Development of a Novel AntibodyBased Assay for Simultaneous
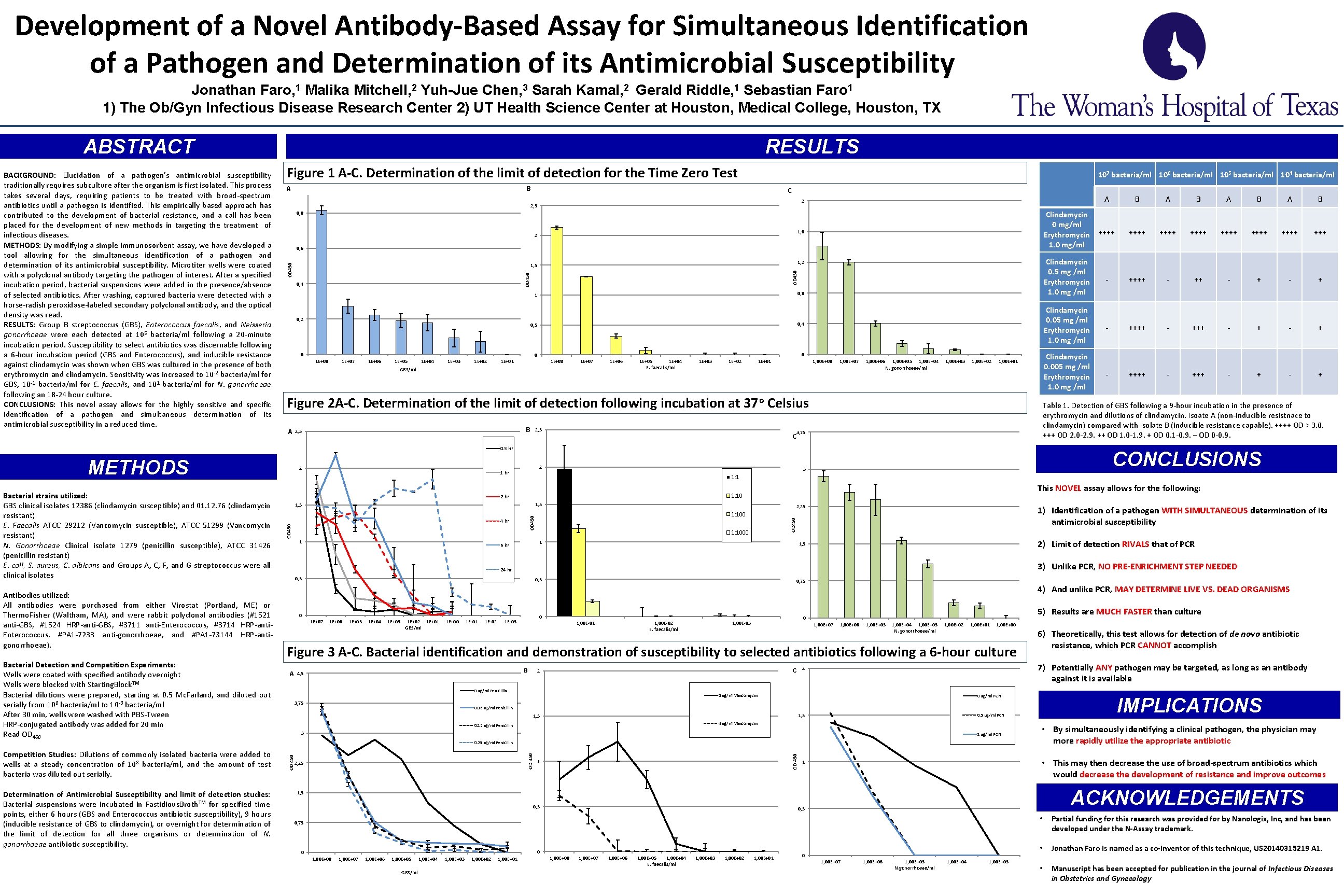
Development of a Novel Antibody-Based Assay for Simultaneous Identification of a Pathogen and Determination of its Antimicrobial Susceptibility Jonathan Faro, 1 Malika Mitchell, 2 Yuh-Jue Chen, 3 Sarah Kamal, 2 Gerald Riddle, 1 Sebastian Faro 1 1) The Ob/Gyn Infectious Disease Research Center 2) UT Health Science Center at Houston, Medical College, Houston, TX ABSTRACT Figure 1 A-C. Determination of the limit of detection for the Time Zero Test A 107 bacteria/ml 106 bacteria/ml 105 bacteria/ml 104 bacteria/ml B C 0, 8 1, 6 2 0, 6 1, 2 OD 450 1, 5 0, 4 0, 8 1 0, 2 0, 5 0, 4 0 0 0 1 E+08 1 E+07 1 E+06 1 E+05 1 E+04 1 E+03 1 E+02 1 E+01 1 E+08 1 E+07 1 E+06 GBS/ml 1 E+05 1 E+04 E. faecalis/ml 1 E+03 1 E+02 1, 00 E+08 1 E+01 1, 00 E+07 1, 00 E+06 1, 00 E+05 1, 00 E+04 N. gonorrhoeae/ml 1, 00 E+03 1, 00 E+02 1, 00 E+01 Figure 2 A-C. Determination of the limit of detection following incubation at 37 o Celsius B 2, 5 A 2, 5 3, 75 C Competition Studies: Dilutions of commonly isolated bacteria were added to wells at a steady concentration of 108 bacteria/ml, and the amount of test bacteria was diluted out serially. Determination of Antimicrobial Susceptibility and limit of detection studies: Bacterial suspensions were incubated in Fastidious. Broth. TM for specified timepoints, either 6 hours (GBS and Enterococcus antibiotic susceptibility), 9 hours (inducible resistance of GBS to clindamycin), or overnight for determination of the limit of detection for all three organisms or determination of N. gonorrhoeae antibiotic susceptibility. OD 450 4 hr OD 450 2, 25 1: 1000 1 6 hr 0, 5 0 1 E+03 1 E+02 1 E+01 GBS/ml 1 E+00 1 E-01 1 E-02 0, 75 0 1 E-03 1, 00 E-01 B 1, 00 E-02 E. faecalis/ml 1, 00 E+07 1, 00 E+06 1, 00 E+05 1, 00 E+04 1, 00 E+03 N. gonorrhoeae/ml 1, 00 E+02 1, 00 E+01 1, 00 E+00 0 ug/ml Vancomycin 0 ug/ml PCN 1, 5 3 1 ug/ml PCN 0. 25 ug/ml Penicillin 1 0, 5 GBS/ml 1, 00 E+04 1, 00 E+03 1, 00 E+02 1, 00 E+01 ++++ Clindamycin 0. 5 mg /ml Erythromycin 1. 0 mg /ml - ++++ - + - + Clindamycin 0. 05 mg /ml Erythromycin 1. 0 mg /ml - ++++ - + - + Clindamycin 0. 005 mg /ml Erythromycin 1. 0 mg /ml - ++++ - + - + ACKNOWLEDGEMENTS 0, 5 0 1, 00 E+05 ++++ • This may then decrease the use of broad-spectrum antibiotics which would decrease the development of resistance and improve outcomes 1 0, 75 1, 00 E+06 ++++ • By simultaneously identifying a clinical pathogen, the physician may more rapidly utilize the appropriate antibiotic 1, 5 1, 00 E+07 ++++ IMPLICATIONS 0. 