Determination of biomolecular structure NMR spectroscopy Xray Crystallography

Determination of bio-molecular structure

NMR spectroscopy X-ray Crystallography

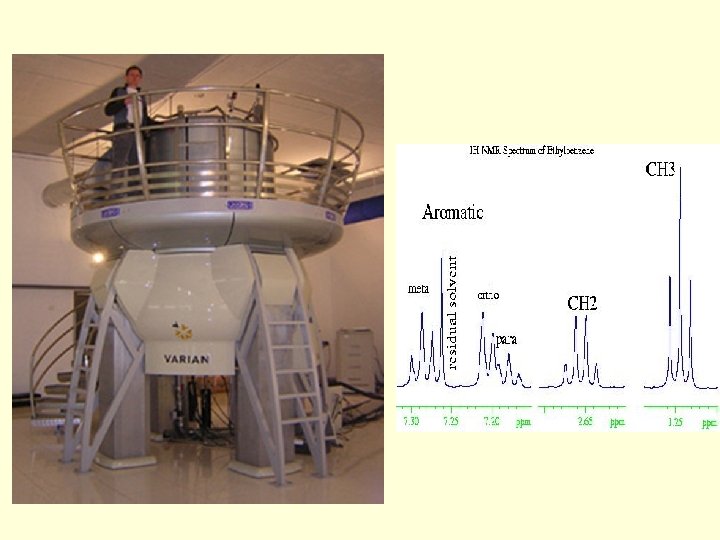

Why NMR spectroscopy is good for you especially in the biological sciences – biomolecular data at atomic resolution directly from recording a spectrum – mapping binding sites in protein-protein or generally protein ligand interaction – measurement of acid and base p. Ka values – characterization of fold and stability of biomolecules – determination of 3 D structures of proteins and nucleic acids

Cont… – characterization of biomolecular dynamics from picosecond to days – fast analysis of the impact of mutations on protein structure/dynamics – developing the methodology might get you a nobel prize: Richard Ernst managed to do so in 1991 and Kurt Wuethrich followed in 2002!

The Road…. . • Purcell had worked on the development and application of RADAR during World War II at Massachusetts Institute of Technology's Radiation Laboratory. • His work during that project on the production and detection of radiofrequency energy, and on the absorption of such energy by matter, preceded his discovery of NMR and probably contributed to his understanding of it and related phenomena.

• They noticed that magnetic nuclei, like 1 H and 31 P, could absorb RF energy when placed in a magnetic field of a specific strength. When this absorption occurs the nucleus is described as being in resonance. • Interestingly, for analytical scientists, different atoms within a molecule resonate at different frequencies at a given field strength. • The observation of the resonance frequencies of a molecule allows a user to discover structural information about the molecule.
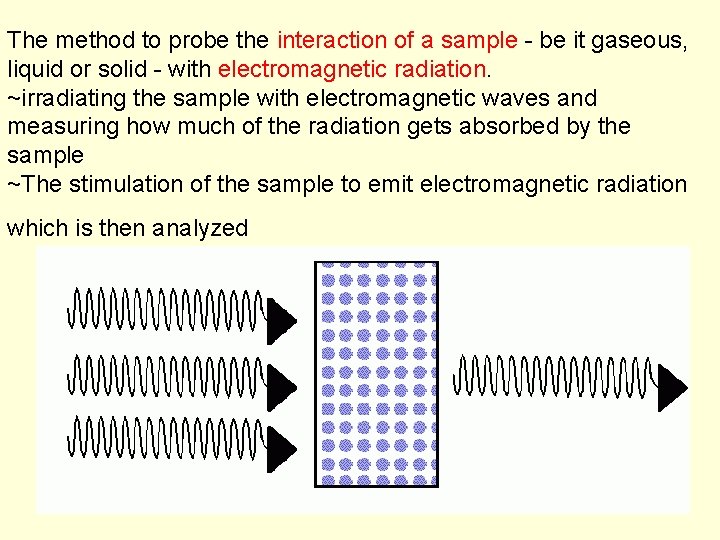
The method to probe the interaction of a sample - be it gaseous, liquid or solid - with electromagnetic radiation. ~irradiating the sample with electromagnetic waves and measuring how much of the radiation gets absorbed by the sample ~The stimulation of the sample to emit electromagnetic radiation which is then analyzed

Basics…. Nuclear makeup The nucleus of an atom consists of protons and neutrons bound by the strong force. These are further composed of subatomic fundamental particles known as quarks
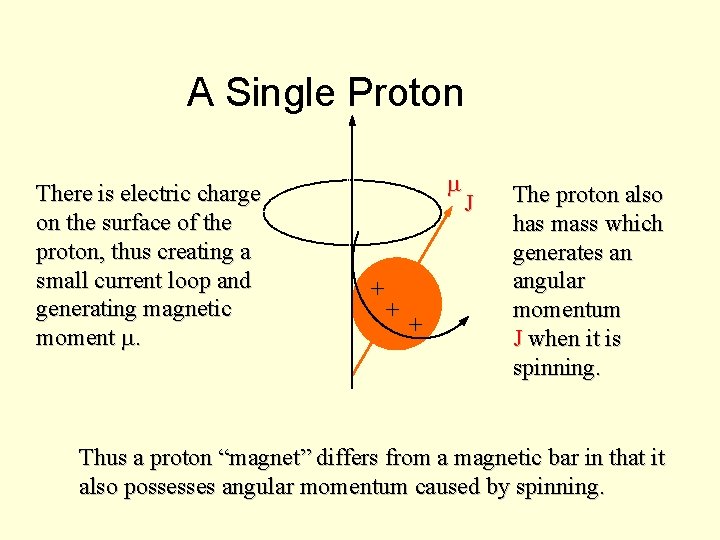
A Single Proton There is electric charge on the surface of the proton, thus creating a small current loop and generating magnetic moment m. m + + + J The proton also has mass which generates an angular momentum J when it is spinning. Thus a proton “magnet” differs from a magnetic bar in that it also possesses angular momentum caused by spinning.
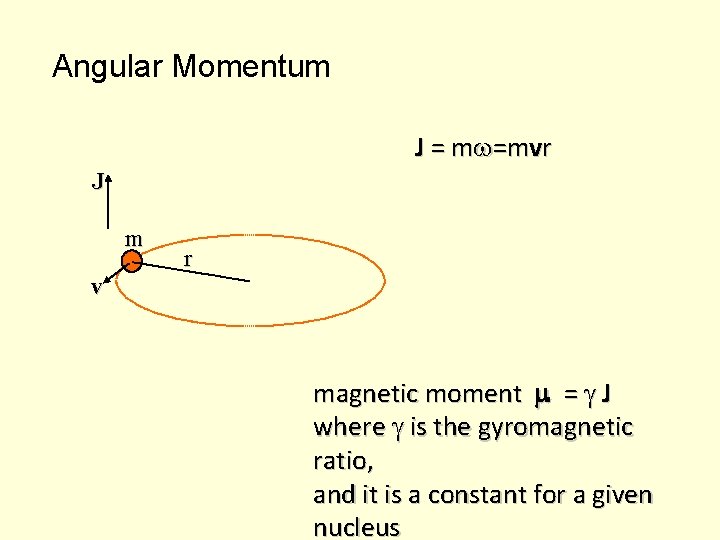
Angular Momentum J = m v r J m r v magnetic moment = J where is the gyromagnetic ratio, and it is a constant for a given nucleus

