Design of Biosensors and Arrays Toxicity Screening James

Design of Biosensors and Arrays: Toxicity Screening James F. Rusling Departments of Chemistry & Pharmacology University of Connecticut Storrs, CT, USA
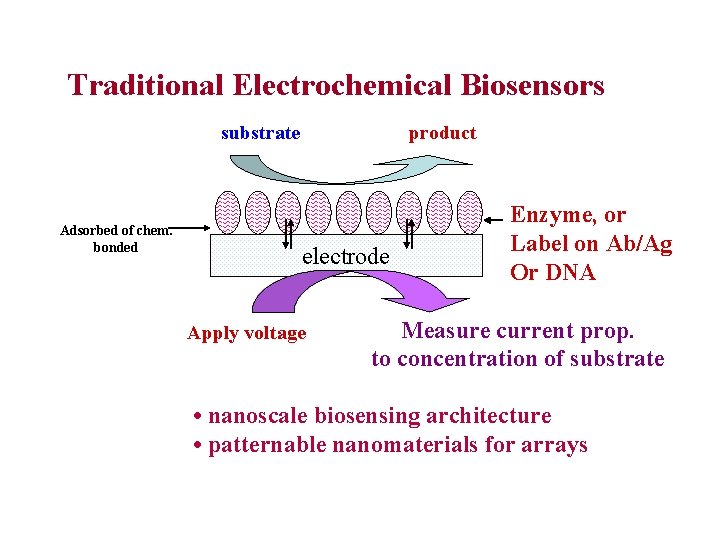
Traditional Electrochemical Biosensors substrate Adsorbed of chem. bonded product electrode Apply voltage Enzyme, or Label on Ab/Ag Or DNA Measure current prop. to concentration of substrate • nanoscale biosensing architecture • patternable nanomaterials for arrays

