Detoxification of Organophosphates by INPsurface display systems Contents

Detoxification of Organophosphates by INP-surface display systems

Contents : 1. 2. 3. 4. 5. 6. 7. Organophosphates , Organophosphorus hydrolase and Methyl Parathion hydrolase Microbial cell surface display systems Ice nucleation protein INP based cell surface display system Various approaches employed for detoxification of organophosphates Conclusion References

1. Organophosphates (OPs) , Organophosphorus hydrolase (OPH) and Methyl Parathion hydrolase(MPH) : It is one of the most popular group of pesticides and insecticides Ø Share structural similarities to chemical warfare agents like sarin, soman, etc Ø Neurotoxic in nature Ø OPH and MPH capable of hydrolysing broad range of OPs. Ø In whole cells, OPH resides inside the cell and thus outermembrane acts as permeability barrier Ø Thus, surface display OPH on the outermembrane so that it can interact with the OPs Ø
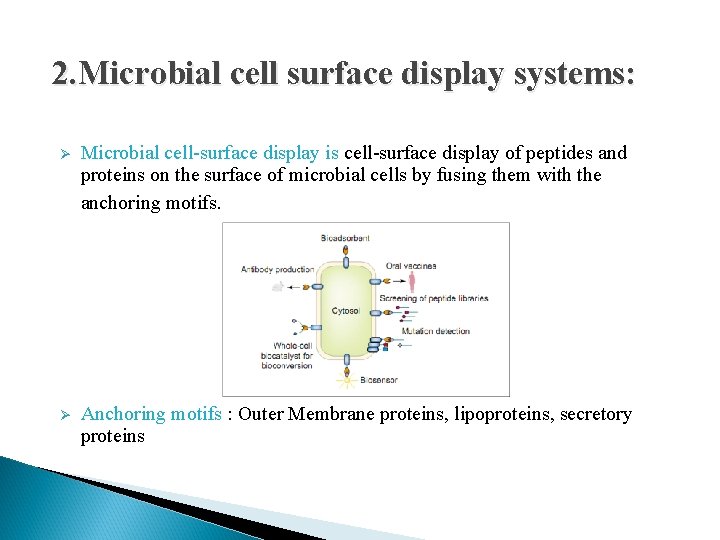
2. Microbial cell surface display systems: Ø Microbial cell-surface display is cell-surface display of peptides and proteins on the surface of microbial cells by fusing them with the anchoring motifs. Ø Anchoring motifs : Outer Membrane proteins, lipoproteins, secretory proteins

3. Ice Nucleation Protein : Ø It causes bacterial ice nucleation and frost injury in plants eg. Pseudomonas syringae and some other Gram negative strains like– Pseudomonas fluorescens Erwinia ananas Erwinia uredovora Xanthomonas campestris Ø It is 1034 – 1567 residues long Contain 3 unique domains : 1. non-repetitive N-terminal domain 2. highly repetitive central domain 3. non-repetitive C-terminal domain