• Spin is a fundamental property of nature like electrical charge or mass. Spin (I) comes in multiples of 1/2. Protons, electrons, and neutrons possess spin. Individual unpaired electrons, protons, and neutrons each possesses a spin of 1/2. • Two or more particles with spins having opposite signs can pair up to eliminate the observable manifestations of spin. An example is helium. In nuclear magnetic resonance, it is unpaired nuclear spins that are of importance. • Nuclei with Spin : The shell model for the nucleus tells us that nucleons, just like electrons, fill orbitals. Because nucleons have spin, just like electrons do, their spin can pair up when the orbitals are being filled and cancel out. Almost every element in the periodic table has an isotope with a non zero nuclear spin.

• If an external magnetic field is applied, an energy transfer is possible between the base energy to a higher energy level (generally a single energy gap). • The energy transfer takes place at a wavelength that corresponds to radio frequencies and when the spin returns to its base level, energy is emitted at the same frequency. The signal that matches this transfer is measured in many ways and processed in order to yield an NMR spectrum for the nucleus concerned.
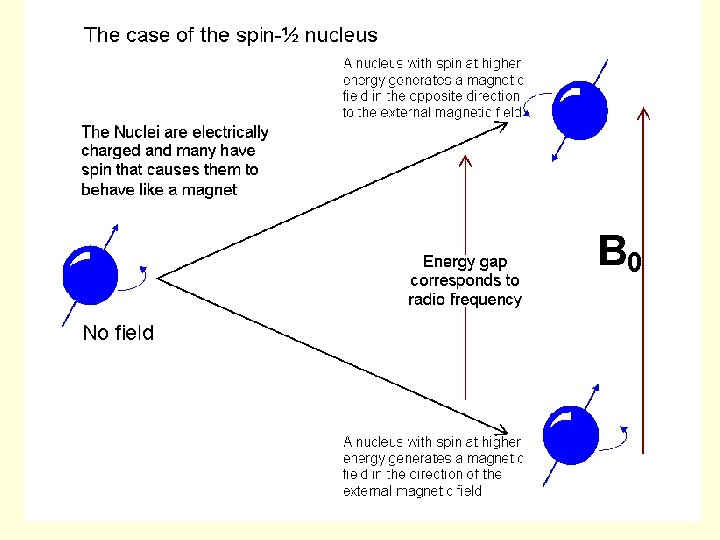
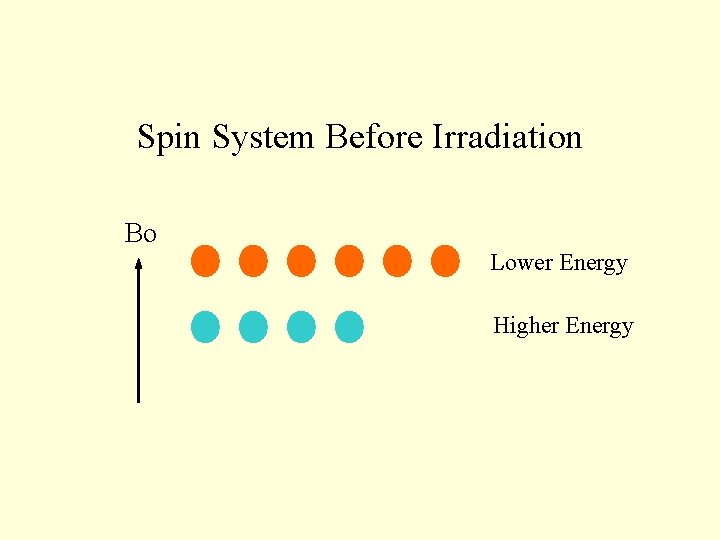
Spin System Before Irradiation Bo Lower Energy Higher Energy
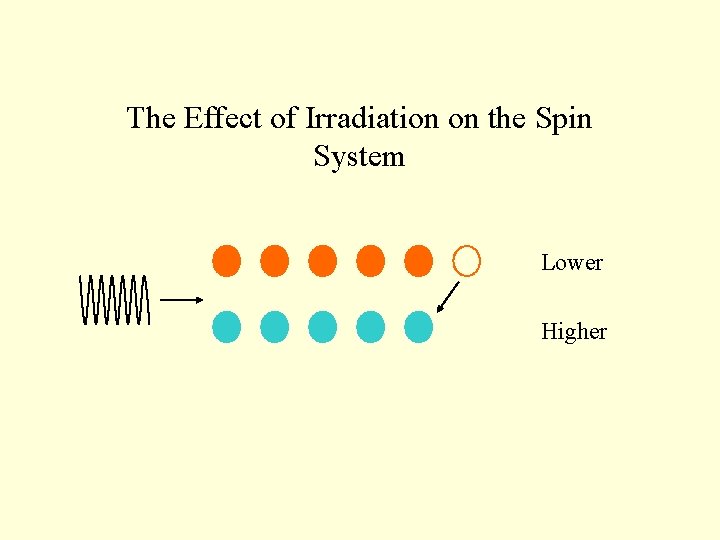
The Effect of Irradiation on the Spin System Lower Higher
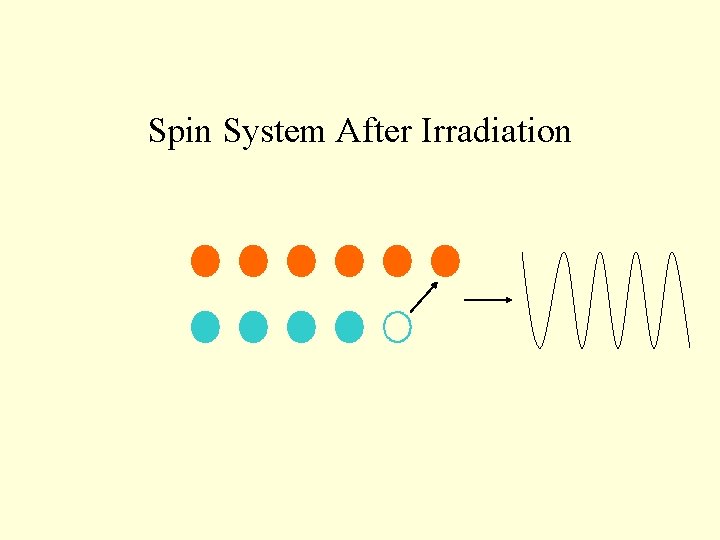
Spin System After Irradiation

