S 47 RPL 33 RPS 19 18 S r. RNA RPS 24 40 S

Hypothesis – Other ribosomal protein (RP) gene mutations may also cause DBA Aim of the Study – To screen remaining 78 ribosomal protein genes for mutations in DBA patients without known RPS 19 and RPS 24 mutations

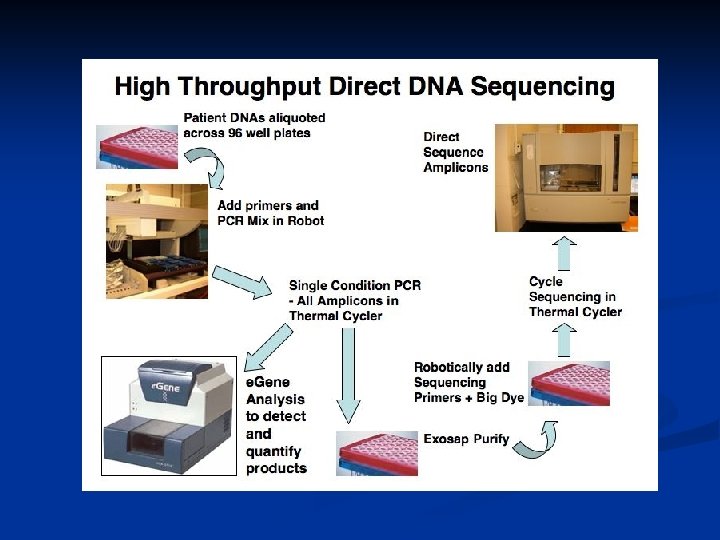
Methods – Screened DNA samples from DBA patients by direct sequencing of exons and intron/exon boundaries using DNA from 96 DBA patients Sequence change identified 1) 2) 3) 4) Sequencing of DNA from an additional 96 patients Search the NCBI and Hap. Map SNP databases Sequencing of DNA from 150 -200 control samples Sequencing of DNA from family members
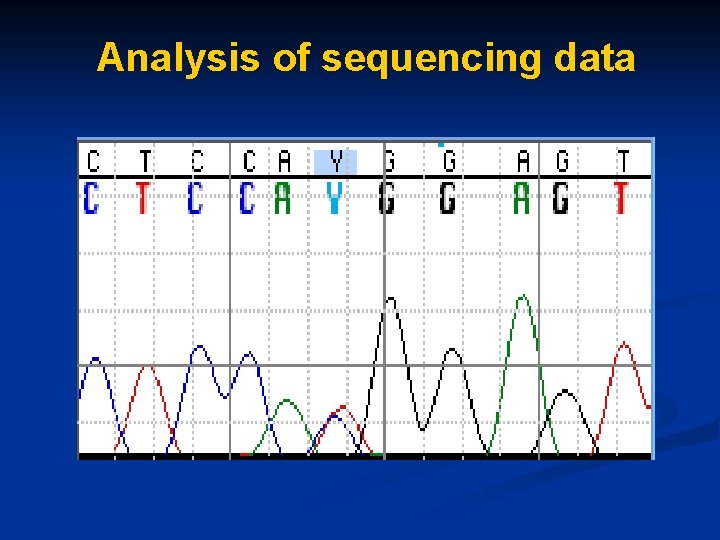
Analysis of sequencing data
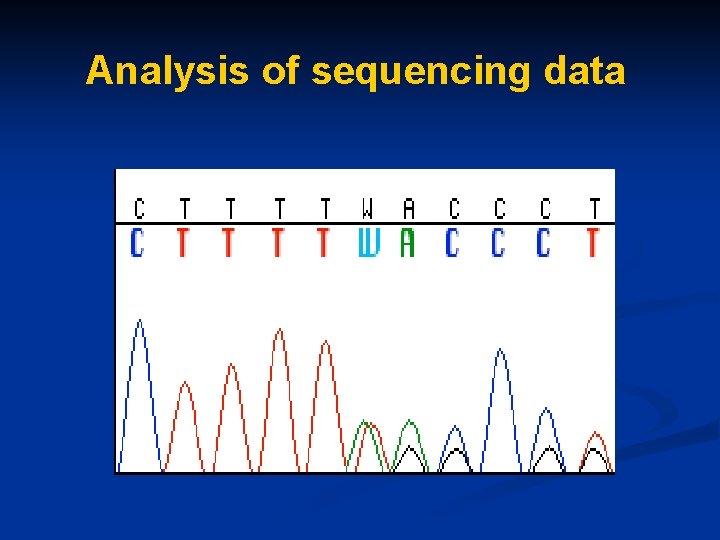
Analysis of sequencing data