4. INP based cell surface display system because : Ø Stable expression Ø Resistant to protease activity Ø Modulatable length of internal repeating units Ø Attachment of eukaryotic cell surface proteins Ø Outer Membrane translocation Ø Can be expressed with high molecular weight proteins Ø Compatible with foreign sequences
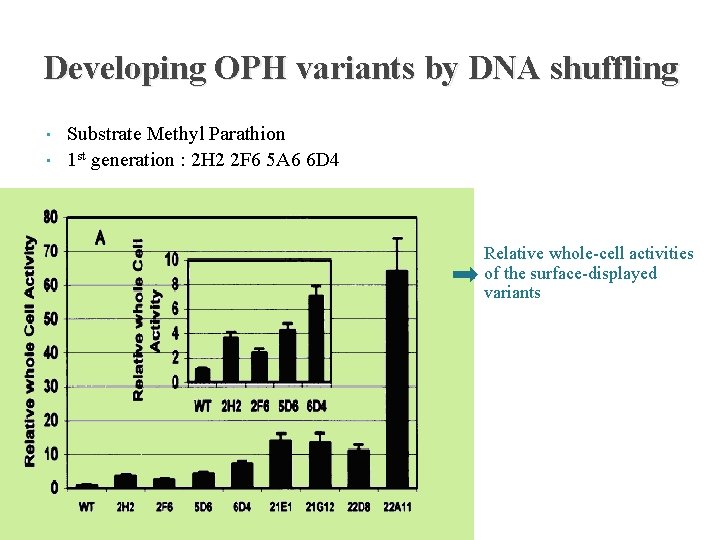
Developing OPH variants by DNA shuffling • • Substrate Methyl Parathion 1 st generation : 2 H 2 2 F 6 5 A 6 6 D 4 Relative whole-cell activities of the surface-displayed variants purified variants
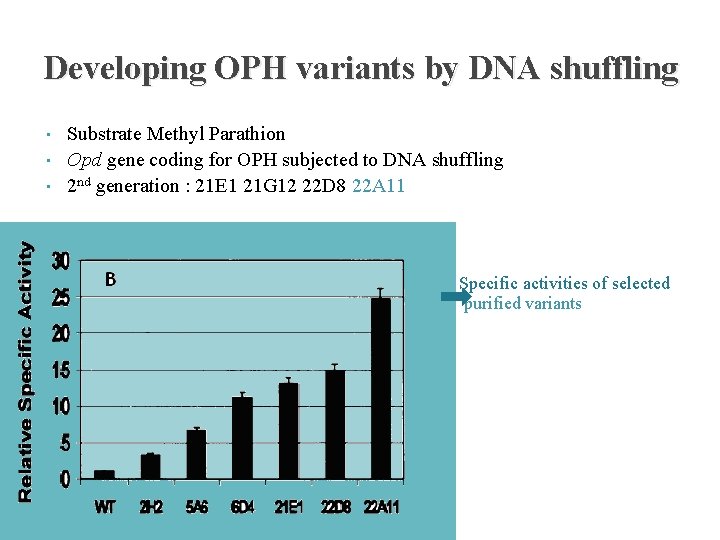
Developing OPH variants by DNA shuffling • • • Substrate Methyl Parathion Opd gene coding for OPH subjected to DNA shuffling 2 nd generation : 21 E 1 21 G 12 22 D 8 22 A 11 Specific activities of selected purified variants Relative whole-cell activities of
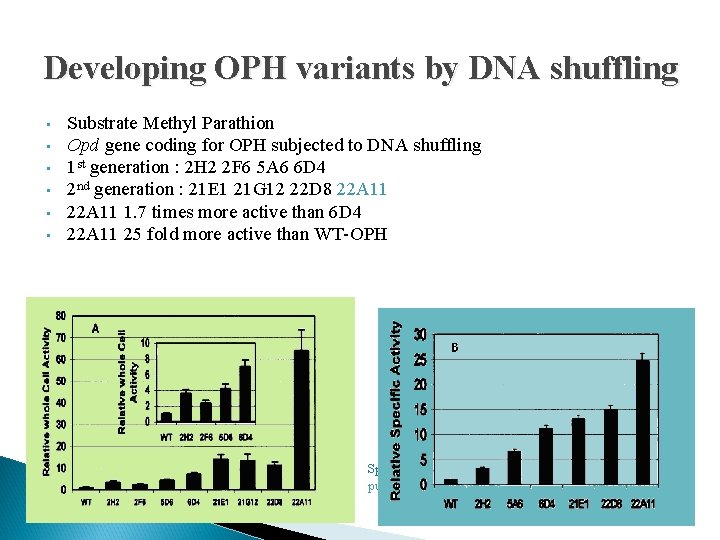
Developing OPH variants by DNA shuffling • • • Substrate Methyl Parathion Opd gene coding for OPH subjected to DNA shuffling 1 st generation : 2 H 2 2 F 6 5 A 6 6 D 4 2 nd generation : 21 E 1 21 G 12 22 D 8 22 A 11 1. 7 times more active than 6 D 4 22 A 11 25 fold more active than WT-OPH Relative whole-cell activities of the surface-displayed variants. Specific activities of selected purified variants
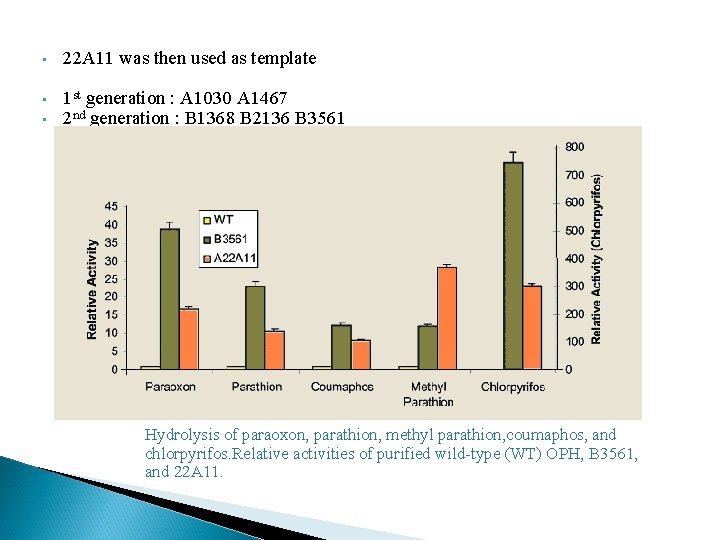
• 22 A 11 was then used as template • • 1 st generation : A 1030 A 1467 2 nd generation : B 1368 B 2136 B 3561 Hydrolysis of paraoxon, parathion, methyl parathion, coumaphos, and chlorpyrifos. Relative activities of purified wild-type (WT) OPH, B 3561, and 22 A 11.
