Detection Methods Joan B Rose rosejomsu edu Presentation

Detection Methods Joan B. Rose rosejo@msu. edu

Presentation will cover • Types of organisms: bacteria, parasites and viruses • Units of measure • Cultivation (to measure viability) versus molecular tools • Brief overview of environmental sampling • Attributes of methods

Enumeration of Microbes • Quantitative measurements • Quantal measurements (presence/absence) • For Bacteria: CFU=Colony Forming Unit • For Viruses: PFU =Plaque Forming Unit • For Parasites (& sometimes bacteria) cell counts using microscopy • MPNMost Probable Number (statistical method to estimate concentrations with multiple assays at dilutions to extinction [0])
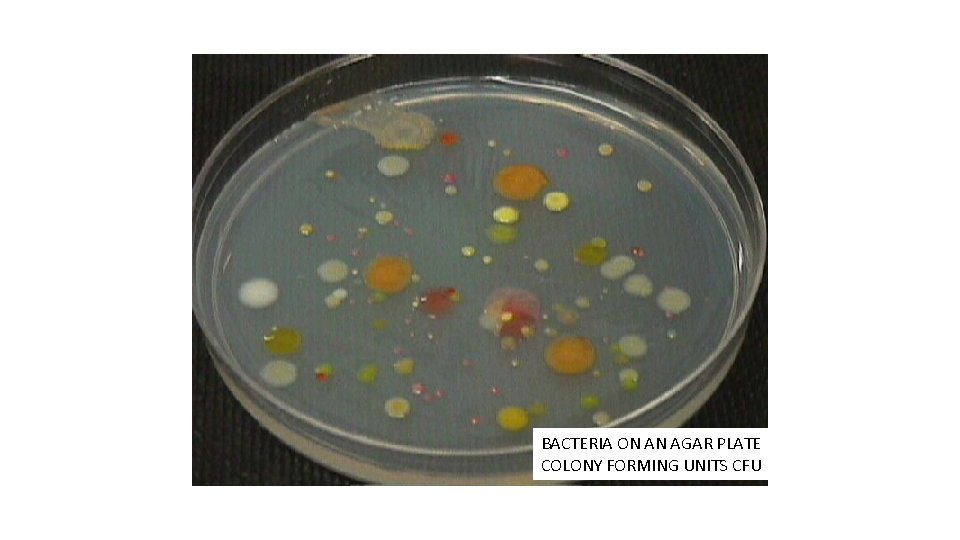
BACTERIA ON AN AGAR PLATE COLONY FORMING UNITS CFU
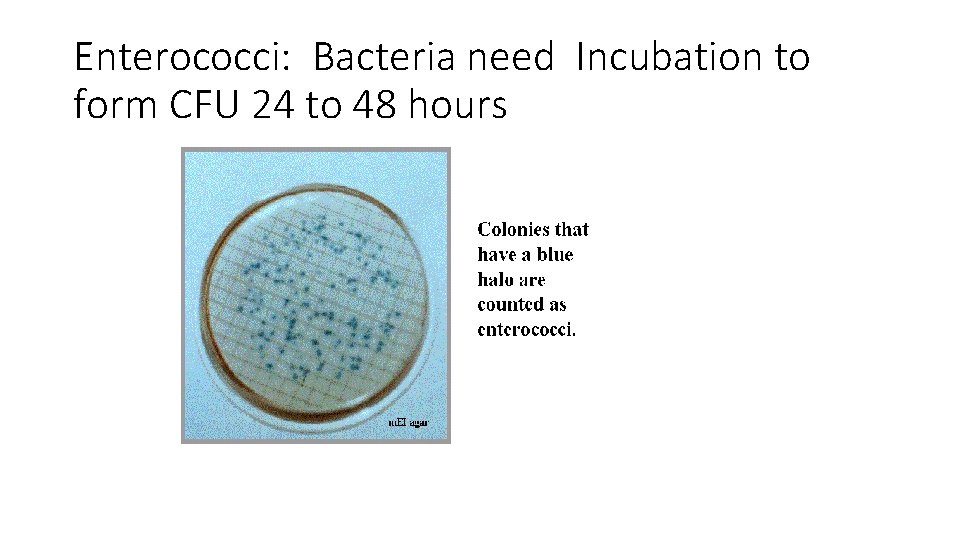
Enterococci: Bacteria need Incubation to form CFU 24 to 48 hours