Chemical Shift is…. In nuclear magnetic resonance (NMR), the chemical shift describes the dependence of nuclear magnetic energy levels on the electronic environment in a molecule

• An atomic nucleus can have a magnetic moment (nuclear spin), which gives rise to different energy levels and resonance frequencies in a magnetic field. • The total magnetic field experienced by a nucleus includes local magnetic fields induced by currents of electrons in the molecular orbitals (note that electrons have a magnetic moment themselves). • The electron distribution of the same type of nucleus (e. g. 1 H, 13 C, 15 N) usually varies according to the local geometry (binding partners, bond lengths, angles between bonds, . . . ), and with it the local magnetic field at each nucleus.

• This is reflected in the spin energy levels (and resonance frequencies). The variations of nuclear magnetic resonance frequencies of the same kind of nucleus, due to variations in the electron distribution, is called the chemical shift. • The size of the chemical shift is given with respect to a reference frequency or reference sample, usually a molecule with a barely distorted electron distribution.
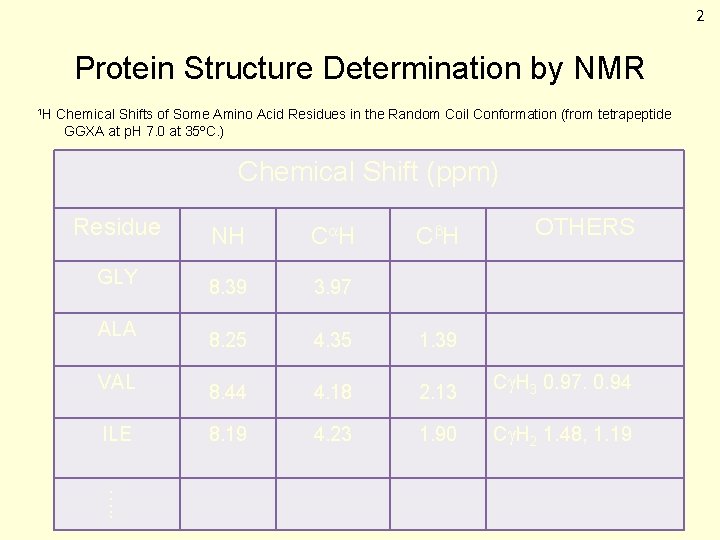
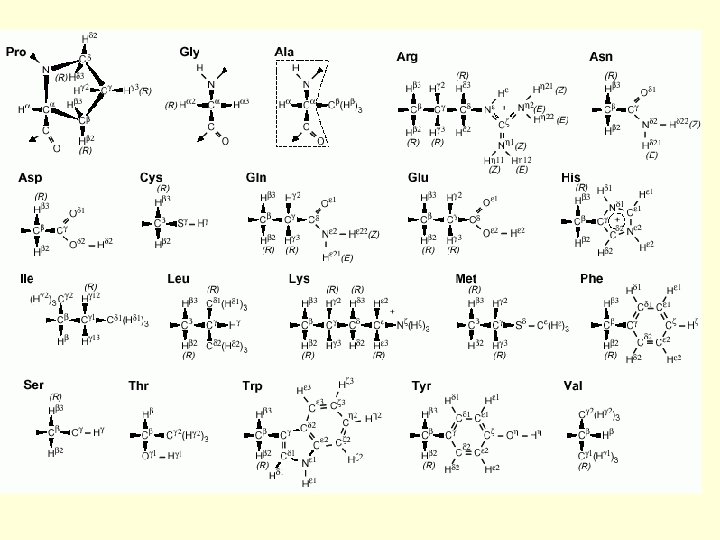
2 Protein Structure Determination by NMR 1 H Chemical Shifts of Some Amino Acid Residues in the Random Coil Conformation (from tetrapeptide GGXA at p. H 7. 0 at 35ºC. ) Chemical Shift (ppm) Residue GLY ALA VAL ILE C H OTHERS NH C H 8. 39 3. 97 8. 25 4. 35 1. 39 8. 44 4. 18 2. 13 C H 3 0. 97. 0. 94 8. 19 4. 23 1. 90 C H 2 1. 48, 1. 19 …. .