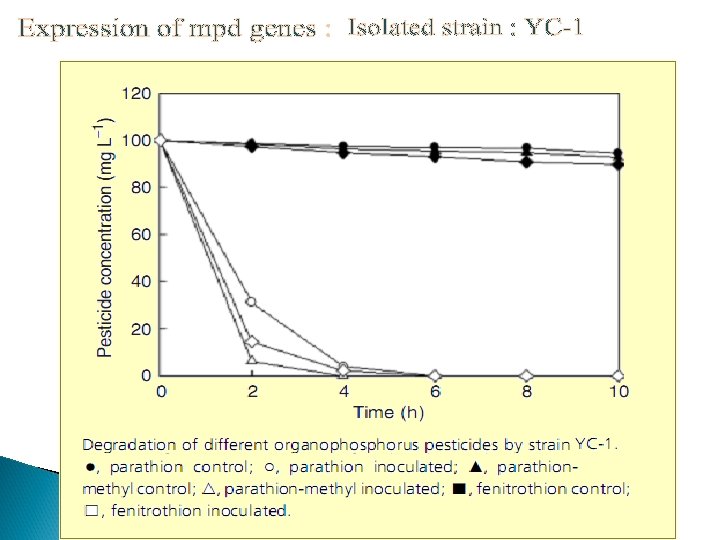
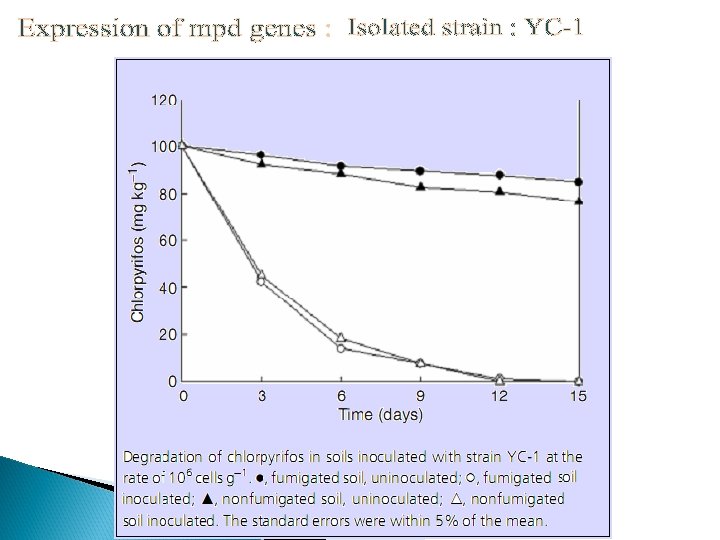
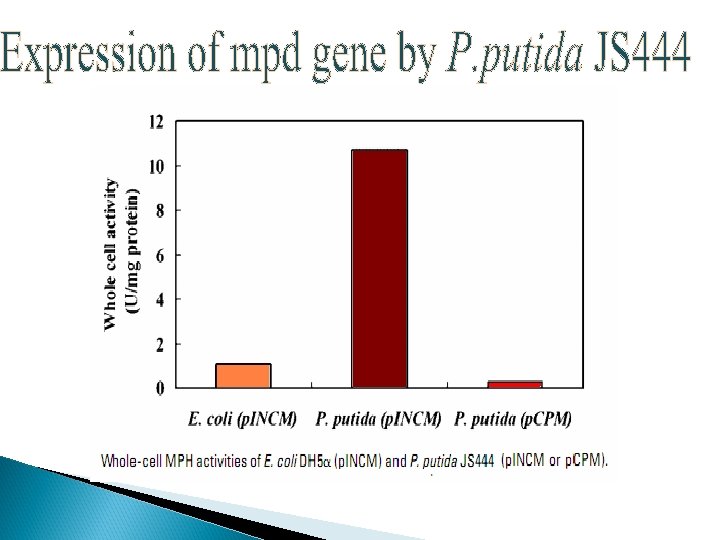
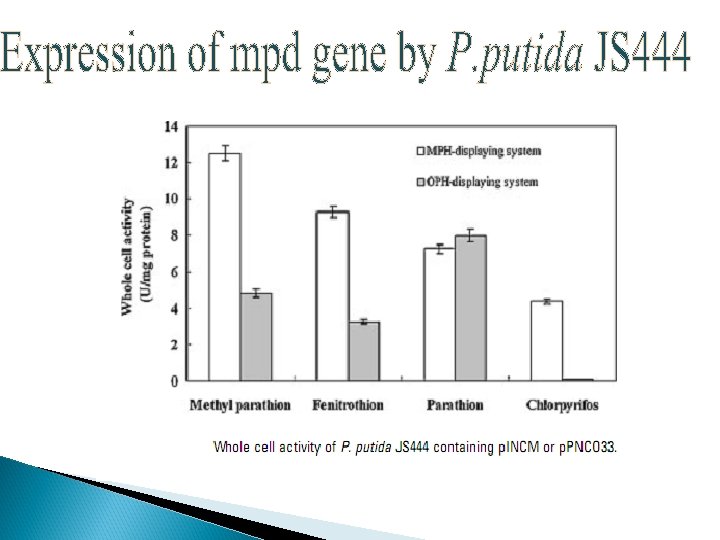
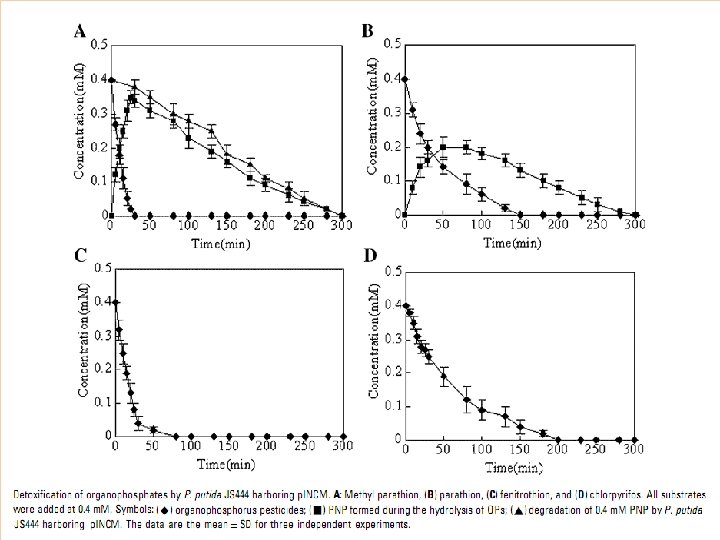

Co-immobilization Ø Ø Ø Immobilization was done on nonwoven cellulosic fiber Co-expression of OPH and CBD on surface