Layer-by-layer Film assembly Lvov, Decher Lvov, Y. in Nalwa, R. W. ; Ed. ; Handbook Of Surfaces And Interfaces Of Materials, Vol. 3. Academic, 2001, pp. 170 -189. Stable, easily prepared, versatile
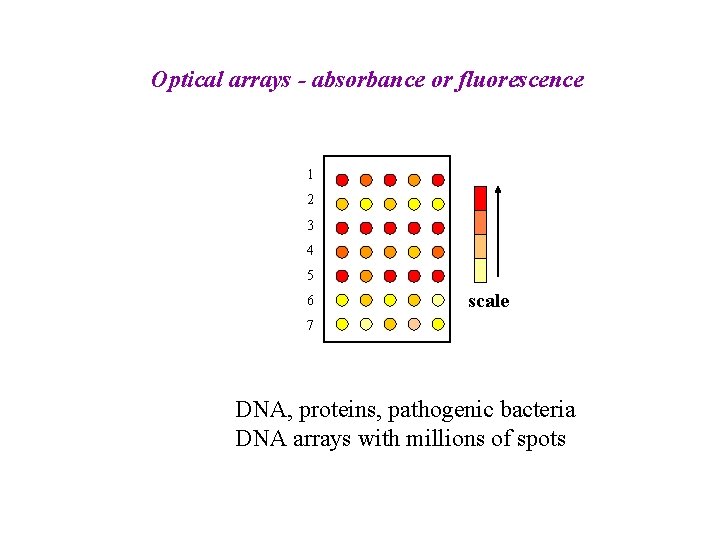
Optical arrays - absorbance or fluorescence 1 2 3 4 5 6 scale 7 DNA, proteins, pathogenic bacteria DNA arrays with millions of spots
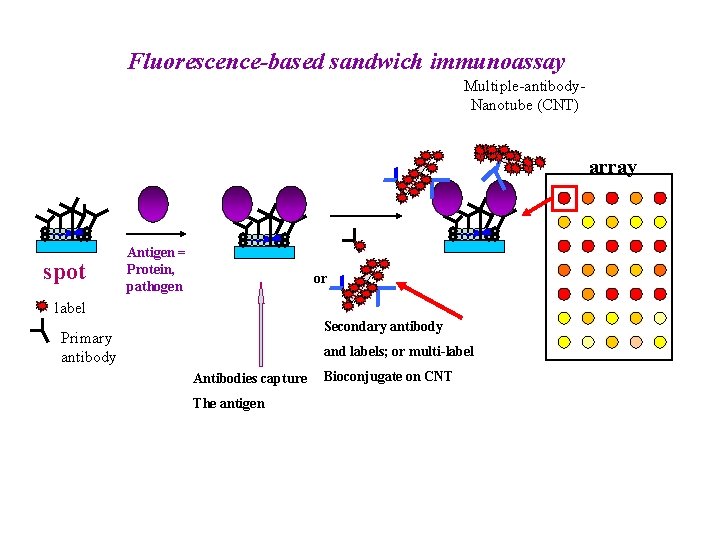
Fluorescence-based sandwich immunoassay Multiple-antibody. Nanotube (CNT) array spot Antigen = Protein, pathogen or label Secondary antibody Primary antibody and labels; or multi-label Antibodies capture The antigen Bioconjugate on CNT
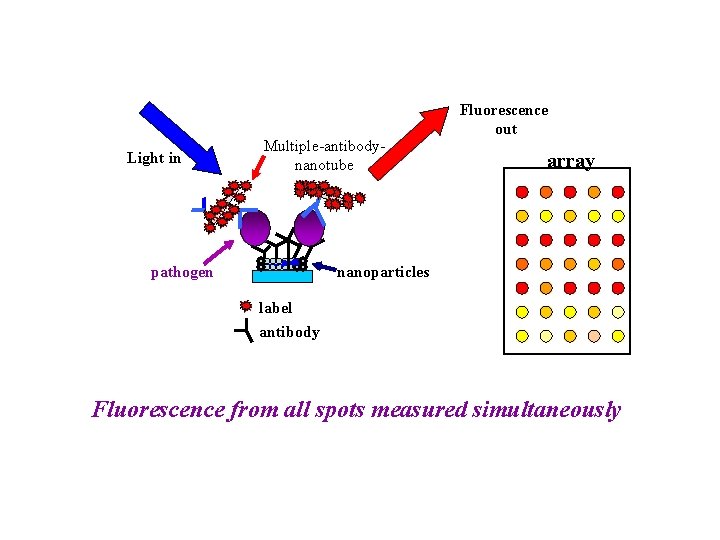
Light in Multiple-antibodynanotube pathogen Fluorescence out array nanoparticles label antibody Fluorescence from all spots measured simultaneously
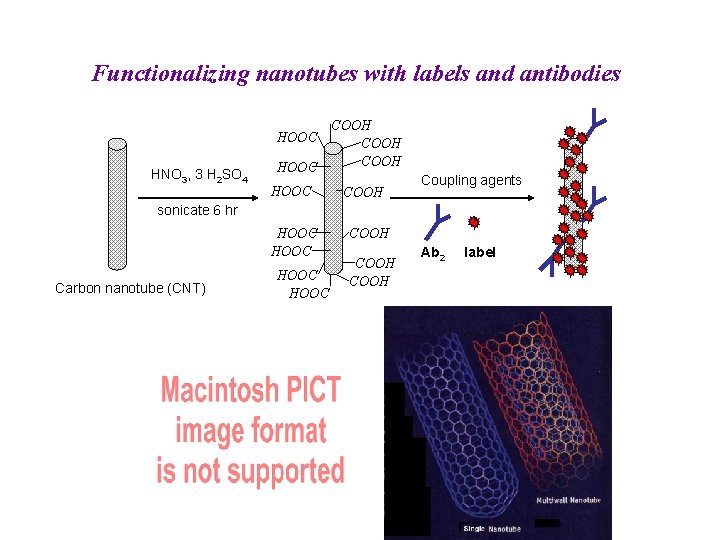
Functionalizing nanotubes with labels and antibodies HOOC HNO 3, 3 H 2 SO 4 HOOC COOH Coupling agents sonicate 6 hr HOOC Carbon nanotube (CNT) HOOC COOH Ab 2 label
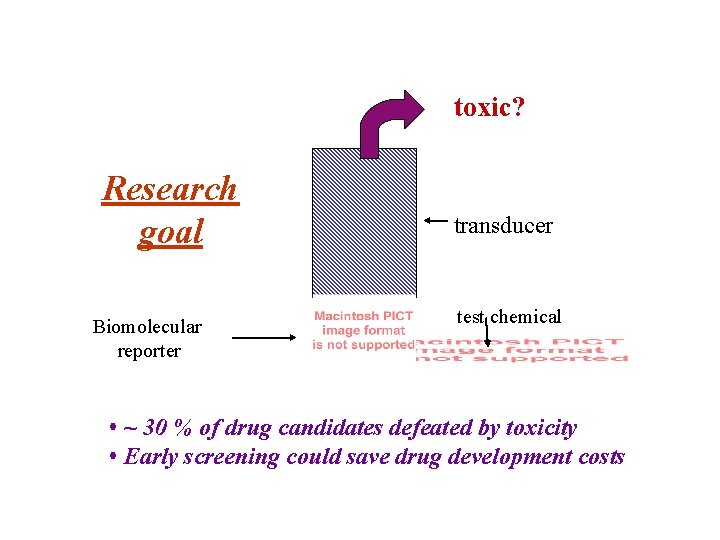
toxic? Research goal Biomolecular reporter transducer test chemical • ~ 30 % of drug candidates defeated by toxicity • Early screening could save drug development costs
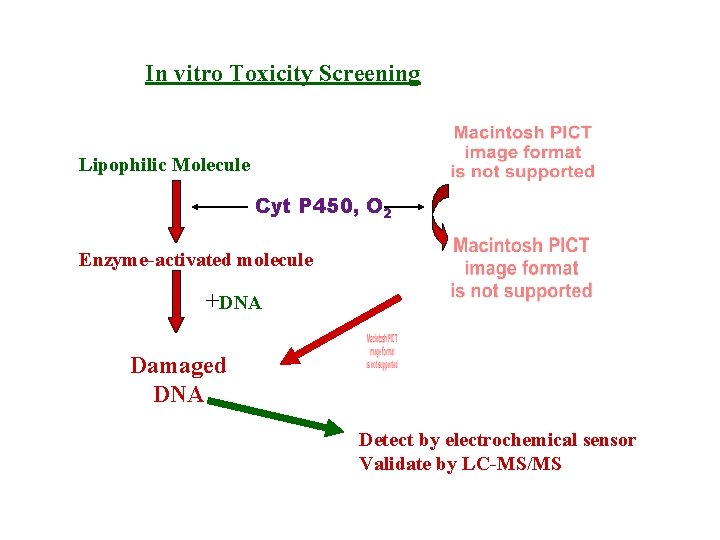
In vitro Toxicity Screening Lipophilic Molecule Cyt P 450, O 2 Enzyme-activated molecule +DNA Damaged DNA Detect by electrochemical sensor Validate by LC-MS/MS