Growth Based Methods: Common Fecal Indicator Organisms for measuring water quality Filtering 100 ml water samples Total coliforms MPN Fecal coliforms E. coli Agar and colonies MPN and colonies
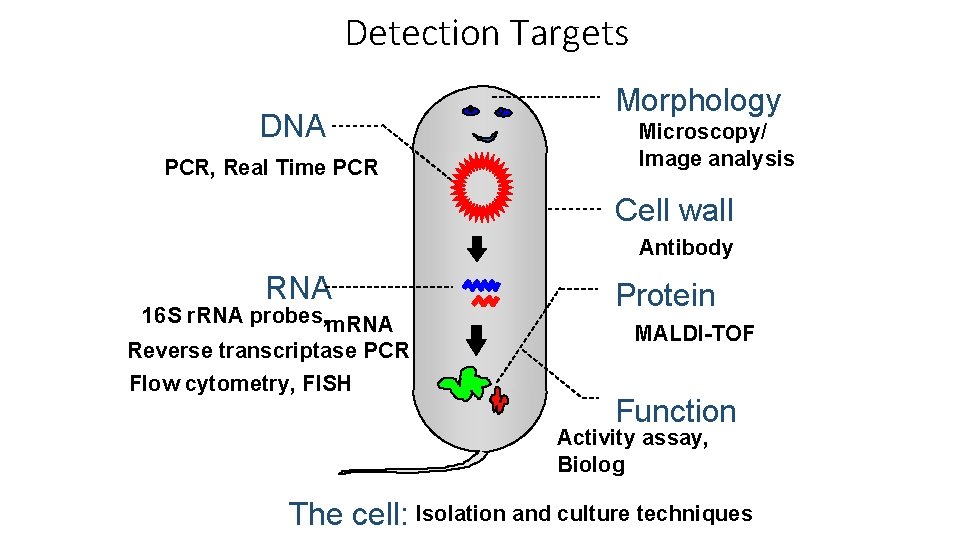
Detection Targets DNA PCR, Real Time PCR Morphology Microscopy/ Image analysis Cell wall Antibody RNA 16 S r. RNA probes, m. RNA Reverse transcriptase PCR Flow cytometry, FISH Protein MALDI-TOF Function Activity assay, Biolog The cell: Isolation and culture techniques
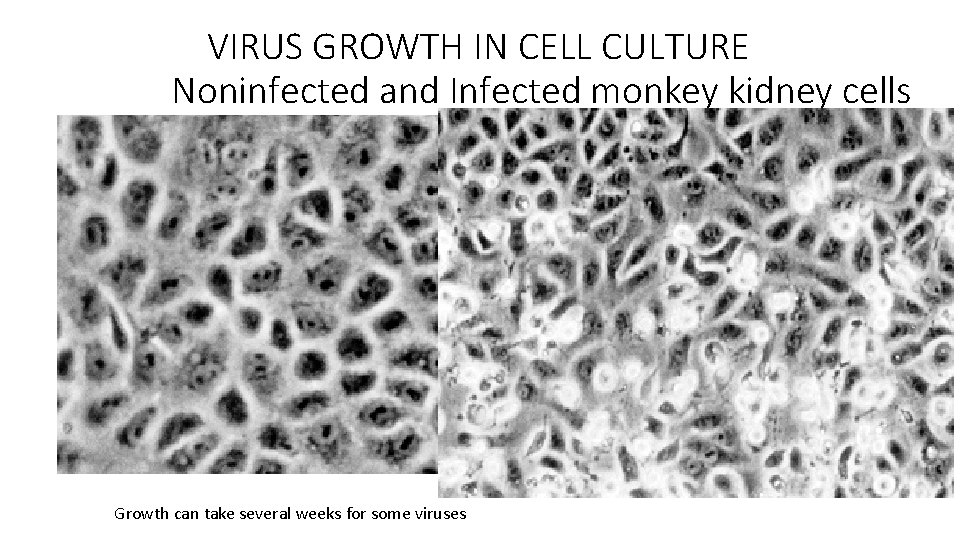