Summary of ribosomal protein genes mutated in DBA Draptchinskaia et al 1999 Gazda et al 2006 Cmejla et al 2007 Farrar et al 2008 Gazda et al 2008 Doherty et al 2010 Gazda et al 2012

Large RP gene deletions in DBA n Dr. Bodine’s group (NIH); 9/51 n n Dr. Hamaguchi’s group (Japan); 7/27 n n RPS 19, RPS 17, RPL 5 and RPL 35 A Dr. Dianzani-Ramenghi’s group (Italy); 14/72 n n RPS 19, RPS 17, RPS 26 and RPL 35 A RPS 19, RPS 17, RPS 26, RPL 5, RPL 11 and RPL 35 A Our own data (BCH); 6/87 n RPS 19, RPS 17, RPS 24, RPS 26 and RPL 15

Ribosomal protein genes and DBA n Ribosomal protein gene mutations and large deletions are known in about 60 -65% of DBA patients n ~35 -40% of patients do not have known pathogenic mutation(s)

Importance of genetic screening in DBA To confirm the clinical diagnosis of DBA n For stem cell transplantation n For reproductive choices (pre-implantation genetic diagnosis) n For future gene therapy n
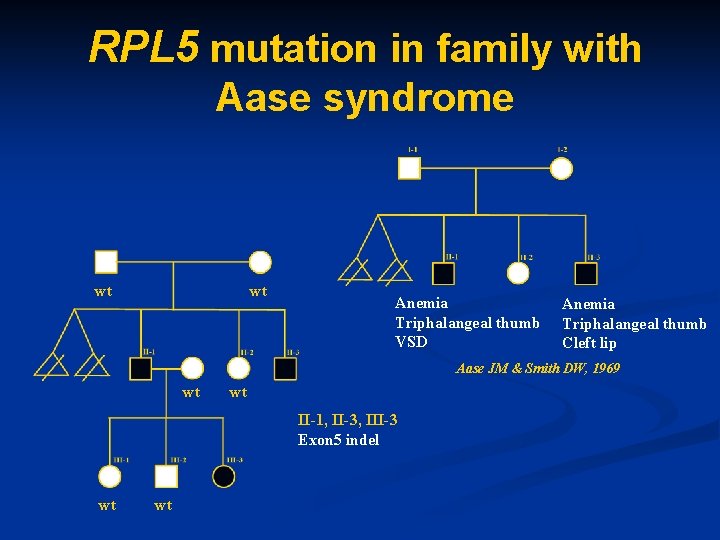
RPL 5 mutation in family with Aase syndrome wt wt Anemia Triphalangeal thumb VSD Anemia Triphalangeal thumb Cleft lip Aase JM & Smith DW, 1969 wt wt II-1, II-3, III-3 Exon 5 indel wt wt
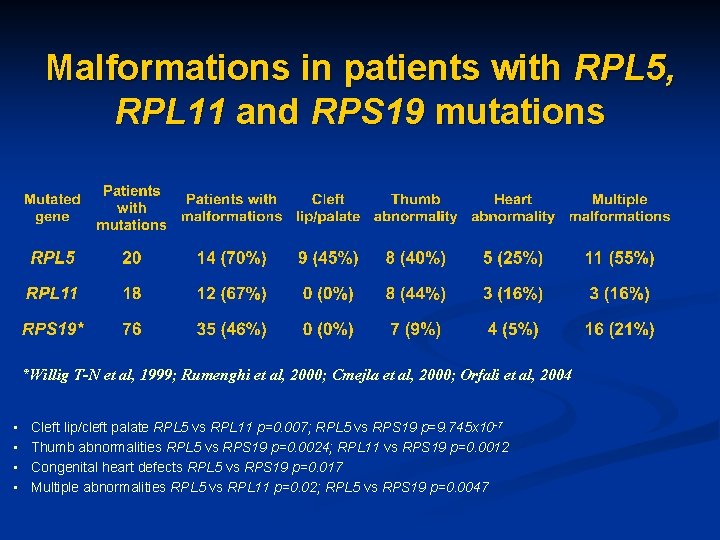
Malformations in patients with RPL 5, RPL 11 and RPS 19 mutations *Willig T-N et al, 1999; Rumenghi et al, 2000; Cmejla et al, 2000; Orfali et al, 2004 • • Cleft lip/cleft palate RPL 5 vs RPL 11 p=0. 007; RPL 5 vs RPS 19 p=9. 745 x 10 -7 Thumb abnormalities RPL 5 vs RPS 19 p=0. 0024; RPL 11 vs RPS 19 p=0. 0012 Congenital heart defects RPL 5 vs RPS 19 p=0. 017 Multiple abnormalities RPL 5 vs RPL 11 p=0. 02; RPL 5 vs RPS 19 p=0. 0047