Collaboration with Prof. John Schenkman, Pharmacology, Uconn Health Center Funding from NIH, NIEHS
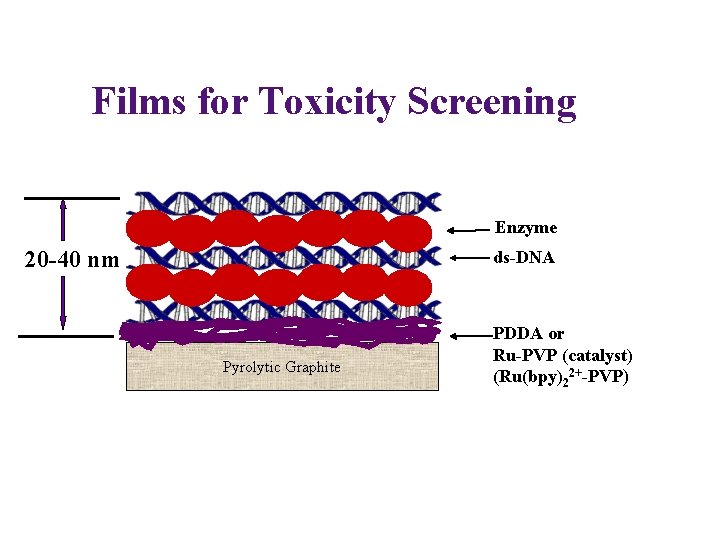
Films for Toxicity Screening Enzyme 20 -40 nm ds-DNA Pyrolytic Graphite PDDA or Ru-PVP (catalyst) (Ru(bpy)22+-PVP)