VIRUS GROWTH IN CELL CULTURE Noninfected and Infected monkey kidney cells Growth can take several weeks for some viruses
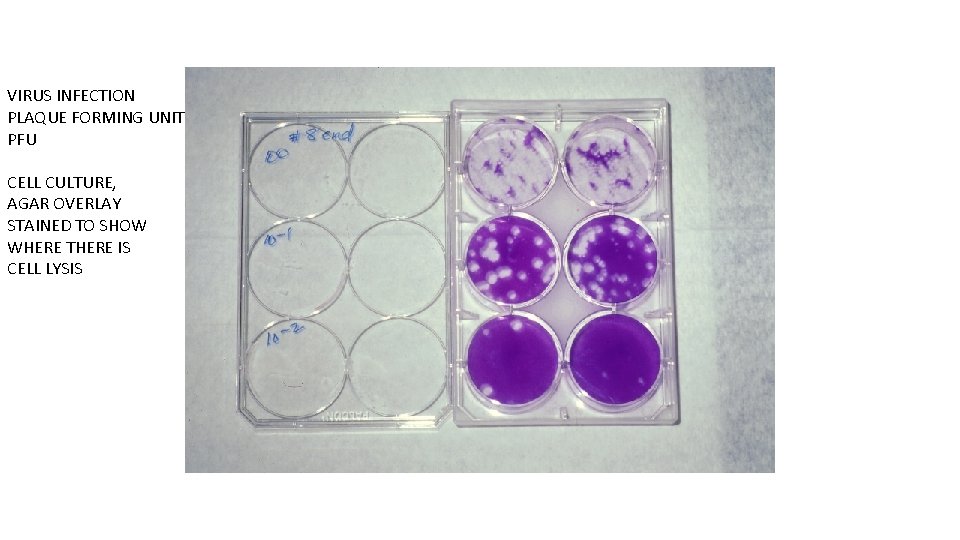
VIRUS INFECTION PLAQUE FORMING UNIT PFU CELL CULTURE, AGAR OVERLAY STAINED TO SHOW WHERE THERE IS CELL LYSIS
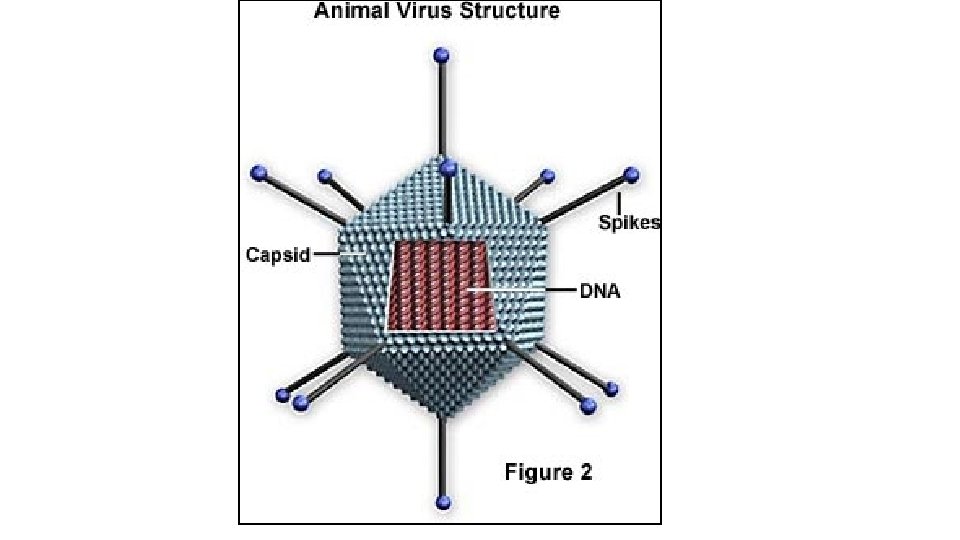
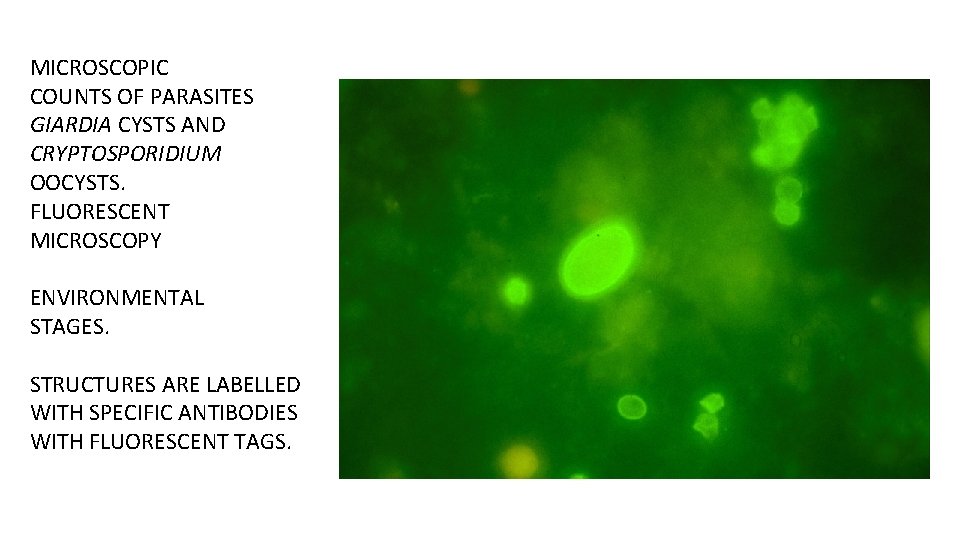
MICROSCOPIC COUNTS OF PARASITES GIARDIA CYSTS AND CRYPTOSPORIDIUM OOCYSTS. FLUORESCENT MICROSCOPY ENVIRONMENTAL STAGES. STRUCTURES ARE LABELLED WITH SPECIFIC ANTIBODIES WITH FLUORESCENT TAGS.

