Design and Use of Repeat Masker Jeremy Buhler

Design and Use of Repeat. Masker Jeremy Buhler HHMI / BIO 4342 Tutorial Workshop

Parts of Repeat. Masker n Programs n n n Smit AFA, Hubley R, and Green P. “Repeat. Masker-Open 3. 0. ” 1996 -2004. http: //www. repeatmasker. org. Cross. Match / WU-BLAST for comparisons Data n Rep. Base library n http: //www. girinst. org

Overview n Sources of repetitive sequence data n How Repeat. Masker finds repeats n Issues and limitations

Data Source n Uses a library of known repeat seqs n Supplied by Rep. Base project n Repeats in Rep. Base are carefully curated, typically by hand.

An example summary report for a repeat family published by Rep. Base

Consensus Sequences n A repeat family is usually summarized by a consensus sequence accgataggtatacgtatca-tttacgatac atcgct-ggtttacgcgtcaattcaggatgc accggt-tgtttacgtagcaatctaggatac accgat-ggtttacgtatcaatttaggatac

Why Consensus Sequences? n n Faster to compare one sequence to genome than many Consensus can actually be better than individual instances for discovering new copies of a repeat.

Utility of Consensus New repeat copy is closer to consensus than actggt to other copies! acacgt 3 3 acaggt tcaggc 2 4 atagct

Types of Repeats in Library n Interspersed (Alu, LINE, MIR, …) n Simple (agag, atcatcatc, …) n Micro- and mini-satellites n Noncoding RNAs (t. RNA, r. RNA, sn. RNA, …) n Common contaminants (E. coli, vectors)
Overview n Sources of repetitive sequence data n How Repeat. Masker finds repeats n Issues and limitations

The Basics n Uses BLAST-like tool to compare libraries to query sequence n Cross-Match (P. Green) – traditional n WU-BLAST (W. Gish) – 10 x faster!

Partial Repeats n n n Repeat. Masker will cheerfully report an incomplete match to a repeat. Detects best-conserved parts Some repeats (retroposons) typically incomplete
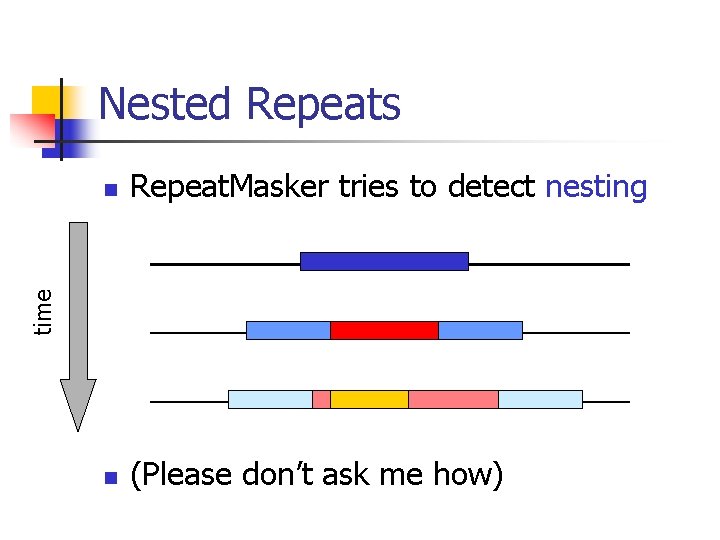
Nested Repeats Repeat. Masker tries to detect nesting n (Please don’t ask me how) time n

Overview n Sources of repetitive sequence data n How Repeat. Masker finds repeats n Issues and limitations

Library Choice n n n Make sure to use correct libraries for your target species (Commonly used organisms have preselected library lists) Danger: mis-identifications!

Incomplete Masking n n Highly diverged repeats can be tough to find Might leave ends of a repeat unmasked (masked) n BLAST hit Is this really a new feature?

Use the Right Tool n Tandem repeats and duplications n n n RNA n n Dust (short) TRF (long) t. RNAScan, Infernal, … Low-copy (chr-specific, inverted, …) n BLAST?

In conclusion… Hey, let’s be careful out there!
- Slides: 18