Layer-by-layer Film assembly Lvov, Decher Lvov, Y. in Nalwa, R. W. ; Ed. ; Handbook Of Surfaces And Interfaces Of Materials, Vol. 3. Academic, 2001, pp. 170 -189. Stable, easily prepared, versatile
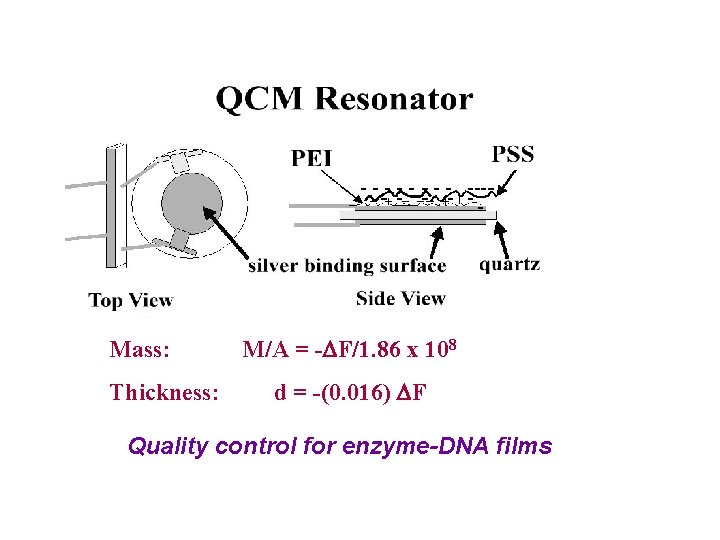
Mass: Thickness: M/A = -DF/1. 86 x 108 d = -(0. 016) DF Quality control for enzyme-DNA films

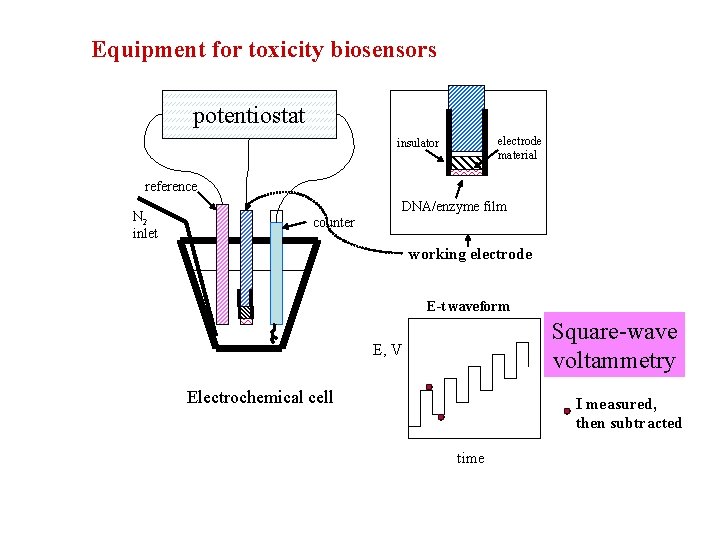
Equipment for toxicity biosensors potentiostat electrode material insulator reference N 2 inlet DNA/enzyme film counter working electrode E-t waveform Square-wave voltammetry E, V Electrochemical cell I measured, then subtracted time
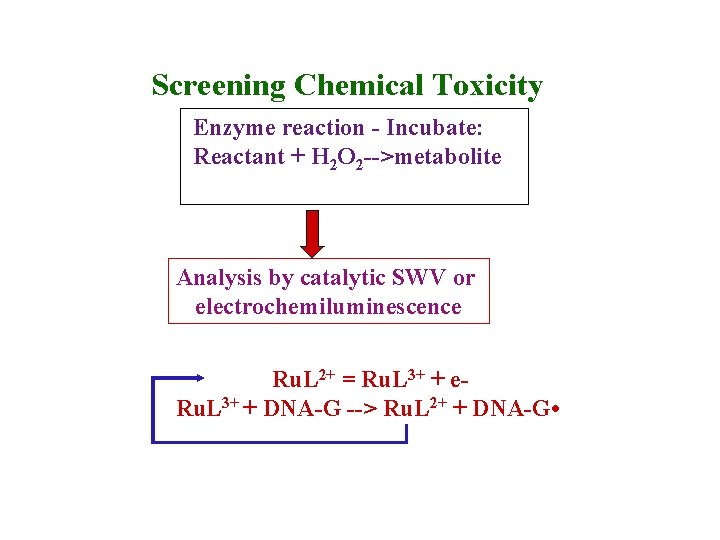
Screening Chemical Toxicity Enzyme reaction - Incubate: Reactant + H 2 O 2 -->metabolite Analysis by catalytic SWV or electrochemiluminescence Ru. L 2+ = Ru. L 3+ + e. Ru. L 3+ + DNA-G --> Ru. L 2+ + DNA-G •