Put it together…… • Several nuclei have intrinsic magnetic moment • Concentrated homogenous solution of a protein(or NA) - a very powerful magnetic field – spin orientation • By applying RF magnetic fields to the sample, we can measure the energy absorbed at the frequency corresponding to the jump between two allowed spin orientation

• Each atom has a characteristic resonance which depends on its structure as well as environment • These subtle absorbance difference btw the same atom in different environments make it possible to identify which resonance corresponds to which of the protein atoms

• If two atoms are close in space, magnetic interactions between their spins can be measured. • The intensity of the interaction decays rapidly with the distance between them…this we can exploit to map short distances between interacting atoms

• So the result will be set of lower and upper limits for the distance btw pairs of atoms • Finally if no. of constraint is sufficient, there will be finite no. of possible conformations of the protein compatible with the data
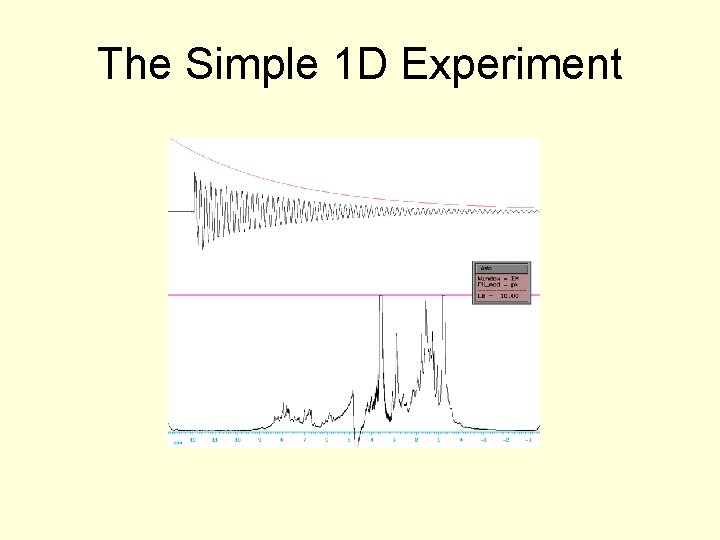
The Simple 1 D Experiment
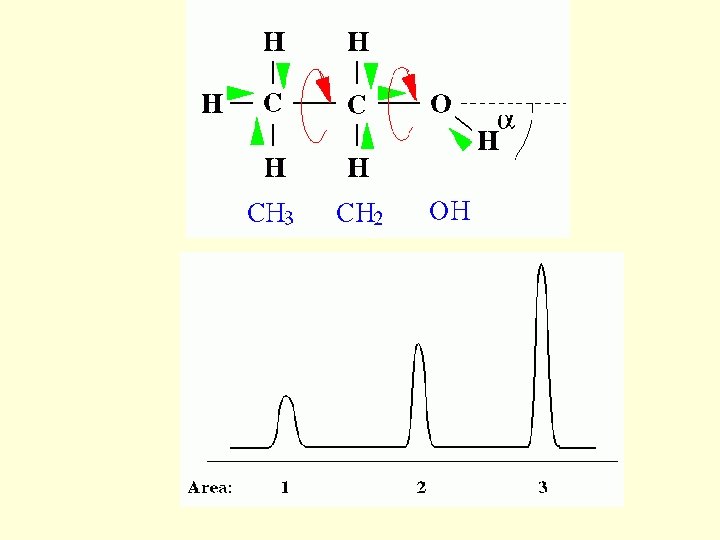
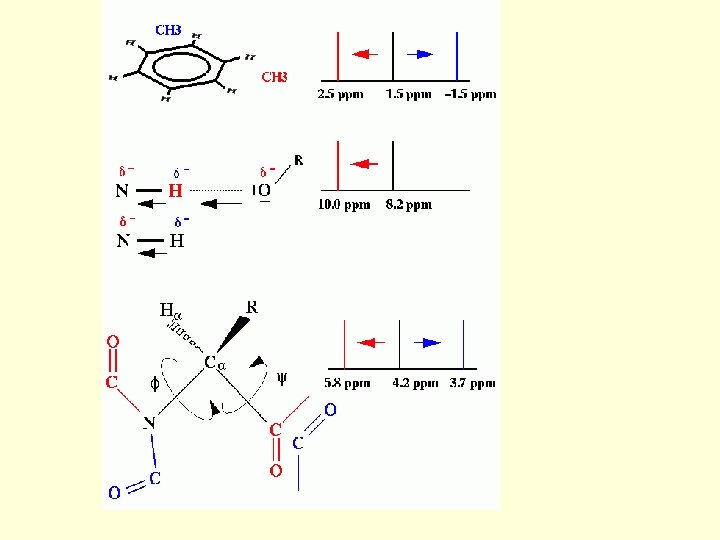

NMR spectroscopy 1. Chemical Shift: Interaction of the nuclear spins with the electronic field surrounding it. 2. Spin-Spin Coupling: Interactions of one nuclear spin with the other through chemical bond. Two Dimensional NMR 1. COrrelated Spectroscop. Y 2. TOtal Correlated Spectroscop. Y 3. Nuclear Overhauser Effect Spectroscop. Y
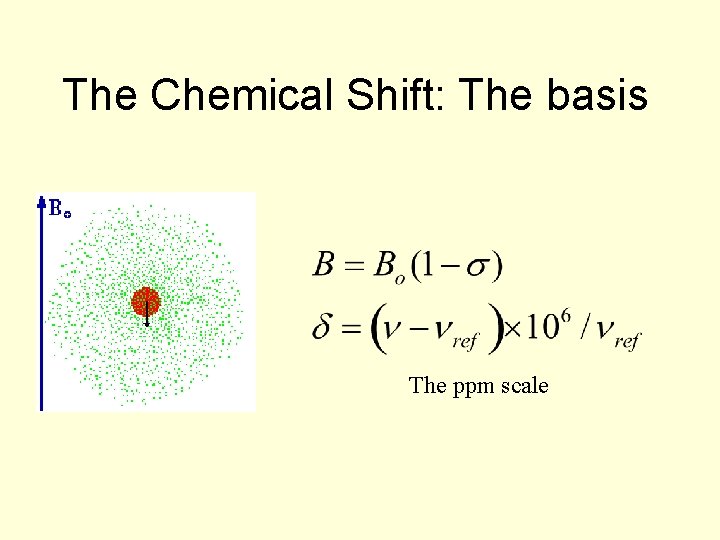
The Chemical Shift: The basis The ppm scale
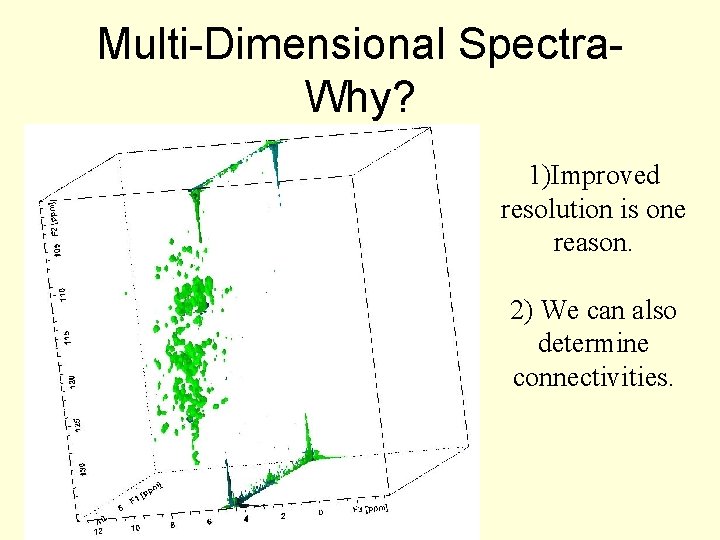
Multi-Dimensional Spectra. Why? 1)Improved resolution is one reason. 2) We can also determine connectivities.