Viruses Three Dimensions of Detection Rotavirus Hepatitis A Norwalk virus Protozoa EPA: Indicator organisms E. coli Fecal enterococci Bacteroides spp. Bifidobacteirum spp. Clostridium spp. Eubacterium spp. Background Organism (List of phgylogentic groups) CDC Category B- Water Yersinia enterocolitica Campylobacter Listeria monocytogenes Salmonella (Newport and DT 104) Pathogenic vibrios Shigella Diarrheagenic E. coli Breadth: Species Cryptosporidium Giardia ce an ce, n Fu n tio : ic iot st i s e vi en rul r tib an c 9 8 1 7 H 1 1 H : : 1 H 1 2 12 11 26: 157 K 1 O O i l i i co col col. co. . . E. E E Depth: Strain
Polymerase Chain Reaction (PCR) • Developed in 1985 by Kary Mullis • Dr. Mullis received the Nobel Prize in Chemistry in 1993 • PCR is considered as one of the most important discoveries in molecular biology

What is PCR? • Enzymatic reaction that makes many copies of DNA from single molecule • 2 n copies of DNA from single molecule where n = No. of cycles • So, 35 cycles of PCR would yield 235 copies of DNA
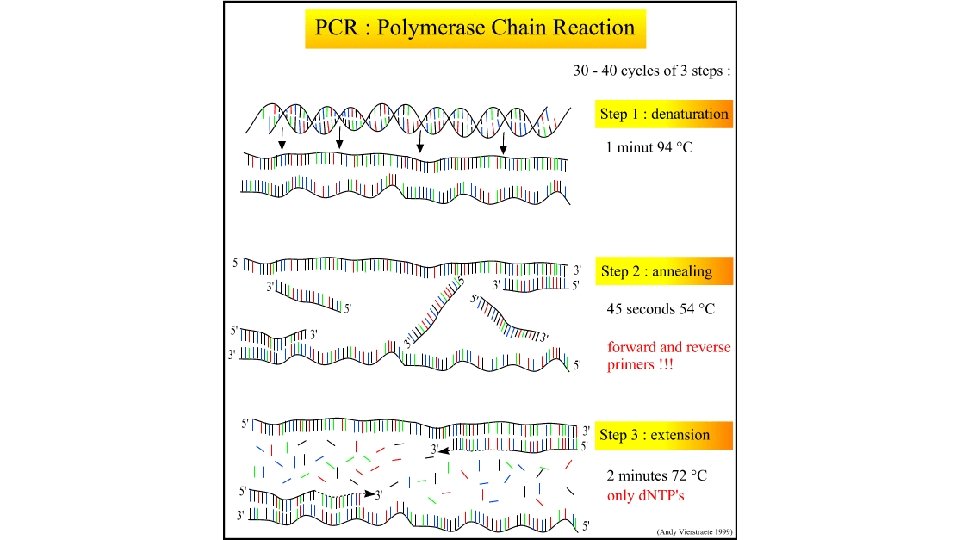
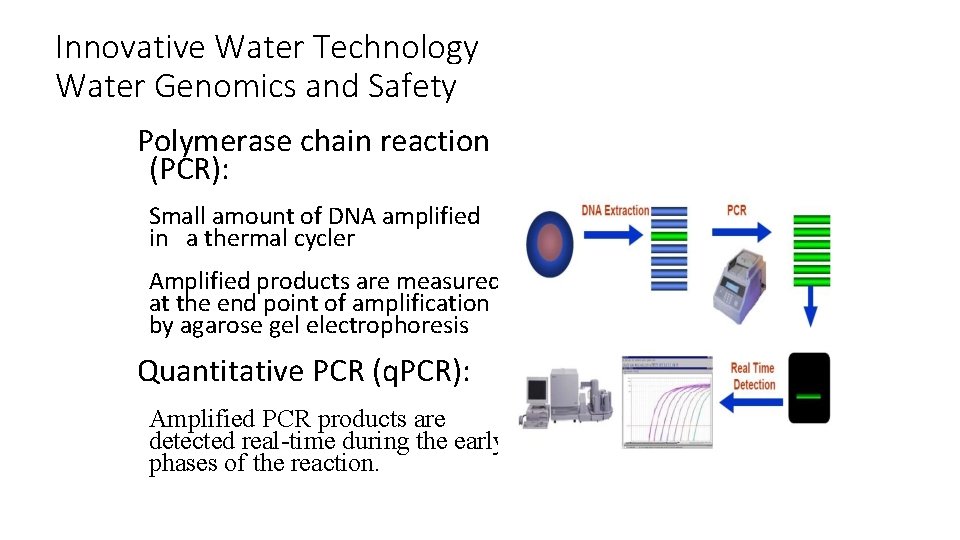
Innovative Water Technology Water Genomics and Safety Polymerase chain reaction (PCR): Small amount of DNA amplified in a thermal cycler Amplified products are measured at the end point of amplification by agarose gel electrophoresis Quantitative PCR (q. PCR): Amplified PCR products are detected real-time during the early phases of the reaction. 16

Applications ØAllows for amplification and identification of specific gene sequences from specific organisms (i. e. bacteria, parasites, viruses, etc. ) that would otherwise be undetectable ØProvides information for HAZ ID ØProvides information for sources (HAZ) ØUsed for Forensic studies (Bioterrorism) ØUsed for molecular epidemiology (outbreak investigation)
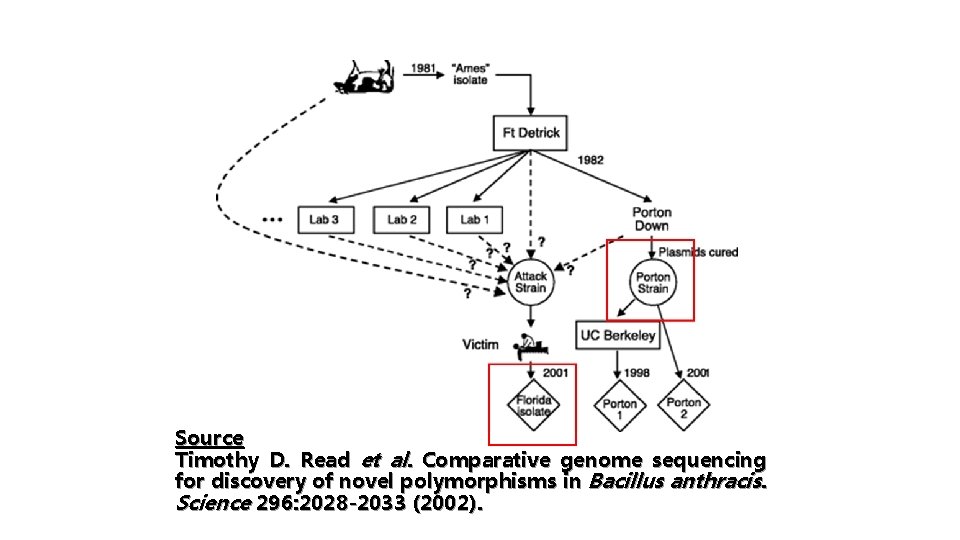
Source Timothy D. Read et al. Comparative genome sequencing for discovery of novel polymorphisms in Bacillus anthracis. Science 296: 2028 -2033 (2002).