Enzyme/DNA films Peak increase measures damage of DNA by enzymegenerated metabolite

Sensor response vs. reaction time + H 2 O 2
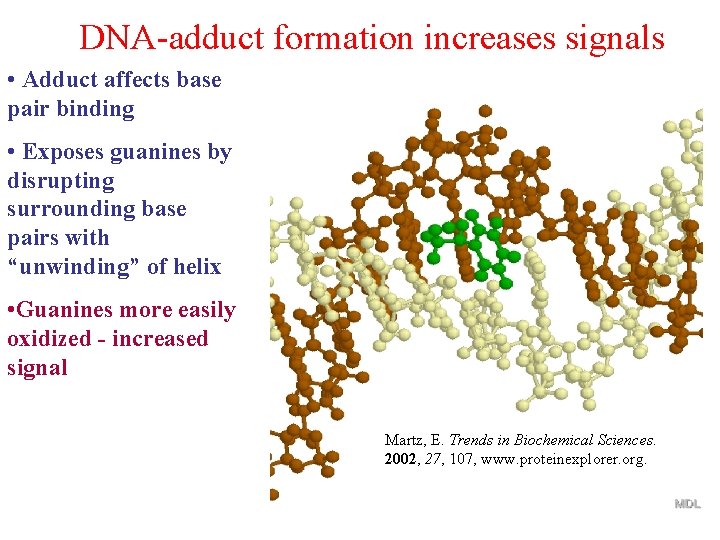
DNA-adduct formation increases signals • Adduct affects base pair binding • Exposes guanines by disrupting surrounding base pairs with “unwinding” of helix • Guanines more easily oxidized - increased signal Martz, E. Trends in Biochemical Sciences. 2002, 27, 107, www. proteinexplorer. org.

Comparison of toxicity sensors with LC-MS For DNA damage by methylmethane sulfonate


Equipment for ECL toxicity sensors ECL = electrochemiluminence Catalytic ECL polymer

Incubations with styrene oxide DNA +

Incubation of Ru-PVP/DNA films with styrene oxide

Direct ECL generation from DNA RVP-Ru. L 2+ = PVP-Ru. L 3+ + e. PVP-Ru. L 3+ + DNA-G --> PVP-Ru. L 2+ + DNA-G • Then? PVP-Ru. L 3+ oxidizes DNA-G • to give Photoexcited PVP-[Ru. L 2+]* Or DNA-G • reduces PVP-Ru. L 2+ to PVP-Ru. L+, PVP-Ru. L 3+ + PVP-Ru. L+ --> PVP-[Ru. L 2+]*
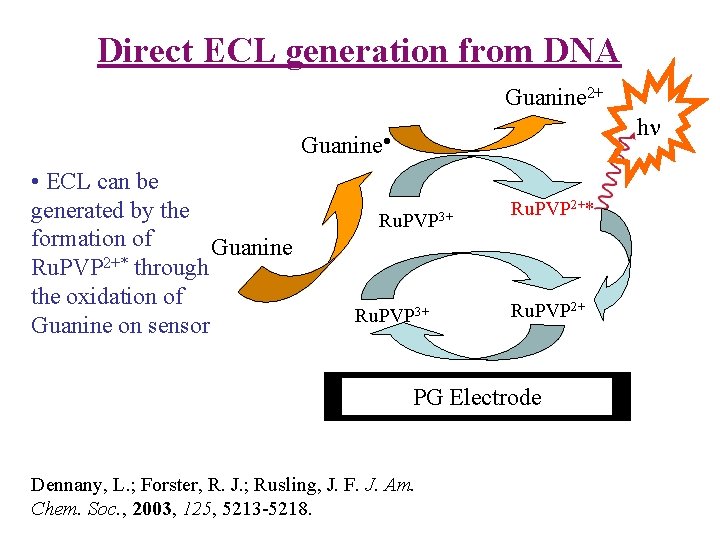
Direct ECL generation from DNA Guanine 2+ h Guanine • ECL can be generated by the formation of Guanine 2+* Ru. PVP through the oxidation of Guanine on sensor Ru. PVP 3+ Ru. PVP 2+* Ru. PVP 2+ PG Electrode Dennany, L. ; Forster, R. J. ; Rusling, J. F. J. Am. Chem. Soc. , 2003, 125, 5213 -5218.
![Benzo[a]Pyrene-DNA Adduct Formation • BP Metabolism much more convoluted BP • Stable adducts formed Benzo[a]Pyrene-DNA Adduct Formation • BP Metabolism much more convoluted BP • Stable adducts formed](http://slidetodoc.com/presentation_image_h2/062b772676e2ee72a0492c6d4f55e5b8/image-27.jpg)
Benzo[a]Pyrene-DNA Adduct Formation • BP Metabolism much more convoluted BP • Stable adducts formed from anti. BPDE 1, 2 7, 8 epoxide • Depurinating adducts formed from oneelectron oxidized BP 2 anti-BPDE 9, 10 epoxide Cation radical • G T Transversions of DNA casued by depurinated bases 2 N 2 d. G adduct- Osborne, M. et al. Benzopyrenes. Cambridge Univ. Press, 1987. 1 Neilson, A. H. PAH’s and Other Related Compounds, Spriger, 1998. 2 N 7 d. G adduct- C 8 d. G adduct- A 7 d. G adduct-
![Arrays: Which Liver Cytochrome P 450 s generate toxic Benzo[a]pyrene Metabolites? Arrays: Which Liver Cytochrome P 450 s generate toxic Benzo[a]pyrene Metabolites?](http://slidetodoc.com/presentation_image_h2/062b772676e2ee72a0492c6d4f55e5b8/image-28.jpg)
Arrays: Which Liver Cytochrome P 450 s generate toxic Benzo[a]pyrene Metabolites?