Mutations of ribosomal protein genes in DBA n n n Mutations of RPS 19, RPL 5, RPL 11, RPS 10, RPS 26 and RPL 35 A are common causes of Diamond-Blackfan anemia, while RPS 24, RPS 7, RPS 17, and RPL 26 are sporadically mutated in DBA. All mutations are heterozygous and present in ~55% of patients. Mutations in RPL 5 are associated with multiple physical abnormalities including triphalangeal thumbs and cleft lip/cleft palate, while RPL 11 mutations are predominantly associated with isolated abnormal thumbs Large deletions are present in ~ 5 -10% of patients. RPL 15 is a novel gene associated with DBA.

Mutations of ribosomal protein genes in DBA n n Majority are nonsense, splice site or frameshift (insertions, deletions) Heterozygous (present on one copy of the gene) and indicate autosomal dominant inheritance

Karyogram of a human female
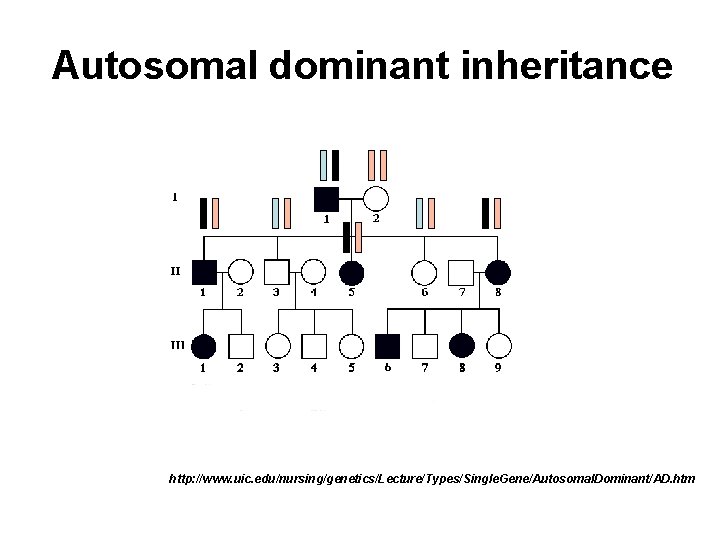
Autosomal dominant inheritance http: //www. uic. edu/nursing/genetics/Lecture/Types/Single. Gene/Autosomal. Dominant/AD. htm
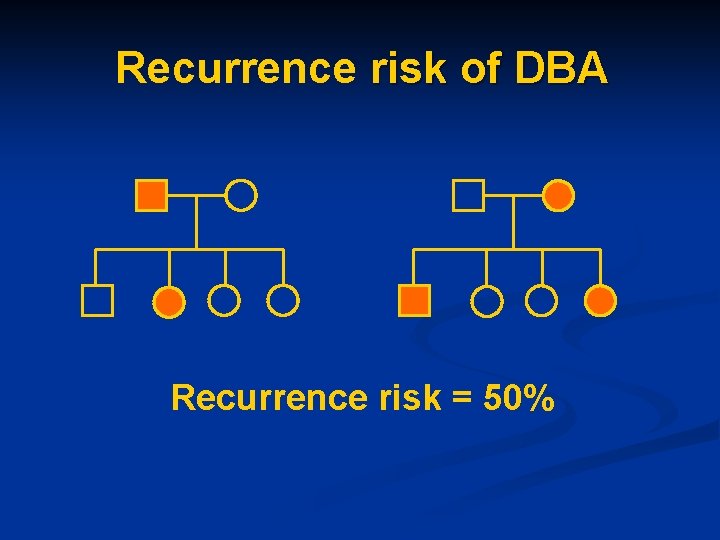
Recurrence risk of DBA Recurrence risk = 50%
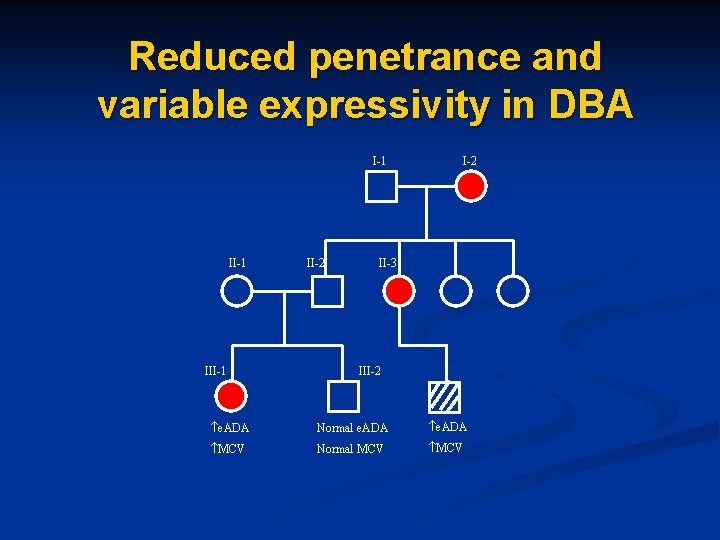
Reduced penetrance and variable expressivity in DBA I-1 III-1 II-2 M I-2 II-3 III-2 e. ADA Normal e. ADA MCV Normal MCV
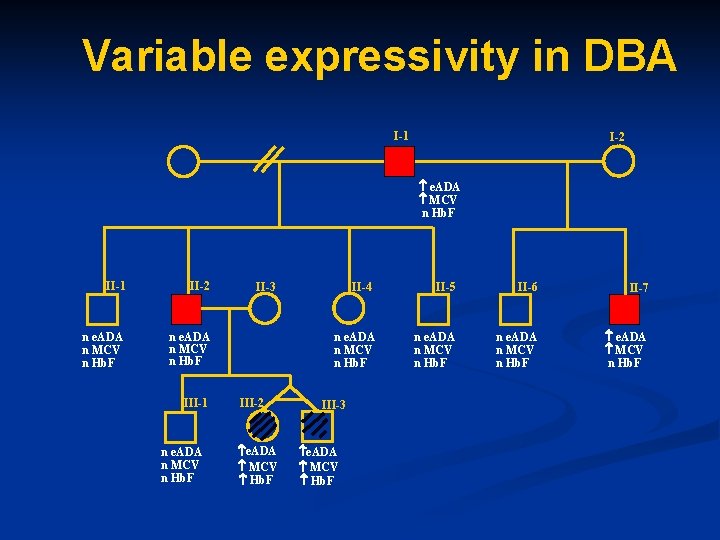
Variable expressivity in DBA I-1 I-2 e. ADA MCV n Hb. F II-1 II-2 n e. ADA n MCV n Hb. F III-1 n e. ADA n MCV n Hb. F II-3 II-4 n e. ADA n MCV n Hb. F III-2 e. ADA MCV Hb. F III-3 e. ADA MCV Hb. F II-5 II-6 II-7 n e. ADA n MCV n Hb. F e. ADA MCV n Hb. F