Steps Involved in the Detection of Environmental Pathogens Sample Collection Concentration Purification Replication Identification Quantification Isolation and/ or Characterization

Cultural/Standard Methods for the Detection of Pathogens in the Environment • Virus • Concentration/separation • Cell culture (one cell line does not detect all) • PCR identification • Bacteria • Concentration/separation • Enrichment media • Selective media • Biochemical tests, serology, immunochemical

Methods for the Detection of Pathogens • Protozoa • Concentration or elution • Purification • Differential centrifugation • Immune Magnetic Separation • Stain with monoclonal antibodies • Observe under UV light • Examine for characteristics • Shape and size • Internal structures

Typical Sample Volumes for Water • Indicator Bacteria & Pathogenic Bacteria • 100 to 1000 ml • Viruses • Raw Sewage 1 -5 liters • Treated sewage 40 liters • Surface waters 400 liters • Ground and Drinking water 400 to 2000 liters • Protozoa • Treated sewage 4 liters • Surface waters 10 to 100 liters • Drinking water 10 to 100 liters

Cultural/Standard Methods for the Detection of Pathogens in the Environment • Virus • Concentration/separation • Cell culture (one cell line does not detect all) • Serology (identification) • Bacteria • Concentration/separation • Enrichment media • Selective media • Biochemical tests, serology, immunochemical

Methods for the Detection of Pathogens • Protozoa • Concentration or elution • Purification • Differential centrifugation • Immune Magnetic Separation • Stain with monoclonal antibodies • Observe under UV light • Examine for characteristics • Shape and size • Internal structures

Typical Sample Volumes for Water • Bacteria • 100 ml • Viruses • Raw Sewage 1 -5 liters • Treated sewage 40 liters • Surface waters 400 liters • Drinking water 400 to 2000 liters • Protozoa • Treated sewage 4 liters • Surface waters 10 to 100 liters • Drinking water 10 to 100 liters
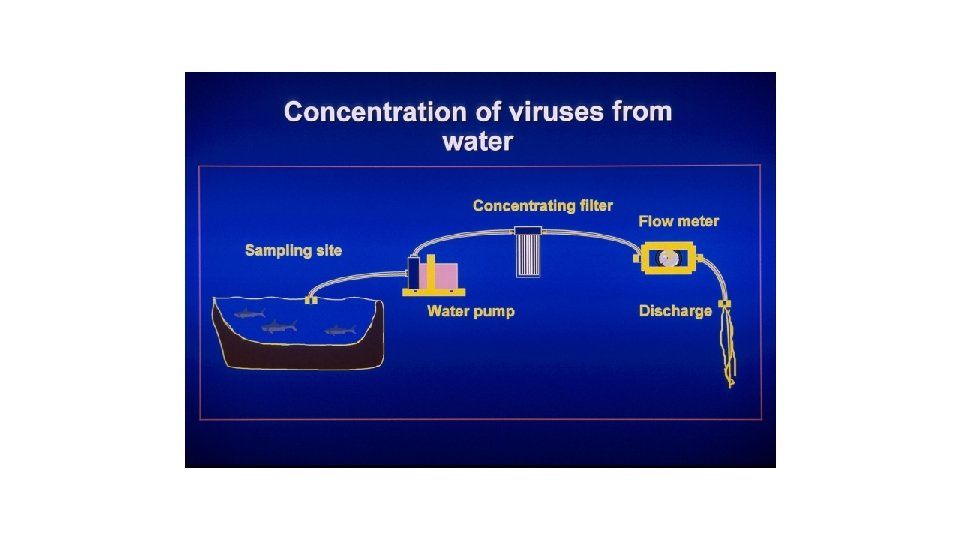
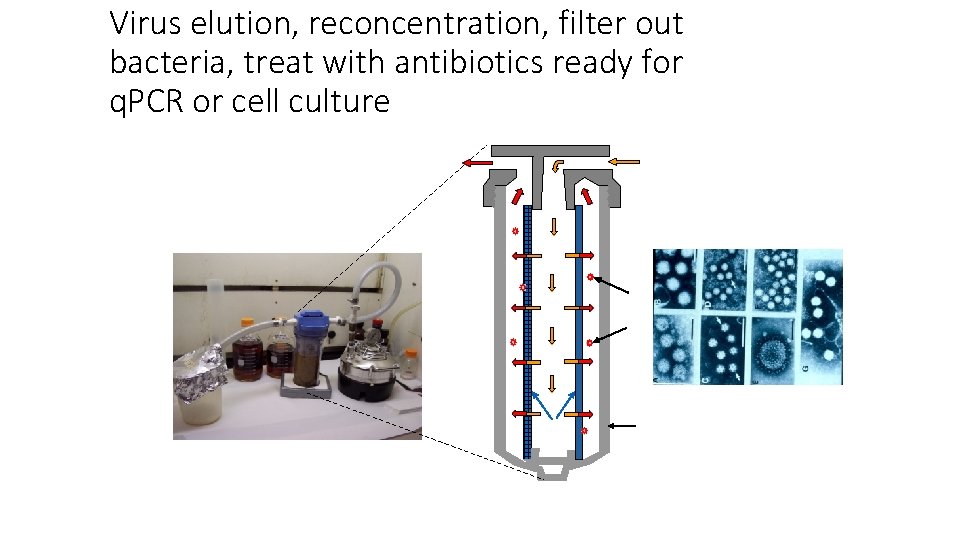
Virus elution, reconcentration, filter out bacteria, treat with antibiotics ready for q. PCR or cell culture 1. 5% beef extract (p. H 9. 5) Viral particles were eluted from 1 MDS filter using 1. 5% beef extract (p. H 9. 5) 1 MDS filter Dr, Xagoraraki, MSU Cartridge housing