but different binding motifs Recombinant strain : XL 1 -Blue/p. UCBD/p. PNCO 33
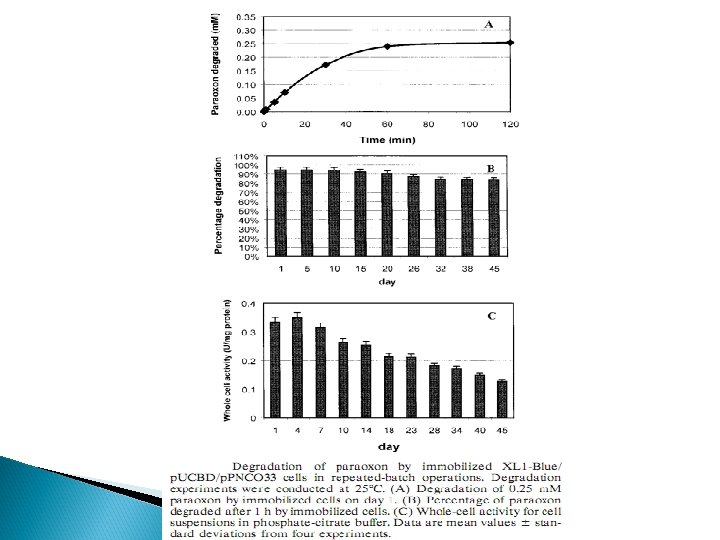

Co-translocation Express OPH and MPH both. OP hydrolase activity : Table : OP hydrolase activities from different subcellular fractions of E. coli. XL 1 Blue/p. PNCO 33/p. UTM 18 and E. coli. XL 1 -Blue/p. CPO/p. UM 18