5 ug/ml PCN 4 ug/ml Vancomycin 0. 12 ug/ml Penicillin 1, 00 E+08 ++++ 7) Potentially ANY pathogen may be targeted, as long as an antibody against it is available C 2 2 1, 5 0 ++++ 6) Theoretically, this test allows for detection of de novo antibiotic resistance, which PCR CANNOT accomplish 0. 06 ug/ml Penicillin 2, 25 ++++ 5) Results are MUCH FASTER than culture 0 1, 00 E-03 0 ug/ml Penicillin 3, 75 Clindamycin 0 mg/ml Erythromycin 1. 0 mg/ml 4) And unlike PCR, MAY DETERMINE LIVE VS. DEAD ORGANISMS Figure 3 A-C. Bacterial identification and demonstration of susceptibility to selected antibiotics following a 6 -hour culture A 4, 5 B 3) Unlike PCR, NO PRE-ENRICHMENT STEP NEEDED 0, 5 1 E+04 A 2) Limit of detection RIVALS that of PCR 24 hr 1 E+05 B 1) Identification of a pathogen WITH SIMULTANEOUS determination of its antimicrobial susceptibility 1, 5 OD 450 Bacterial Detection and Competition Experiments: Wells were coated with specified antibody overnight Wells were blocked with Starting. Block. TM Bacterial dilutions were prepared, starting at 0. 5 Mc. Farland, and diluted out serially from 108 bacteria/ml to 10 -3 bacteria/ml After 30 min, wells were washed with PBS-Tween HRP-conjugated antibody was added for 20 min Read OD 450 1, 5 1 E+06 A This NOVEL assay allows for the following: 1: 10 1, 5 1 E+07 B CONCLUSIONS 3 2 hr 1 A 1: 1 OD 450 Antibodies utilized: All antibodies were purchased from either Virostat (Portland, ME) or Thermo. Fisher (Waltham, MA), and were rabbit polyclonal antibodies (#1521 anti-GBS, #1524 HRP-anti-GBS, #3711 anti-Enterococcus, #3714 HRP-anti. Enterococcus, #PA 1 -7233 anti-gonorrhoeae, and #PA 1 -73144 HRP-antigonorrhoeae). 1 hr OD 450 Bacterial strains utilized: GBS clinical isolates 12386 (clindamycin susceptible) and 01. 12. 76 (clindamycin resistant) E. Faecalis ATCC 29212 (Vancomycin susceptible), ATCC 51299 (Vancomycin resistant) N. Gonorrhoeae Clinical isolate 1279 (penicillin susceptible), ATCC 31426 (penicillin resistant) E. coli, S. aureus, C. albicans and Groups A, C, F, and G streptococcus were all clinical isolates 2 OD 450 METHODS B Table 1. Detection of GBS following a 9 -hour incubation in the presence of erythromycin and dilutions of clindamycin. Isoate A (non-inducible resistnace to clindamycin) compared with Isolate B (inducible resistance capable). ++++ OD > 3. 