Peristaltic pump Filtrate/waste Low-cost ultrafiltration system with disposable hollow fiber filters Ultrafilter (MWCO 30 k. Da) - Size-exclusion Feed Retentate Sample/ret entate reservoir - Wide range of viruses can be recovered in a single water sample

Aerosol Sampling • Impingers • Impact- Anderson Aerosol • Filters • High volume fluid samplers • Electrostatic participators • Settling plates • Efficacy affected by • Collection media • Relative humidity • Desiccation
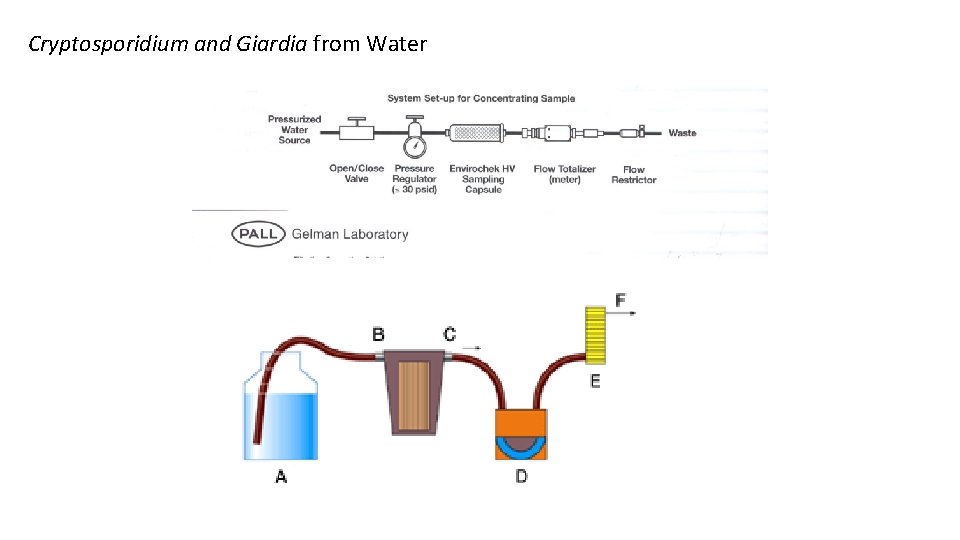
representation of the primary concentration step Cryptosporidium. Graphic and Giardia from Water A: Sample B: Filter inlet C: Filter outlet D: Peristaltic pump E: Flow meter F: To waste


Immunomagnetic Separation of Cryptosporidium parvum oocysts IMS beads captured oocysts
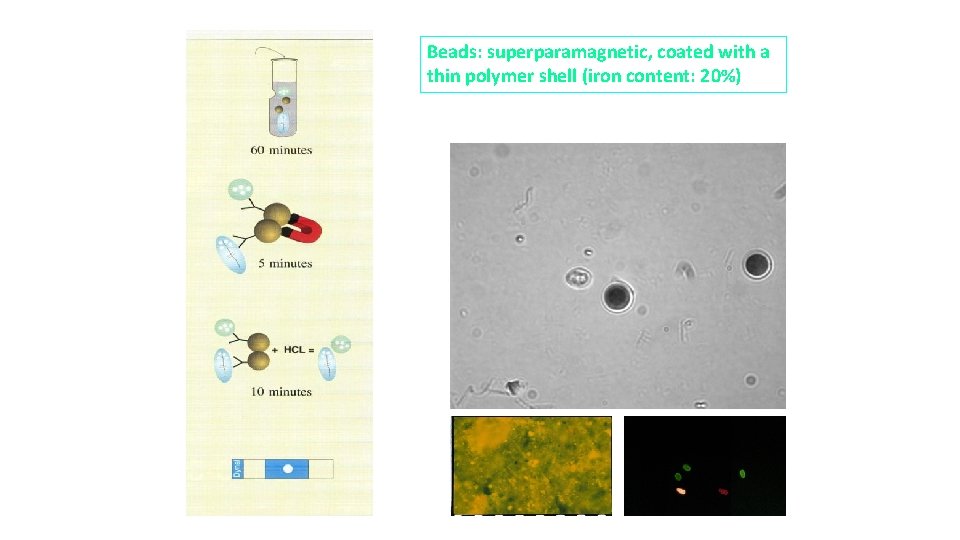
Beads: superparamagnetic, coated with a thin polymer shell (iron content: 20%)

Fomites • WHAT IS A FOMITE ? - Any inanimate object or substance capable of absorbing and transferring infectious organisms. • Examples: Table tops, computers, keyboards, glass, cloths, door knobs, kitchen surfaces and so on

Fomites Studied Acrylic Formica Ceramic tile Cotton Glass Polyester Stainless steel Money