Arrays detect in-vitro DNA damage from metabolites of different enzymes in DNA/enzyme films Rel. turnover rate, 1/min (nmol E) Mb 0. 9 P 450 cam 3. 0 P 450 1 A 2 3. 5 Drug discovery applications

Sensors for oxidative stress via oxidized DNA Os-PVP PSS Pyrolytic Graphite Ru-PVP ECL detection in films: Lynn Dennany, Robert J. Forster, Blanaid White, Malcolm Smyth and James F. Rusling, Am. Chem. Soc. , 2004, 126, 8835 -8841.

Summary: DNA damage detection/toxicity sensors • Catalytic voltammetry and ECL toxicity sensors • sensors produce metabolites, damage DNA • Can detect 5 -10 damaged bases/10, 000 • can detect DNA oxidation - 8 -oxoguanine (1/6000) • Future: extensions to many compounds, cyt P 450 arrays, ECL arrays, drug toxicity
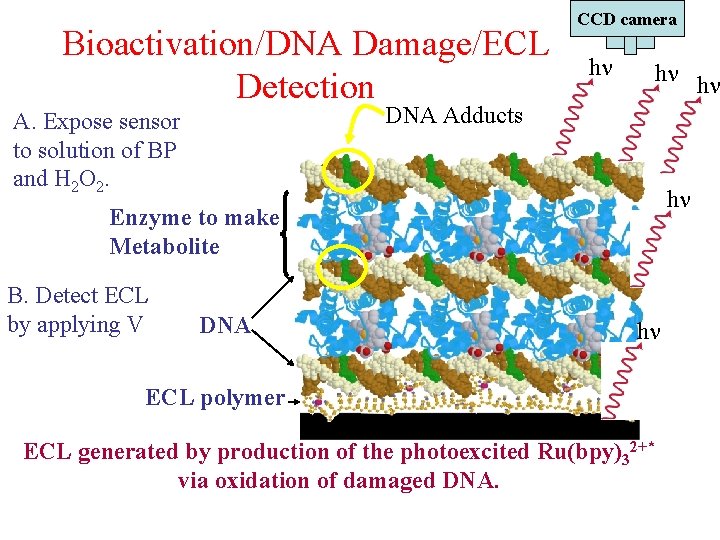
Bioactivation/DNA Damage/ECL Detection CCD camera h h h DNA Adducts A. Expose sensor to solution of BP and H 2 O 2. h Enzyme to make Metabolite B. Detect ECL by applying V DNA h ECL polymer ECL generated by production of the photoexcited Ru(bpy)32+* via oxidation of damaged DNA.
Equipment for ECL Arrays