Flowchart of Tertiary Structure determination 1. Sequence specific assignments Make use of experiments like COSY, TOCSY, etc. 2. Collect restraints from NOE Make use of experiments like NOESY, ROESY, etc… 3. Secondary structure determination Generally characteristic patterns exist for secondary structure in finger printing region 4. The constraints obtained are used in certain distance geometry algorithm to obtain tertiary structure 5. Refinement of the structure using molecular dynamics / molecular modelling
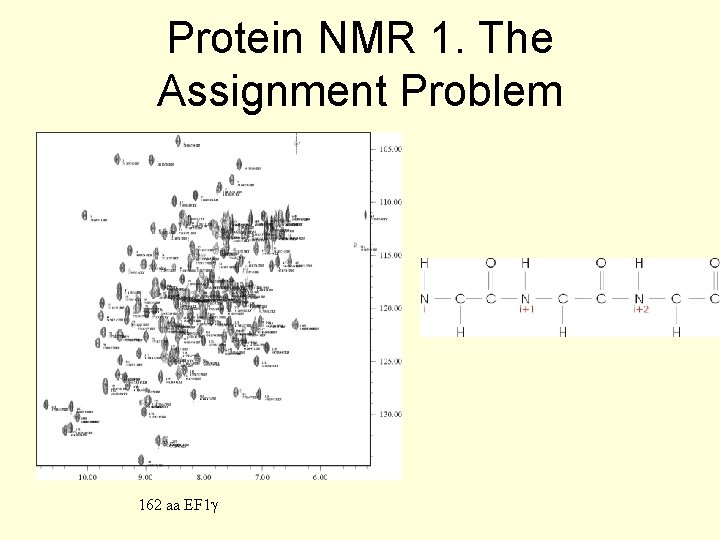
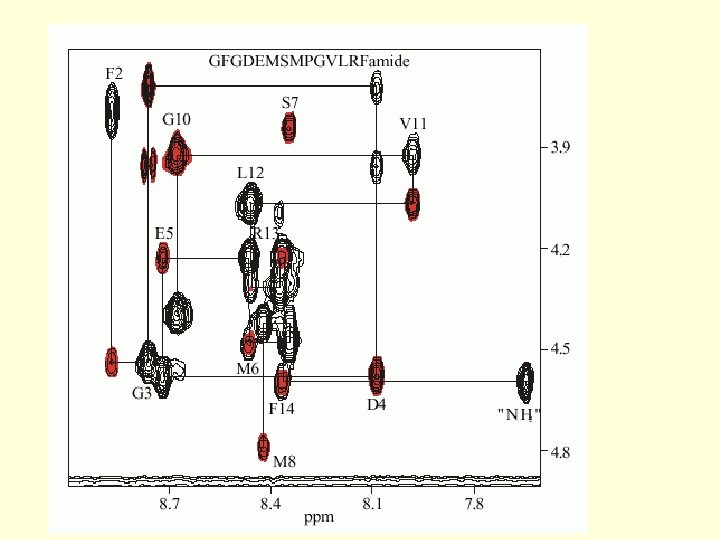
Protein NMR 1. The Assignment Problem 162 aa EF 1
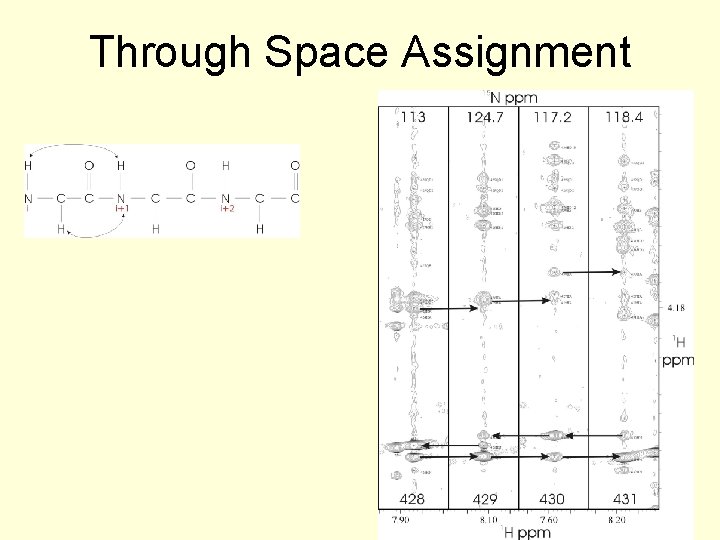
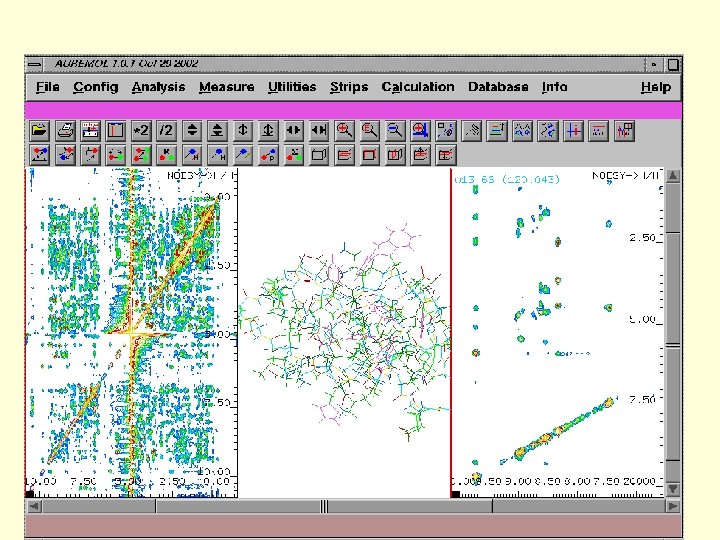
Through Space Assignment
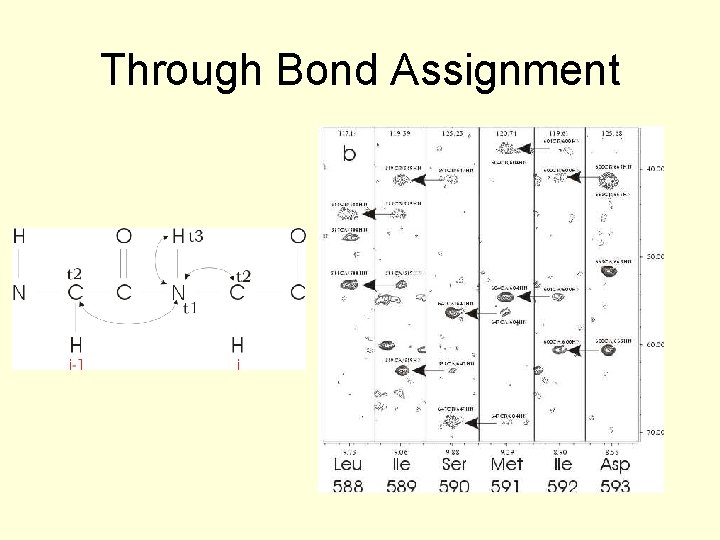
Through Bond Assignment
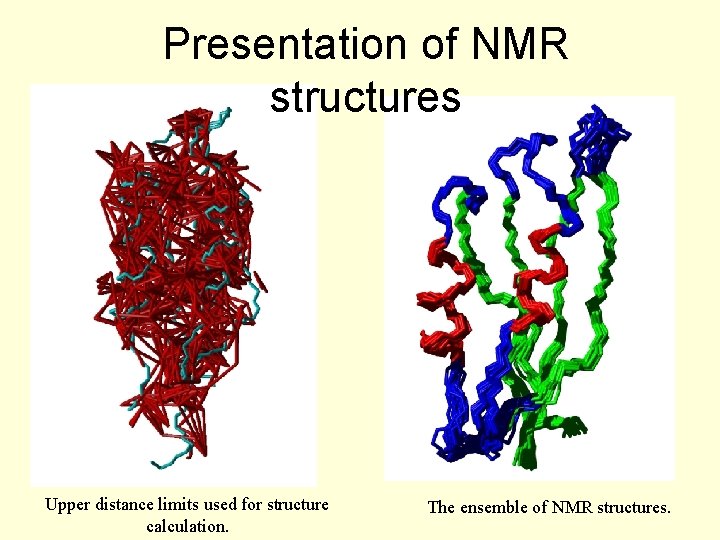
Presentation of NMR structures Upper distance limits used for structure calculation. The ensemble of NMR structures.
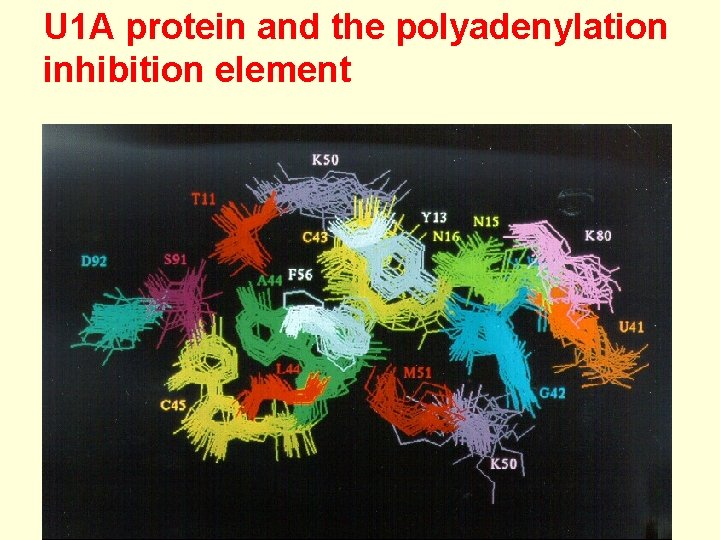
U 1 A protein and the polyadenylation inhibition element