Germline mutations in DBA N e. ADA N MCV M M e. ADA MCV
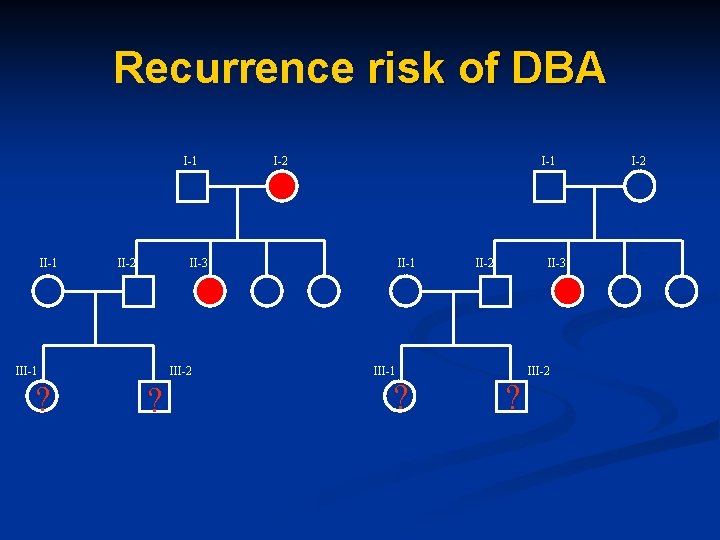
Recurrence risk of DBA I-1 II-2 M III-1 ? I-1 II-3 III-2 ? I-2 II-1 III-1 ? II-2 M ? II-3 III-2

Recurrence risk of DBA Recurrence risk is slightly higher than in general population

Ribosomal protein genes and DBA n Ribosomal protein gene mutations and large deletions are known in about 60 -65% of DBA patients n ~35 -40% of patients do not have known pathogenic mutation(s)

Next step in DBA gene discoveries n Entire exome sequencing- (all exons) all coding regions of ~25, 000 genes

New patients enrolled into our study 11 ribosomal protein gene screening n GATA 1 gene screening n Screening of the new genes by exome sequencing n

Study participation in DBA gene discovery and modifier genes Consent form and Questionnaire – Lindsay Swanson, genetic counselor; ph. 617 -919 -2169; lindsay. swanson@childrens. harvard. edu n Blood draw at local doctor’s office n Blood sample sent to Boston Children’s Hospital n No charge to participate n

Acknowledgements Alan H. Beggs Mee Rie Sheen Natasha Darras Leana Doherty Mike Landowski Chris Buros Roxy Ghazvinian Adrianna Vlachos Jeffrey M. Lipton Eva Atsidaftos ] Genetics/Genomics Children’s Hospital Harvard Medical School Boston, MA, USA Vijay Sankaran Eric Lander Bertil Glader ] DBA Registry Feinstein Institute for Medical Research, Manhasset, NY Colin A. Sieff s Hospital Boston, MA, USA University of London, Sarah E. Ball St. George's London, UK ] ] Stanford University School of Medicine Stanford, CA Charlotte Niemeyer Joerg Meerpohl Children’ Edyta Niewiadomska Michal Matysiak ] Peter E. Newburger University of Massachusetts Medical School, Worcester, MA, USA University Medical School of Warsaw, Poland Broad Institute, Cambridge, MA ] University of Freiburg, Germany DBA Foundation DMA Foundation We thank the physicians, DBA patients and their family members for participating in the study!
- Slides: 32