DEVELOPMENT OF A PCR BASED TEST TO DIFFERENTIATE

DEVELOPMENT OF A PCR BASED TEST TO DIFFERENTIATE PADDLEFISH AND STURGEON EGGS Scott Cooper, Jill Mader, Sue Kittleson, Valerie Hyde, Marc Rott Biology/Microbiology Department, University of Wisconsin-La Crosse, WI 56401

Sample Preparation • Eggs kept on ice until freezing at -80 o. C • Single egg mashed with sterile glass rod in 200 ul buffer (20 m. M Tris, 20 m. M KCl, 0. 5% Tween 20, 5% Chelex, 0. 2 mg/ml proteinase K) • Samples incubated 1. 5 hours at 55 o. C • Samples incubated 8 minutes at 95 o. C • Samples centrifuged at 14, 000 rpm for 10 minutes • An aliquot was removed for PCR amplification (0. 01 -1 ul)
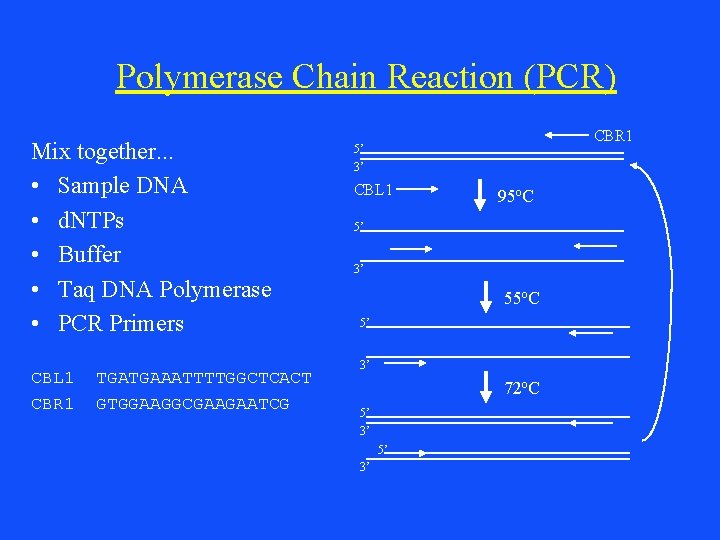
Polymerase Chain Reaction (PCR) Mix together. . . • Sample DNA • d. NTPs • Buffer • Taq DNA Polymerase • PCR Primers CBL 1 CBR 1 TGATGAAATTTTGGCTCACT GTGGAAGGCGAAGAATCG CBR 1 5’ 3’ CBL 1 95 o. C 5’ 3’ 55 o. C 5’ 3’ 72 o. C 5’ 3’

Cytochrome b DNA Sequences • PCR products from paddlefish, shovelnose sturgeon and lake sturgeon were cloned and sequenced. Table 1. Percent DNA Sequence Identity Shovelnose Lake Paddlefish 86. 6% 86. 0% Lake 90. 8%

Restriction Enzyme Digestion Mix together : • 1 ul PCR product • 8 ul buffer and water • 1 ul restriction enzyme Incubate 30 min, 37 o. C. Subject sample to agarose gel electrophoresis for 45 minutes. Stain gel with ethidium bromide and photograph.

Restriction Mapping Restriction enzymes cut DNA at specific sites. A single change in the sequence of that site will prevent cleavage. Example: Hinc. II cuts GTTAAC but not GTAAAC in paddlefish cyt b DNA in sturgeon cyt b DNA Hinc. II Nar. I Hinc. II PA SN LK
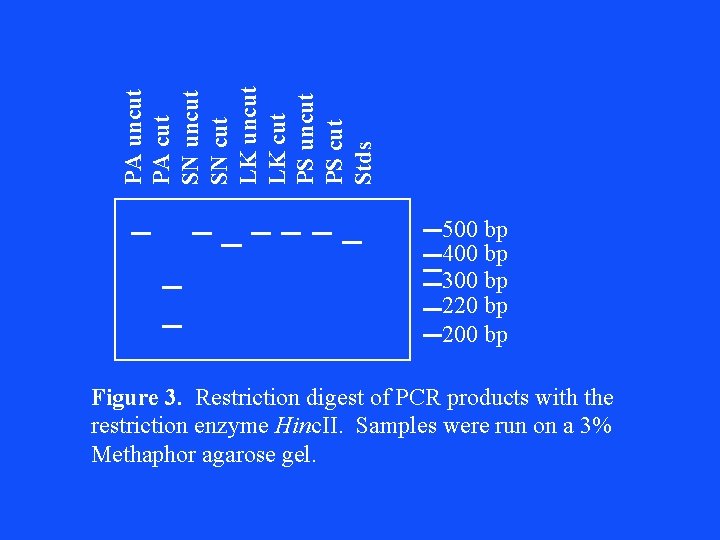
PA uncut PA cut SN uncut SN cut LK uncut LK cut PS uncut PS cut Stds 500 bp 400 bp 300 bp 220 bp 200 bp Figure 3. Restriction digest of PCR products with the restriction enzyme Hinc. II. Samples were run on a 3% Methaphor agarose gel.

PA uncut PA cut SN uncut SN cut LK uncut LK cut PS uncut PS cut Stds 500 bp 400 bp 300 bp 220 bp 200 bp Figure 4. Restriction digest of PCR products with the restriction enzyme Nar. I. Samples were run on a 1% LE agarose gel.

Conclusions • Enough DNA can be isolated from a single fish egg to allow for sucessful PCR amplification. • DNA sequence analysis demosnstrates unique genetic markers in paddlefish, and in shovelnose, pallid and lake sturgeon. • This assay will allow the species identification of a single egg within one day. • To date this assay has been tested on more than thirty fish, from all over the country, with no observed discrepancies in the assay.
- Slides: 9