GOOD BAD Allows probing biomolecules on atomic level: each peak in spectrum = 1 atom Low sensitivity => demand for large amount of sample Determination of protein structures currently up to 60 k. D (~450 Molecular weight limit residues), other studies up to 100 k. D Biomolecular dynamics at atomic resolution Mapping of biomolecule - ligand interactions fast and precise Needs expensive isotope enrichment to study larger molecules / complexes
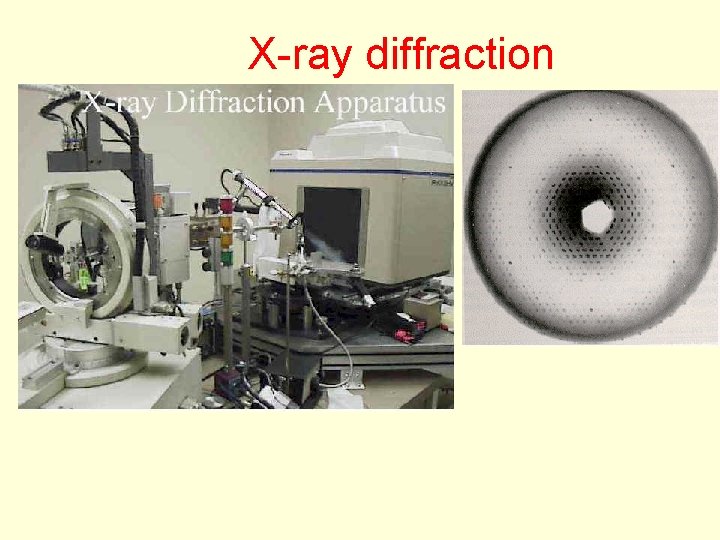
X-ray diffraction
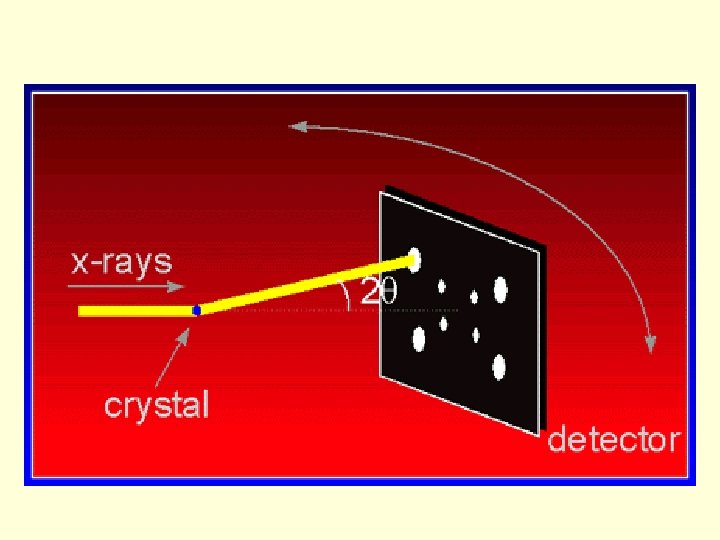