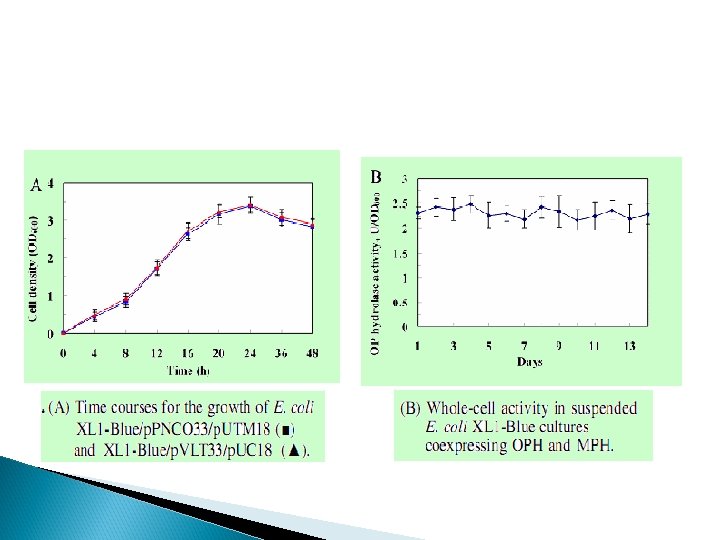
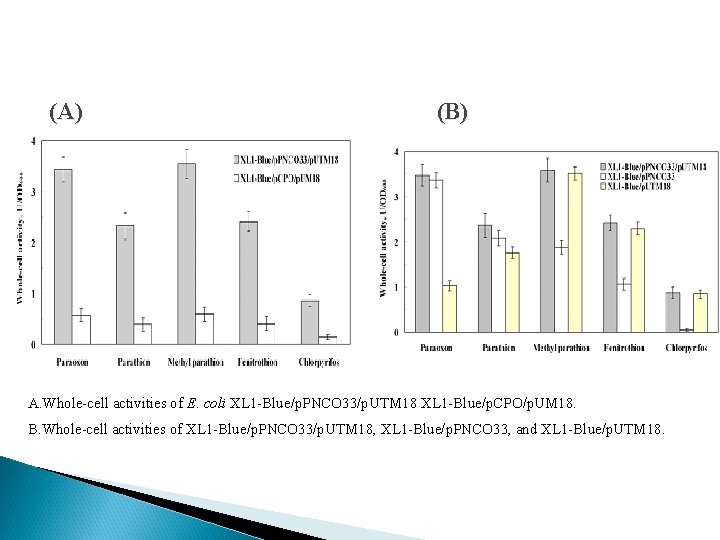
(A) (B) A. Whole-cell activities of E. coli XL 1 -Blue/p. PNCO 33/p. UTM 18 XL 1 -Blue/p. CPO/p. UM 18. B. Whole-cell activities of XL 1 -Blue/p. PNCO 33/p. UTM 18, XL 1 -Blue/p. PNCO 33, and XL 1 -Blue/p. UTM 18.