Scanning Electron Micrographs Stainless Steel Cotton Paper Money
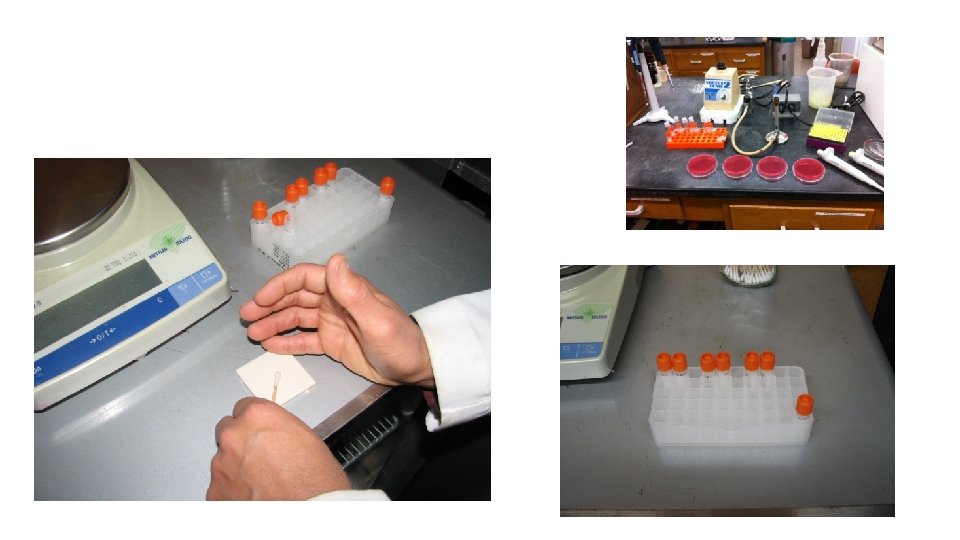