Principles • Why X-rays? – Cu. Ka Wavelength, 1. 5418Å – very similar to distance between bonded carbon atoms • Why electron density? – charge/mass: electrons >>atomic nuclei or protons – electrons acceleration electromagnetic radiation • Why crystals? – Huge number of molecules in same orientation
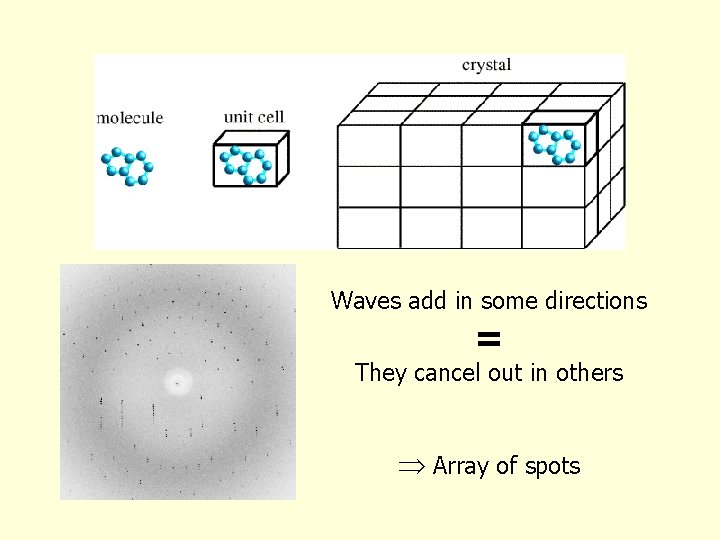
Waves add in some directions = They cancel out in others Array of spots
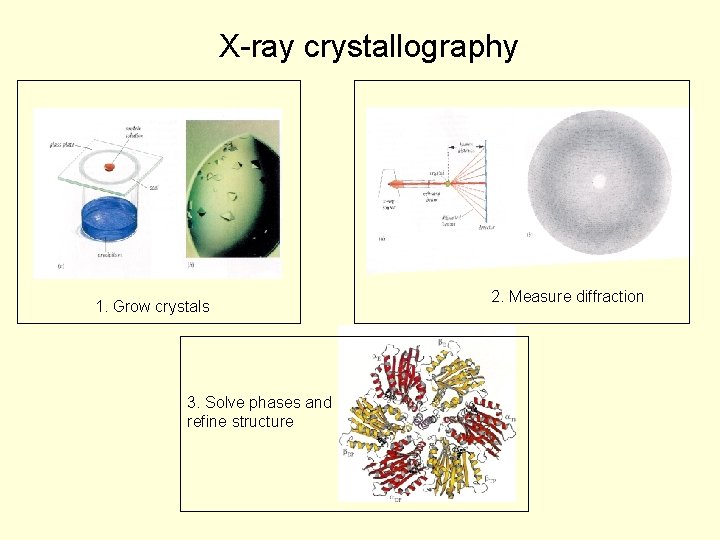
X-ray crystallography 1. Grow crystals 3. Solve phases and refine structure 2. Measure diffraction
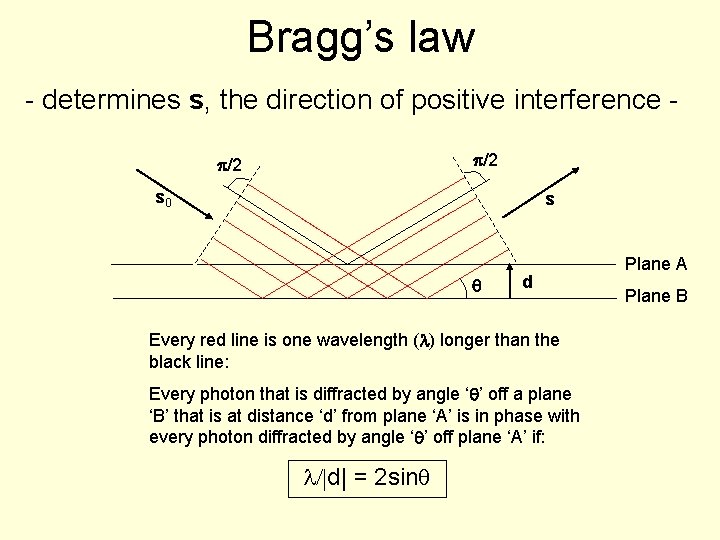
Bragg’s law - determines s, the direction of positive interference p/2 s 0 s q d Every red line is one wavelength (l) longer than the black line: Every photon that is diffracted by angle ‘q’ off a plane ‘B’ that is at distance ‘d’ from plane ‘A’ is in phase with every photon diffracted by angle ‘q’ off plane ‘A’ if: l/|d| = 2 sinq Plane A Plane B
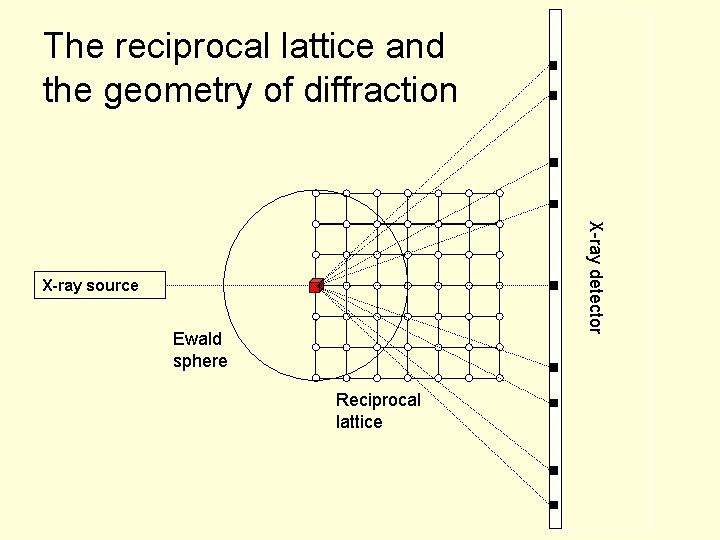
The reciprocal lattice and the geometry of diffraction X-ray detector X-ray source Ewald sphere Reciprocal lattice
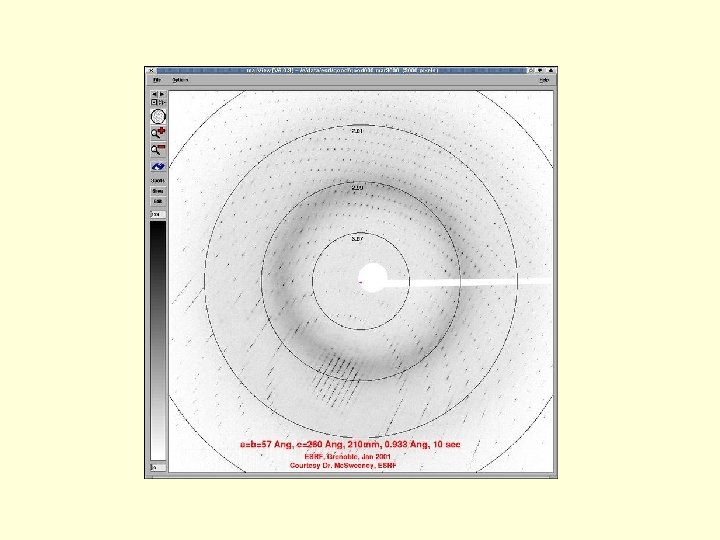