Conclusion : OPs being synthetic in nature are not degraded easily Ø Other Methods : chemical incineration , Mutation , Enzymatic degradation , whole cells as biocatalysts Ø By products can be used as sole Carbon and Phosphorus sources ( Degradation brought about by P. putida ) Ø Plant pathogen used for degradative purpose of neurotoxic agents Ø
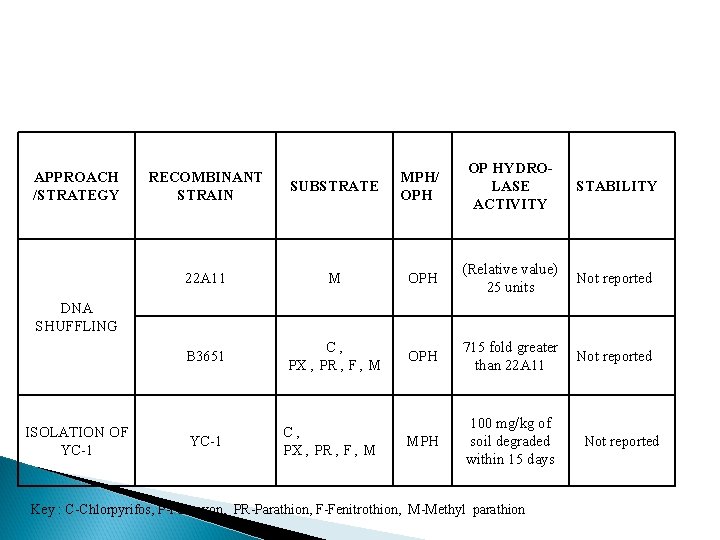
APPROACH /STRATEGY SUBSTRATE MPH/ OPH OP HYDROLASE ACTIVITY STABILITY 22 A 11 M OPH (Relative value) 25 units Not reported B 3651 C, PX , PR , F , M OPH 715 fold greater than 22 A 11 Not reported MPH 100 mg/kg of soil degraded within 15 days RECOMBINANT STRAIN DNA SHUFFLING ISOLATION OF YC-1 C, PX , PR , F , M Key : C-Chlorpyrifos, P-Paraoxon, PR-Parathion, F-Fenitrothion, M-Methyl parathion Not reported
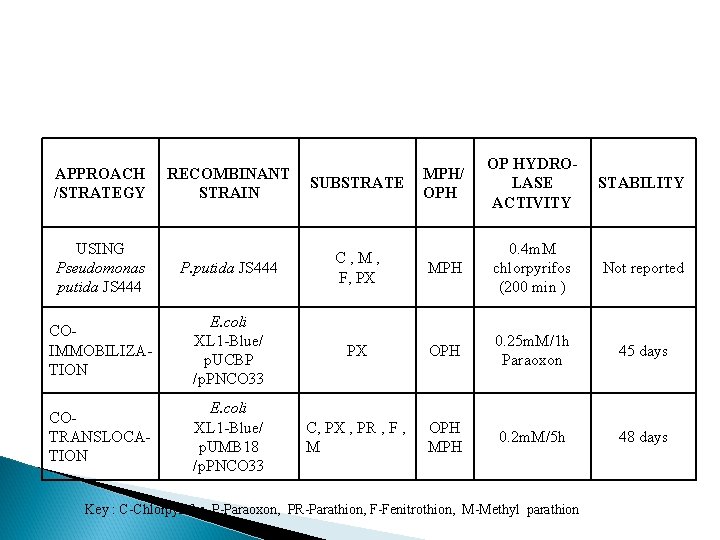
MPH/ OPH OP HYDROLASE ACTIVITY STABILITY P. putida JS 444 C, M, F, PX MPH 0. 4 m. M chlorpyrifos (200 min ) Not reported COIMMOBILIZATION E. coli XL 1 -Blue/ p. UCBP /p. PNCO 33 PX OPH 0. 25 m. M/1 h Paraoxon 45 days COTRANSLOCATION E. coli XL 1 -Blue/ p. UMB 18 /p. PNCO 33 C, PX , PR , F , M OPH MPH 0. 2 m. M/5 h 48 days APPROACH /STRATEGY RECOMBINANT STRAIN SUBSTRATE USING Pseudomonas putida JS 444 Key : C-Chlorpyrifos, P-Paraoxon, PR-Parathion, F-Fenitrothion, M-Methyl parathion

Thank You !
- Slides: 24