The ECL Array on graphite w/ Enzyme Exposed to BP + H 2 O 2 • Cyt P 450 1 B 1 enzyme in all spots (1 mg/m. L) • 49 spots allows monitoring damage response of cyt 1 B 1 to BP (100 M) activated with H 2 O 2 (0. 5 m. M) + control • Apply voltage to generate ECL; ± 10% reproducibility Eli Hvastkovs
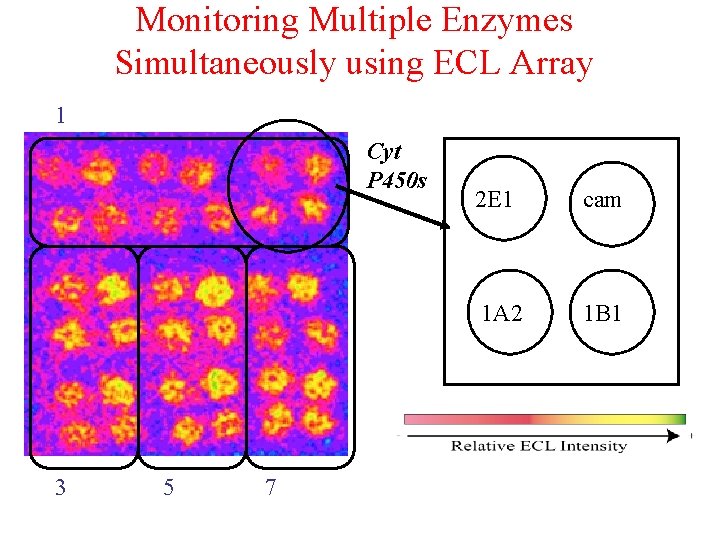
Monitoring Multiple Enzymes Simultaneously using ECL Array 1 Cyt P 450 s 3 5 7 2 E 1 cam 1 A 2 1 B 1
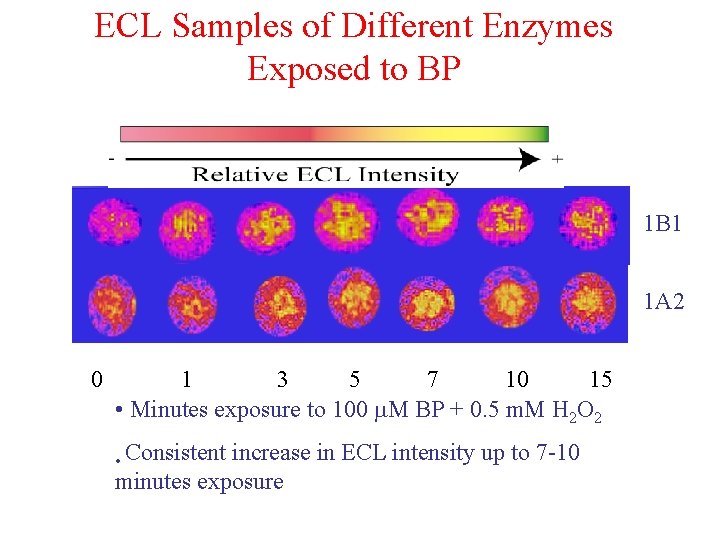
ECL Samples of Different Enzymes Exposed to BP 1 B 1 1 A 2 0 1 3 5 7 10 15 • Minutes exposure to 100 M BP + 0. 5 m. M H 2 O 2 • Consistent increase in ECL intensity up to 7 -10 minutes exposure

Toxicity screening sensors and arrays • Electrochemical, ECL, LC-MS toxicity sensors • Sensors produce metabolites, detect DNA damage • Detect relative rates of DNA damage beginning at 5 -10 damaged bases/10, 000 - in several min. • Multienzyme arrays • Cap. LC-MS rates of DNA damage with fmol detection limits of nucleobase adducts • Future: commercialization? MS kinetics arrays?

Future of bioanalysis with arrays • high throughput, speed • DNA microarrays for genetic analysis - most developed • proteomics arrays based on fluorescence • cyt P 450 genotoxicity arrays • future - arrays for disease diagnostics and treatment • future - electrochemical and ECL arrays for proteins

Thanks to NIH, NSF and ARO for funding! Thanks to all our coworkers and collaborators http: //web. uconn. edu/rusling/ Thanks to YOU for listening! Thanks to intangible creative factors +
- Slides: 39