Requirement of X-ray diffraction 1. The molecule must be crystallized, and the crystals must be singular (not 2 or more stuck together) and of perfect quality 2. The phases of the waves which formed each spot must also be determined in order to produce an electron density map. (Isomorphous replacement) 3. One then interprets this image by building a model of the protein to fit the map

Fourier theory & the Phase Problem
Fourier Theory • Diffraction pattern related to object • Mathematical operation called Fourier Transform • Can be inverted to give pattern of electron density • Requires amplitude and phase of diffracted waves The Phase Problem
The Phase Problem • What we need: • What we have: • What we can get: • What we miss: • Phase and Amplitude of diffracted waves • Number of X-ray photons in each spot • Number of Photons Intensity Amplitude 2 Phase angles • Relative phase has been lost for different spots

NMR X-ray crystallography short time scale, protein folding long time scale, static structure solution, purity single crystal, purity < 20 k. D, domain any size, domain, complex atomic nuclei, chemical bonds electron density resolution limit 2 -3. 5Å primary structure must be known

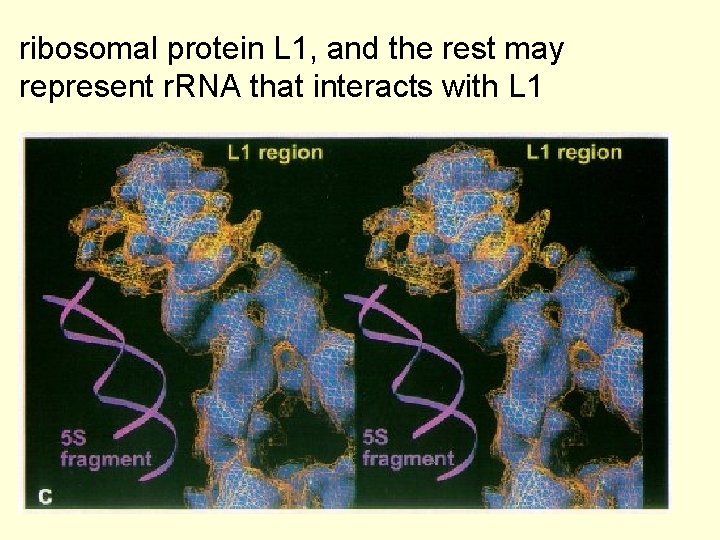
ribosomal protein L 1, and the rest may represent r. RNA that interacts with L 1
- Slides: 58