Development of virus methods for sampling fomites Agents Bacteriophage P 22 Fomites Acrylic, Stainless Steel, Laminate Fomite Area 100 cm 2, 1000 cm 2 Sample Concentration 0. 5 – 55 PFU/cm 2 Application Medium Phosphate Buffered Saline Tween-80 (PBST), Trypticase Soy Broth (TSB), Water Sampling Method Pre-moistened Wipe, Kimwipe, Cotton Swab Application Method 50 drops of 1µL Time of Recovery 0 min and 20 min (dry)
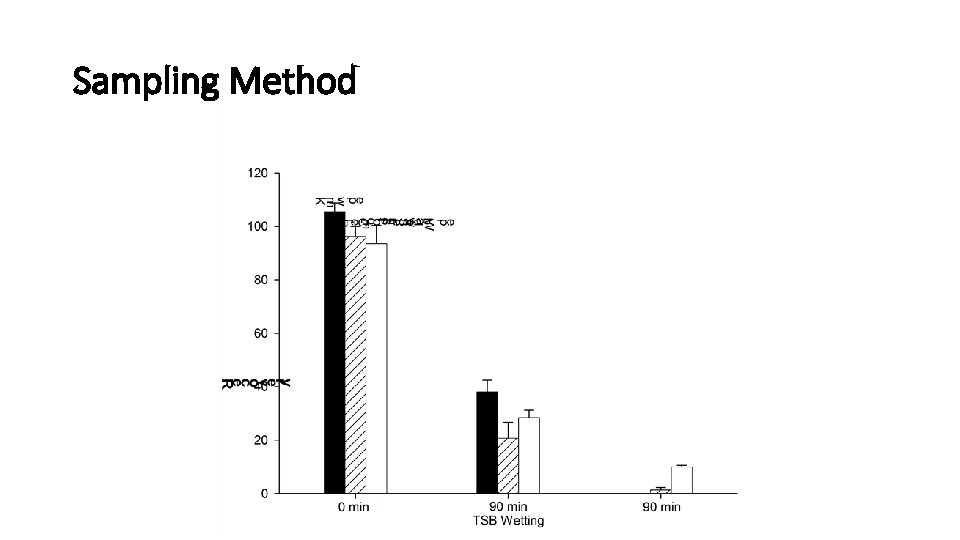
Sampling Method
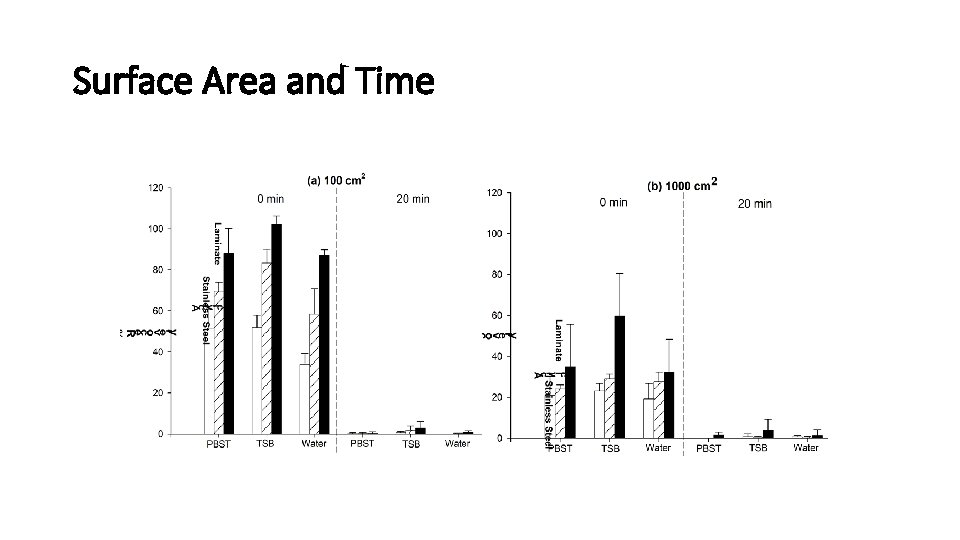
Surface Area and Time
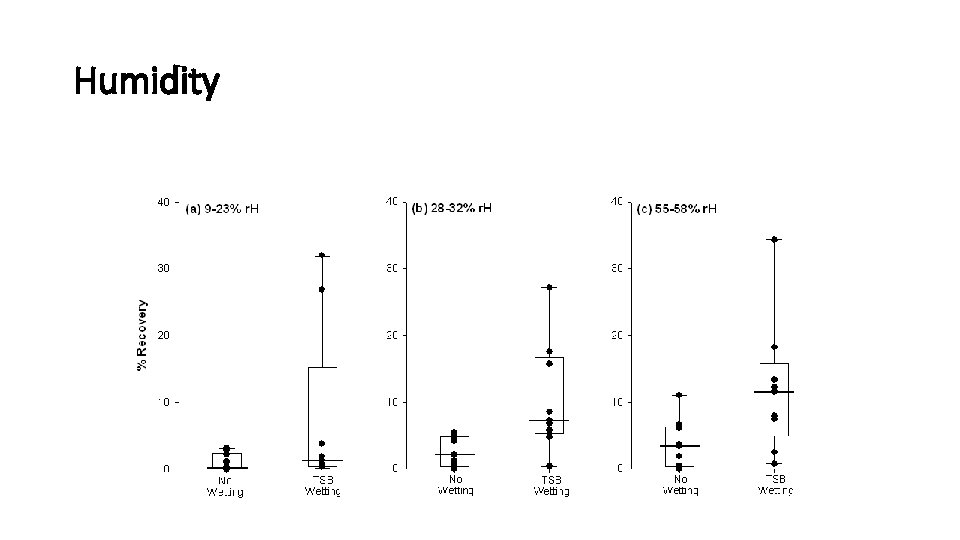
Humidity
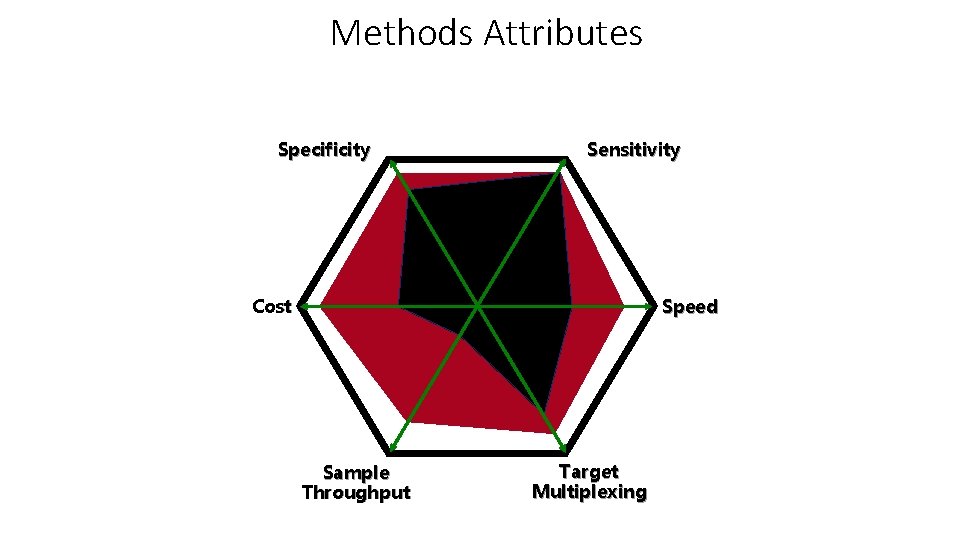
Methods Attributes Specificity Sensitivity Cost Speed Sample Throughput Target Multiplexing

Issues in Detection How low can you go? And No method is 100% • Sensitivity: Probability of a positive test result (or presence of the correct organism) given the true presence of that organism. Here you can calculate the False positives. • Specificity: Probability of a negative test result (or absence of the organism) given the true absence of the organism. Here you calculate the False negatives. • Accuracy: is the closeness of the measurement to the true quantity in the sample. • Precision: measures the reproducibility of a test. • Recovery rate: measures of what proportion of the original concentration of the organism can be recovered from a sample. • Detection limit: is the lowest concentration where a true positive is still identifiable. Volume and presence of interfering substances influence these TWO ways to test new approaches • Compare to a golden standard • Seeded recovery methods (ratio of recovered concentration to a known concentration added to the matrix)

Methods and QMRA • Used to identify the Hazard for Hazard Identification Where specificity is important; may be used in clinical but also environmental samples. • Used to measure the Dose in Dose-response experiments. Where quantification, accuracy and precision becomes important. In many cases cultivation methods are used. • Used in exposure assessment to address sources, concentrations, transport, and persistence. Where viability measures as well as quantification are important and ability to use in environmental samples.
- Slides: 45