0. +++ OD 2. 0 -2. 9. ++ OD 1. 0 -1. 9. + OD 0. 1 -0. 9. – OD 0 -0. 9. 0. 5 hr 2 A 2 2, 5 OD 450 BACKGROUND: Elucidation of a pathogen’s antimicrobial susceptibility traditionally requires subculture after the organism is first isolated. This process takes several days, requiring patients to be treated with broad-spectrum antibiotics until a pathogen is identified. This empirically based approach has contributed to the development of bacterial resistance, and a call has been placed for the development of new methods in targeting the treatment of infectious diseases. METHODS: By modifying a simple immunosorbent assay, we have developed a tool allowing for the simultaneous identification of a pathogen and determination of its antimicrobial susceptibility. Microtiter wells were coated with a polyclonal antibody targeting the pathogen of interest. After a specified incubation period, bacterial suspensions were added in the presence/absence of selected antibiotics. After washing, captured bacteria were detected with a horse-radish peroxidase-labeled secondary polyclonal antibody, and the optical density was read. RESULTS: Group B streptococcus (GBS), Enterococcus faecalis, and Neisseria gonorrhoeae were each detected at 105 bacteria/ml following a 20 -minute incubation period. Susceptibility to select antibiotics was discernable following a 6 -hour incubation period (GBS and Enterococcus), and inducible resistance against clindamycin was shown when GBS was cultured in the presence of both erythromycin and clindamycin. Sensitivity was increased to 10 -2 bacteria/ml for GBS, 10 -1 bacteria/ml for E. faecalis, and 101 bacteria/ml for N. gonorrhoeae following an 18 -24 hour culture. CONCLUSIONS: This novel assay allows for the highly sensitive and specific identification of a pathogen and simultaneous determination of its antimicrobial susceptibility in a reduced time. RESULTS 1, 00 E+08 1, 00 E+07 1, 00 E+06 1, 00 E+05 1, 00 E+04 E. faecalis/ml 1, 00 E+03 1, 00 E+02 1, 00 E+01 0 1, 00 E+07 1, 00 E+06 1, 00 E+05 N. gonorrhoeae/ml 1, 00 E+04 1, 00 E+03 • Partial funding for this research was provided for by Nanologix, Inc, and has been developed under the N-Assay trademark. • Jonathan Faro is named as a co-inventor of this technique, US 20140315219 A 1. • Manuscript has been accepted for publication in the journal of Infectious Diseases in Obstetrics and Gynecology
- Slides: 1